barbieQ_paper_Figure3_FigureS3: Type I
error rate and power using Mixture data
Liyang Fei
Initiate: 2025-09-07
Last update: 2026-01-11
Last updated: 2026-01-11
Checks: 5 2
Knit directory: public_barcode_count/
This reproducible R Markdown analysis was created with workflowr (version 1.7.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of
the R Markdown file created these results, you’ll want to first commit
it to the Git repo. If you’re still working on the analysis, you can
ignore this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
The global environment had objects present when the code in the R
Markdown file was run. These objects can affect the analysis in your R
Markdown file in unknown ways. For reproduciblity it’s best to always
run the code in an empty environment. Use wflow_publish or
wflow_build to ensure that the code is always run in an
empty environment.
The following objects were defined in the global environment when these results were created:
| Name | Class | Size |
|---|---|---|
| module | function | 5.6 Kb |
The command set.seed(20250112) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version ecc9192. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .RData
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: public_barcode_count.Rproj
Unstaged changes:
Modified: analysis/barbieQ_paper_Figure3.Rmd
Modified: output/f3.png
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/barbieQ_paper_Figure3.Rmd)
and HTML (docs/barbieQ_paper_Figure3.html) files. If you’ve
configured a remote Git repository (see ?wflow_git_remote),
click on the hyperlinks in the table below to view the files as they
were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | 670258c | FeiLiyang | 2026-01-07 | minor change |
| Rmd | f48add2 | FeiLiyang | 2026-01-07 | supple analyses |
| Rmd | 34f5894 | FeiLiyang | 2026-01-01 | reorder f3 |
| html | 34f5894 | FeiLiyang | 2026-01-01 | reorder f3 |
| html | a2e7a36 | FeiLiyang | 2025-12-17 | fix yml |
| Rmd | 7b7813d | FeiLiyang | 2025-12-17 | update f3 fs4 |
| html | 7b7813d | FeiLiyang | 2025-12-17 | update f3 fs4 |
| Rmd | c2766f9 | Liyang Fei | 2025-10-07 | remake figures |
1 Load Dependencies
library(readxl)
library(magrittr)
library(dplyr)
library(tidyr) # for pivot_longer
library(tibble) # for rownames_to _column
library(knitr) # for kable()
library(data.table) # for data.table()
library(SummarizedExperiment)
library(S4Vectors)
library(grid)
library(ggplot2)
library(ggbreak)
library(ggnewscale) # ggnewscale::new_scale_fill()
library(patchwork)
library(scales)
library(ggVennDiagram)
library(ComplexHeatmap)
library(magick) # image_read()
library(eulerr)
library(edgeR)
source("analysis/plotBarcodeHistogram.R") ## accommodated from bartools::plotBarcodehistogram
source("analysis/F3_simulation.R") ## for negative simulation
source("analysis/ggplot_theme.R") ## setting ggplot theme2 Install
barbieQ
Installing the latest devel version of barbieQ from
GitHub.
if (!requireNamespace("barbieQ", quietly = TRUE)) {
remotes::install_github("Oshlack/barbieQ")
}Warning: replacing previous import 'data.table::first' by 'dplyr::first' when
loading 'barbieQ'Warning: replacing previous import 'data.table::last' by 'dplyr::last' when
loading 'barbieQ'Warning: replacing previous import 'data.table::between' by 'dplyr::between'
when loading 'barbieQ'Warning: replacing previous import 'dplyr::as_data_frame' by
'igraph::as_data_frame' when loading 'barbieQ'Warning: replacing previous import 'dplyr::groups' by 'igraph::groups' when
loading 'barbieQ'Warning: replacing previous import 'dplyr::union' by 'igraph::union' when
loading 'barbieQ'library(barbieQ)Check the version of barbieQ.
packageVersion("barbieQ")[1] '1.1.3'3 Set seeds
set.seed(2025)4 Load data
load("output/mixture_barbieQ.rda")5 F3A: Experimental schematic
schematic <- image_read("output/f3a_schematic.png")
f3a <- rasterGrob(schematic, interpolate = FALSE)
wrap_elements(full = f3a)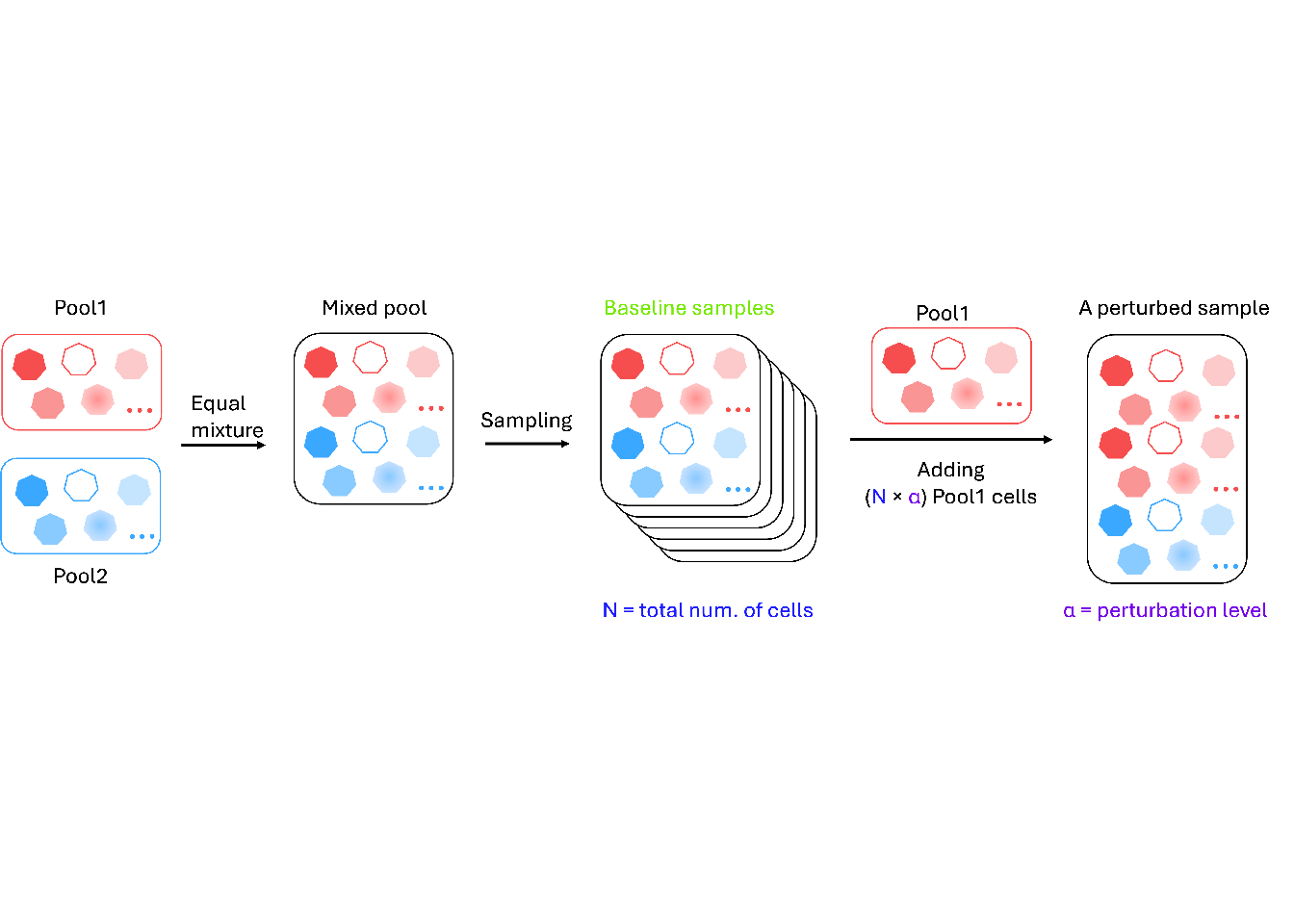
| Version | Author | Date |
|---|---|---|
| dfa737a | FeiLiyang | 2025-12-17 |
6 F3B: Plot barcode proportion and sample annotation
plotBarcodeHistogram, sourced from
“plotBarcodeHistogram.R”, is customized from the bartools
package.
## pull out sample designs
design_mixed <- mixture_plus1_top$sampleMetadata %>% as.data.frame()
design_mixed$sample <- colnames(mixture_plus1_top)
str(design_mixed)'data.frame': 38 obs. of 5 variables:
$ Subset : Factor w/ 4 levels "premix1","premix2",..: 1 2 3 3 3 3 3 3 3 3 ...
$ BarcodeSize : num NA NA 660000 660000 330000 330000 160000 160000 80000 80000 ...
$ Perturbation: num NA NA 0 0 0 0 0 0 0 0 ...
$ TechRep : Factor w/ 2 levels "1","2": 1 1 1 2 1 2 1 2 1 2 ...
$ sample : chr "P1" "P2" "null_660.1" "null_660.2" ...design_mixed$y = 1.05
color_mapping <- mixture_plus1_top@metadata$factorColors
## order samples
order_sample <- design_mixed %>% with(order(Subset, Perturbation, BarcodeSize))
mixed_dge <- DGEList(
counts = assay(mixture_plus1_top),
group = mixture_plus1_top$sampleMetadata$Subset)
## pull out ggplot data
p_mixed <- plotBarcodeHistogram(mixed_dge, orderSamples = colnames(mixture_plus1_top)[order_sample])Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"barcode"` instead of `.data$barcode`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"value"` instead of `.data$value`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"name"` instead of `.data$name`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"freq"` instead of `.data$freq`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.p_mixed_illus <- p_mixed +
## label subset
ggnewscale::new_scale_fill() +
geom_tile(data = design_mixed, aes(x = sample,y = y, fill = Subset), height = 0.04, inherit.aes = FALSE, color = "white") +
scale_fill_manual(values = color_mapping$Subset, labels = c(null = "baseline", perturb = "perturbed")) +
labs(fill = "Subset") +
guides(fill = guide_legend(ncol = 2)) +
## label perturbations
ggnewscale::new_scale_fill() +
geom_tile(data = design_mixed, aes(x = sample,y = y+0.05, fill = as.factor(Perturbation)), height = 0.04, inherit.aes = FALSE, color = "white") +
scale_fill_manual(values = color_mapping$Perturbation) +
labs(fill = "Perturbation level") +
guides(fill = guide_legend(ncol = 2)) +
## label libsize
ggnewscale::new_scale_fill() +
geom_tile(data = design_mixed, aes(x = sample,y = y+0.10, fill = as.factor(BarcodeSize)), height = 0.04, inherit.aes = FALSE, color = "white") +
scale_fill_manual(values = color_mapping$BarcodeSize)+
labs(fill = "Num. of cells") +
guides(fill = guide_legend(ncol = 2)) +
labs(x = "Sample", y = "Barcode proportion") +
theme_minimal() +
scale_y_continuous(breaks = seq(0, 1, by = 0.2)) +
theme(axis.text.x = element_blank(), panel.grid.major.x = element_blank())
f3b <- p_mixed_illus +
theme(legend.title = element_text(size = 12), legend.text = element_text(size = 11), aspect.ratio = 1,
axis.title = element_text(size = 13), axis.text = element_text(size = 12)) +
guides(fill = guide_legend(ncol = 2))
f3bWarning: The `guide` argument in `scale_*()` cannot be `FALSE`. This was deprecated in
ggplot2 3.3.4.
ℹ Please use "none" instead.
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.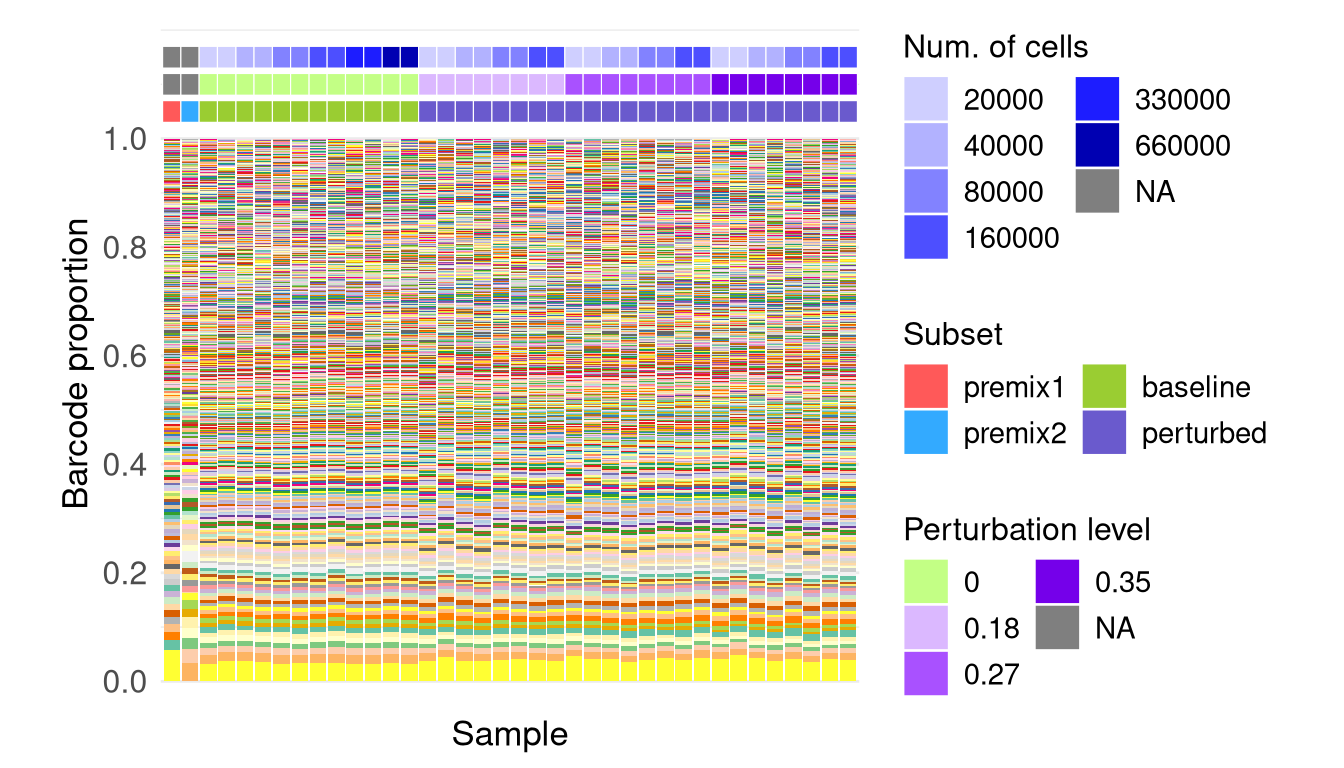
7 FPR
Using mixture_plus1_top data, where an offset (one) was added to the raw count, based on which proportion was calculated. However the “top” barcodes were filtered based on the raw count data.
Here we subset the 12 “null” samples for evaluating FPR of diff_prop, and diff_occ methods.
Three transformations with dif_prop were evaluated separately.
- avoid running this chunk during rendering.
global_all_loopswas saved from the first run.
nulls <- mixture_plus1_top[, mixture_plus1_top$sampleMetadata$Subset == "null"]
results_negsi <- suppressMessages({
random_sampling(barbieQ = nulls, loop_times = 100)
})
## `random_sampling()` save the simulations to global environment
## as `global_all_loops`
save(global_all_loops, file = "output/mixture_negative_simulation.rda")
results_negsi[[1]] <- results_negsi[[1]] + ggtitle("Prop_asin, perturb_null")
results_negsi[[2]] <- results_negsi[[2]] + ggtitle("Prop_logit, perturb_null")
results_negsi[[3]] <- results_negsi[[3]] + ggtitle("Prop_noTrans, perturb_null")
results_negsi[[4]] <- results_negsi[[4]] + ggtitle("Occ_firth, perturb_null")
results_negsi[[5]] <- results_negsi[[5]] + ggtitle("Prop_asin, perturb_null")
results_negsi[[6]] <- results_negsi[[6]] + ggtitle("Prop_logit, perturb_null")
results_negsi[[7]] <- results_negsi[[7]] + ggtitle("Prop_noTrans, perturb_null")
results_negsi[[8]] <- results_negsi[[8]] + ggtitle("Occ_firth, perturb_null")
wrap_plots(results_negsi[1:8], ncol = 4)7.1 load results
## loading random sampling results to avoid running it repeatedly
load("output/mixture_negative_simulation.rda") # extract results from the loops
end_sampling <- 6
## extract FPR
all_FPR <- lapply(seq(3:end_sampling), function(n) {
global_all_loops[[n]]$FPR
})
df_FPR <- do.call(rbind, all_FPR)
## extract P.Val
all_P.Val <- lapply(seq(3:end_sampling), function(n) {
global_all_loops[[n]]$P.Val
})
df_P.Val <- do.call(rbind, all_P.Val)
## extract MA
all_MA <- lapply(seq(3:end_sampling), function(n) {
global_all_loops[[n]]$MA
})
df_MA <- do.call(rbind, all_MA)
all_Group_Vec <- lapply(seq(3:end_sampling), function(n) {
global_all_loops[[n]]$Group_Vec
})
df_Group_Vec <- do.call(rbind, all_Group_Vec)
stats_barcodes <- cbind(df_P.Val[,1:4], df_MA)7.2 F3C: Type I Error Rate
- Diff_prop
df_FPR$N_Samples <- as.factor(df_FPR$N_Samples)
## reshape the data to fit methods for testing
df_FPR_long <- df_FPR %>%
pivot_longer(cols = starts_with("Prop_"),
names_to = "Method",
values_to = "Pval_Prop") %>%
mutate(Method = factor(Method, levels = c("Prop_asin", "Prop_logit", "Prop_noTrans"),
labels = c("asin-sqrt", "logit", "no trans.")))
P_FPR_add1_prop <- ggplot(
df_FPR_long, aes(x = factor(N_Samples), y = Pval_Prop, color = Method)) +
geom_boxplot(outlier.shape = 1, position = position_dodge(width = 0.7)) +
# geom_jitter(size = 1, position = position_dodge(width = 0.7)) +
geom_hline(yintercept = 0.05, linetype = "dashed", color = "black") +
# coord_cartesian(ylim = c(0, 0.2)) +
labs(
y = "Fraction of P-Value < 0.05",
x = "Number of samples per group",
title = "Type I error rate",
subtitle = "Differential proportion test") +
# facet_wrap(~Method, scales = "free_y") +
theme_classic() +
theme(legend.position = "top")- Diff_occ. when converting barcode presence / absence, setting the threshold as CPM>10 instead of the default 1, as an offset (one) was added to the raw count preprocessing.
P_FPR_occ <- ggplot(
df_FPR, aes(x = factor(N_Samples), y = Occ_firth)) +
geom_boxplot(outlier.shape = 1, position = position_dodge(width = 0.7)) +
# geom_jitter(size = 1, position = position_dodge(width = 0.7)) +
geom_hline(yintercept = 0.05, linetype = "dashed", color = "black") +
scale_color_manual(values = c("Differential occurrrence test" = "black")) +
# coord_cartesian(ylim = c(0, 0.2)) +
labs(
y = "Fraction of P-Value < 0.05",
x = "Number of samples per group",
titlte = "",
subtitle = "Differential occurrence test") +
theme_classic() +
theme(legend.position = "top")f3c <- (P_FPR_add1_prop + theme_barbieQ_size() + theme(legend.position = "top")) +
(P_FPR_occ + theme_barbieQ_size())
f3cIgnoring unknown labels:
• titlte : ""Warning: No shared levels found between `names(values)` of the manual scale and the
data's colour values.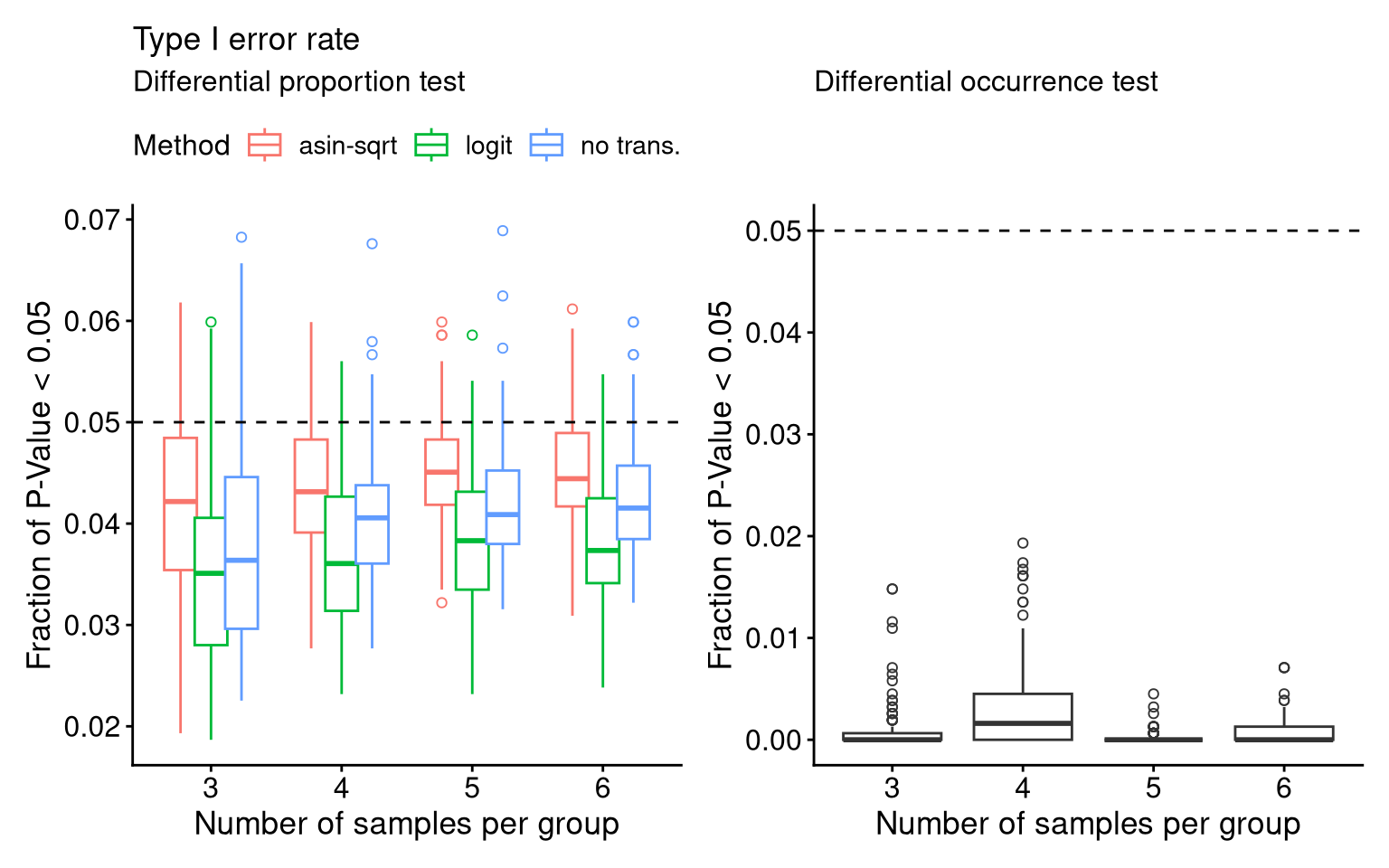
7.3 F3H: Type I Error Rate as a function to N_BC
7.3.1 Generate subsets of barcodes
## with different filtering thresholds, how many "top" barcodes do we get
1:9/10[1] 0.1 0.2 0.3 0.4 0.5 0.6 0.7 0.8 0.9lapply(
c(1:9/10, 0.95),
function(filter_level) {
mixture_plus1 <- tagTopBarcodes(mixture_plus1, nSampleThreshold = 8, proportionThreshold = filter_level)
paste0("filter_thresh=", filter_level, "; N_barcodes=", sum(rowData(mixture_plus1)$isTopBarcode$isTop)) %>% print()
# plotBarcodeSankey(mixture)
return(
sum(rowData(mixture_plus1)$isTopBarcode$isTop)
)
}
) %>% unlist() %>%
setNames(nm = c(1:9/10, 0.95)) ->
filter_to_N[1] "filter_thresh=0.1; N_barcodes=12"
[1] "filter_thresh=0.2; N_barcodes=32"
[1] "filter_thresh=0.3; N_barcodes=64"
[1] "filter_thresh=0.4; N_barcodes=101"
[1] "filter_thresh=0.5; N_barcodes=169"
[1] "filter_thresh=0.6; N_barcodes=269"
[1] "filter_thresh=0.7; N_barcodes=394"
[1] "filter_thresh=0.8; N_barcodes=617"
[1] "filter_thresh=0.9; N_barcodes=1060"
[1] "filter_thresh=0.95; N_barcodes=1553"filter_to_N 0.1 0.2 0.3 0.4 0.5 0.6 0.7 0.8 0.9 0.95
12 32 64 101 169 269 394 617 1060 1553 names(filter_to_N) %>% str() chr [1:10] "0.1" "0.2" "0.3" "0.4" "0.5" "0.6" "0.7" "0.8" "0.9" "0.95"7.3.2 Update proportion
## Compute medians per bin/method/N_Samples
method_labels <- c(
"Prop_asin" = "asin-sqrt",
"Prop_logit" = "logit",
"Prop_noTrans" = "none"
)- avoid running this chunk during rendering.
stats_barcodes_all_levelswas saved from the first run.
stats_barcodes_all_levels <- lapply(
c(1:9/10, 0.95),
function(i) {
stats_barcodes_i <- get_null_sample_by_filter(mixture_plus1, i, loop_times = 100)
stats_barcodes_i$filter_level <- i
stats_barcodes_i$N_BC <- filter_to_N[i %>% as.character()]
return(stats_barcodes_i)
}
)
stats_barcodes_all_levels <- do.call(rbind, stats_barcodes_all_levels)
save(stats_barcodes_all_levels, file = "output/fdr_by_Nbc.rda")load("output/fdr_by_Nbc.rda")
FPR_box <- stats_barcodes_all_levels %>%
pivot_longer(
cols = c(Prop_asin, Prop_logit, Prop_noTrans),
names_to = "method",
values_to = "pvalue"
) %>%
group_by(method, N_Samples, N_BC, Loop_N) %>%
summarize(N_signif = sum(pvalue < 0.05), Frac_signif = sum(pvalue < 0.05) / n(), .groups = "drop") %>%
mutate(Method = recode(method, !!!method_labels))
ggplot(FPR_box, aes(x = as.factor(N_BC), y = Frac_signif, color = Method, group = paste0(N_BC, Method))) +
geom_boxplot(alpha = 0.6, position = position_dodge(), outliers = T) +
labs(
x = paste0("Number of total barcodes after filtering"),
y = "Fraction of P-Value < 0.05", color = "Method", shape = "N_sample", linetype = "N_sample",
title = "Type I Error Rate as a function to number of barcodes"
) +
geom_hline(yintercept = 0.05, linetype = "dashed") +
# facet_wrap(~N_Samples, ncol =4) +
theme_linedraw()+
theme(axis.text.x = element_text(angle = 0, vjust = 0.5, size = 10, hjust = 0.5), aspect.ratio = 0.5) +
theme(
legend.title = element_text(size = 12), legend.text = element_text(size = 11),
axis.title = element_text(size = 13), axis.text = element_text(size = 12)) +
scale_y_continuous(limits = c(0,0.2))Ignoring unknown labels:
• shape : "N_sample"
• linetype : "N_sample"Warning: Removed 80 rows containing non-finite outside the scale range
(`stat_boxplot()`).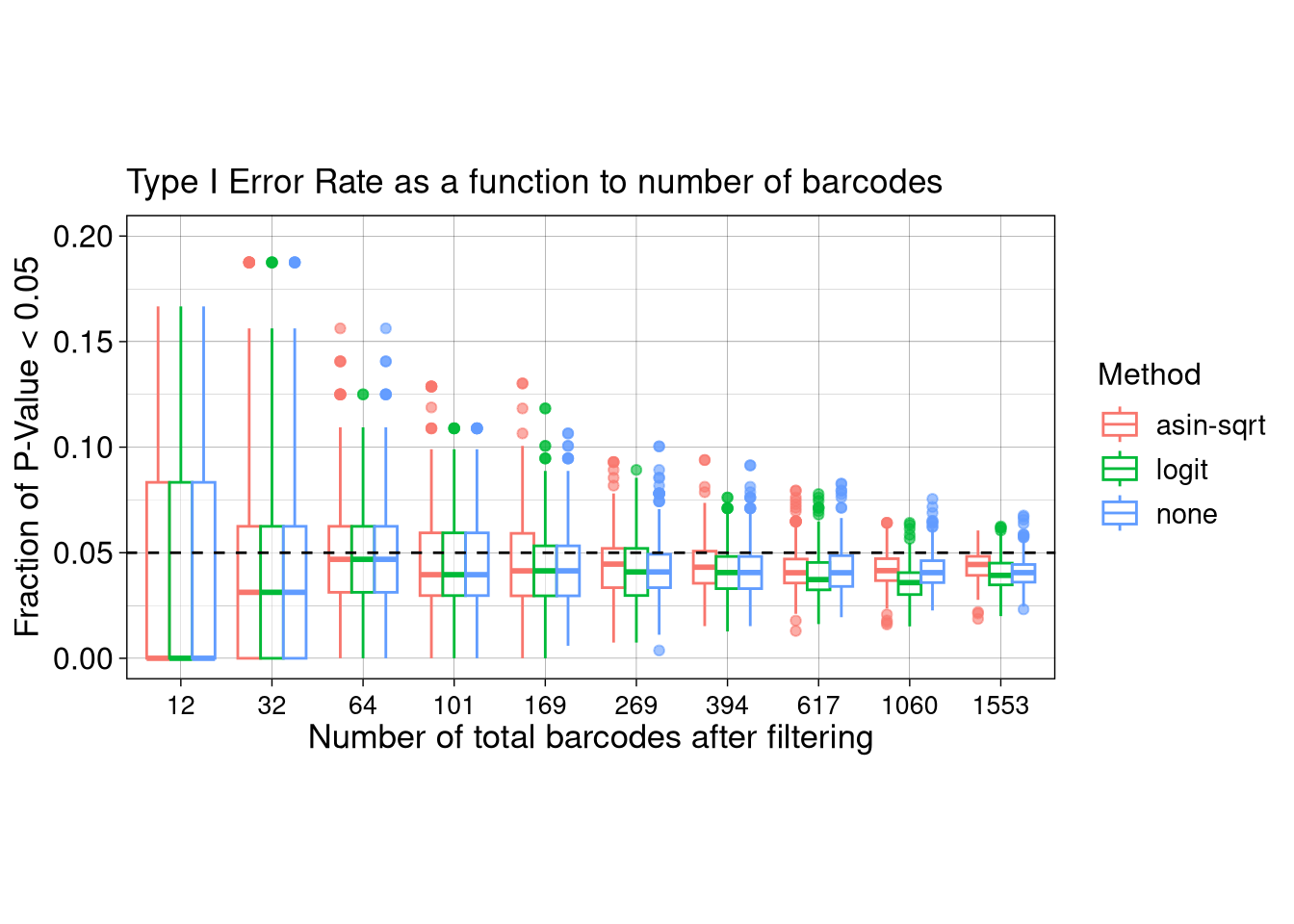
7.3.3 Raw proportion
## saving all p.vals from the N=1553 simulation
# df_P.Val
mixture_plus1 <- tagTopBarcodes(mixture_plus1, nSampleThreshold = 8, proportionThreshold = 0.95)
## get the subset of barcodes
lapply(c(1:9/10, 0.95), function(filter_level) {
barbieQ <- tagTopBarcodes(mixture_plus1, nSampleThreshold = 8, proportionThreshold = filter_level)
tag <- rowData(barbieQ)$isTopBarcode$isTop
tag <- tag[mixture_plus1@elementMetadata$isTopBarcode$isTop]
return(tag)
}) -> BC_set_by_filter
BC_set_by_filter <- do.call(cbind, BC_set_by_filter)
colnames(BC_set_by_filter) <- filter_to_N[c(1:9/10, 0.95) %>% as.character()]
## check
colSums(BC_set_by_filter) 12 32 64 101 169 269 394 617 1060 1553
12 32 64 101 169 269 394 617 1060 1553 mean_prop <- rowMeans(mixture_plus1[mixture_plus1@elementMetadata$isTopBarcode$isTop,mixture_plus1$sampleMetadata$Subset=="null"]@assays@data$proportion)
## testing results obtained from raw proportin in stats_barcodes
df_P.Val_filter_tag <- cbind(
stats_barcodes, # testing results obtained from raw proportin
mean_prop,
BC_set_by_filter[
rep(
seq_len(nrow(BC_set_by_filter)),
length.out = nrow(stats_barcodes)
),
]
)Warning in data.frame(..., check.names = FALSE): row names were found from a
short variable and have been discardedFPR_box_raw <- df_P.Val_filter_tag %>%
pivot_longer(
cols = c(Prop_asin, Prop_logit, Prop_noTrans),
names_to = "method",
values_to = "pvalue"
) %>%
pivot_longer(
cols = as.character(filter_to_N[c(1:9/10, 0.95) %>% as.character()]),
names_to = "N_BC",
values_to = "selectedBy"
) %>%
group_by(method, N_Samples, Loop_N, N_BC) %>%
summarize(
N_signif = sum(pvalue < 0.05 & selectedBy),
Frac_signif = sum(pvalue < 0.05 & selectedBy) / sum(selectedBy), .groups = "drop") %>%
mutate(Method = recode(method, !!!method_labels))
FPR_box_raw$N_BC <- as.numeric(FPR_box_raw$N_BC)
ggplot(FPR_box_raw, aes(x= as.factor(N_BC), y = Frac_signif, color = Method, group = paste0(Method, N_BC))) +
geom_boxplot(alpha = 0.6, position = position_dodge(), outliers = T) +
labs(
x = paste0("Number of total barcodes after filtering"),
y = "Fraction of P-Valuev < 0.05", color = "Method", shape = "N_sample", linetype = "N_sample"
) +
geom_hline(yintercept = 0.05, linetype = "dashed") +
# facet_wrap(~N_Samples, ncol =4) +
theme_linedraw()+
theme(axis.text.x = element_text(angle = 0, vjust = 0.5, size = 10, hjust = 0.5), aspect.ratio = 0.5) +
theme_barbieQ_size() +
scale_y_continuous(limits = c(0,0.2)) ## y scale zoom inIgnoring unknown labels:
• shape : "N_sample"
• linetype : "N_sample"Warning: Removed 85 rows containing non-finite outside the scale range
(`stat_boxplot()`).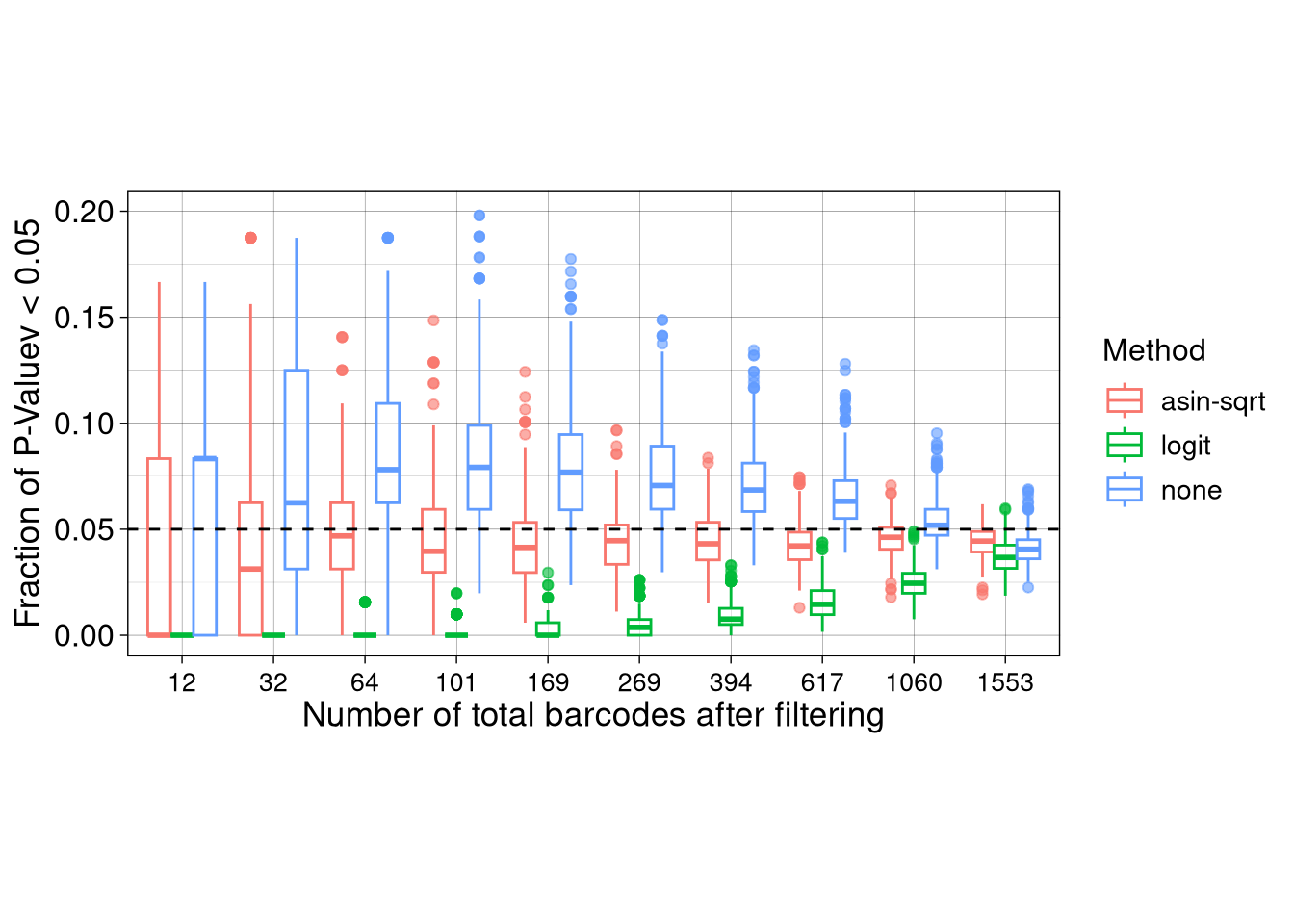
| Version | Author | Date |
|---|---|---|
| 34f5894 | FeiLiyang | 2026-01-01 |
rbind(
data.frame(FPR_box_raw, prop_type = "Raw proportion"),
data.frame(FPR_box, prop_type = "Updated proportion")
) %>%
mutate(prop_type = factor(prop_type, levels = c("Updated proportion", "Raw proportion"))) %>%
## plot
ggplot(aes(x = as.factor(N_BC), y = Frac_signif, color = Method, group = paste0(N_BC, Method))) +
geom_boxplot(alpha = 0.6, position = position_dodge(), outliers = T) +
labs(
x = paste0("Number of total barcodes after filtering"),
y = "Fraction of P-Value < 0.05", color = "Method", shape = "N_sample", linetype = "N_sample",
title = "Type I error rate to number of barcodes"
) +
geom_hline(yintercept = 0.05, linetype = "dashed") +
facet_grid(prop_type~1, switch = "y") +
theme_bw()+
theme_barbieQ_size() +
theme(panel.grid.minor = element_blank(), axis.text.x = element_text(angle = 15, vjust = 0.5, hjust = 0.5),
legend.position = "top",
strip.placement = "outside", strip.text.y = element_text(size = 12),
strip.text.x = element_blank(), strip.background.x = element_blank()) +
scale_y_continuous(limits = c(0,0.2)) -> f3h
## scale_x_discrete(limits = rev) ## reverse x axis
f3hIgnoring unknown labels:
• shape : "N_sample"
• linetype : "N_sample"Warning: Removed 165 rows containing non-finite outside the scale range
(`stat_boxplot()`).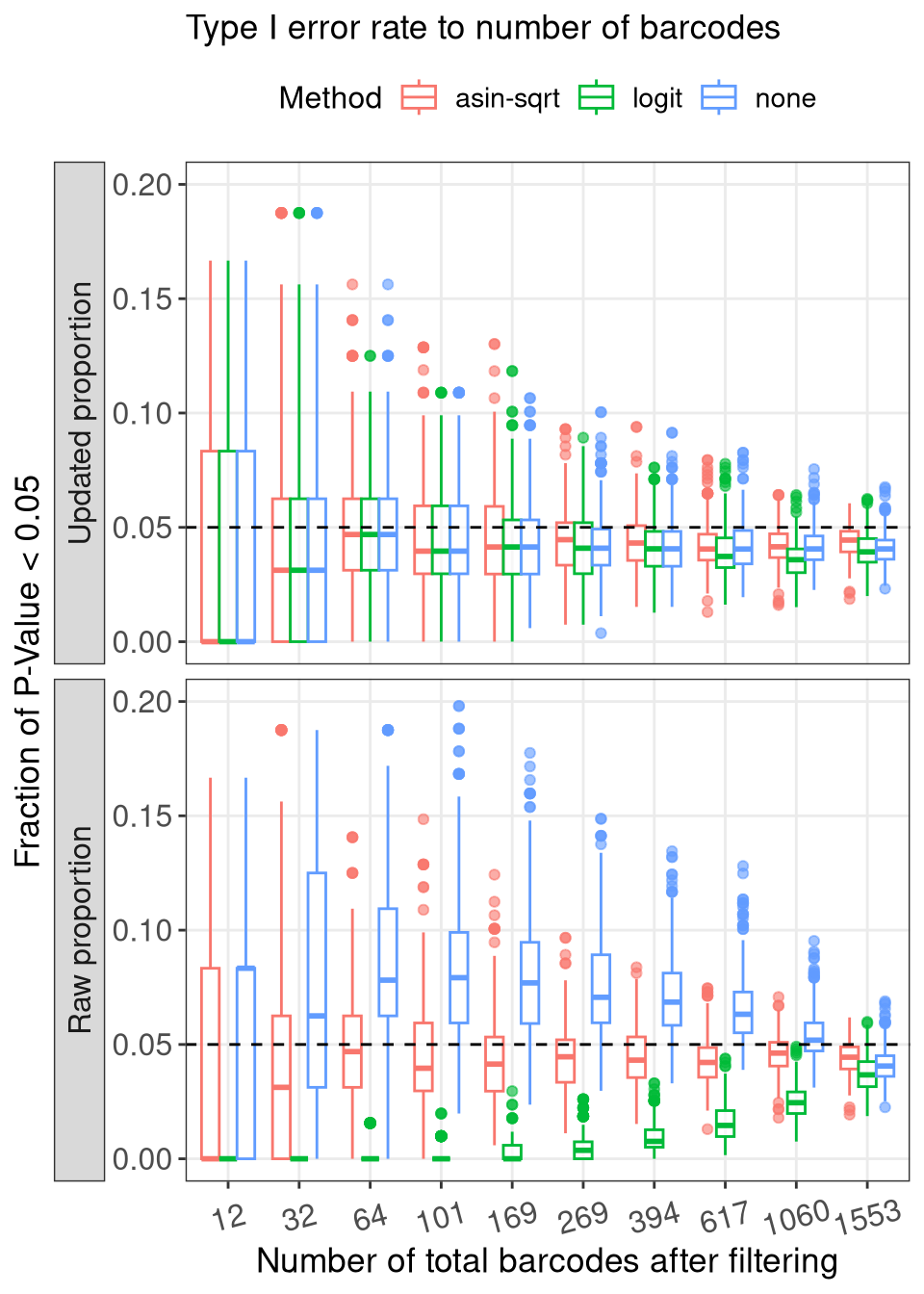
7.4 F3G: Density to N_Barcodes
7.4.1 Updated proportion
## get the subset of barcodes
lapply(c(1:9/10, 0.95), function(filter_level) {
barbieQ <- tagTopBarcodes(mixture_plus1, nSampleThreshold = 8, proportionThreshold = filter_level)
tag <- rowData(barbieQ)$isTopBarcode$isTop
prop_mat <- mixture_plus1[tag,mixture_plus1$sampleMetadata$Subset=="null"]@assays@data$proportion
## update prop
prop_mat <- t(t(prop_mat) / colSums(prop_mat))
df <- data.frame(
prop = as.vector(prop_mat),
N_BC=filter_to_N[as.character(filter_level)]
)
return(df)
}) -> BC_set_prop_filterWarning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discardedBC_set_prop_filter <- do.call(rbind, BC_set_prop_filter)
ggplot(BC_set_prop_filter) +
geom_density(aes(x = prop, color = as.factor(N_BC), group = as.factor(N_BC))) +
scale_x_log10(breaks = 10^seq(-6, -1, by = 1), labels = strip_zeros) +
coord_cartesian(xlim = c(1e-6, 3e-1)) +
labs(color = "Num. BC", x = "Barcode Proportion", y = "Density", subtitle = "with updated proportion") +
theme_barbieQ() +
theme(aspect.ratio = 0.9) -> f3g17.4.2 Raw proportion
## get the subset of barcodes
lapply(c(1:9/10, 0.95), function(filter_level) {
barbieQ <- tagTopBarcodes(mixture_plus1, nSampleThreshold = 8, proportionThreshold = filter_level)
tag <- rowData(barbieQ)$isTopBarcode$isTop
## keep raw prop
prop_mat <- mixture_plus1[tag,mixture_plus1$sampleMetadata$Subset=="null"]@assays@data$proportion
df <- data.frame(
prop = as.vector(prop_mat),
N_BC=filter_to_N[as.character(filter_level)]
)
return(df)
}) -> BC_set_prop_filter_rawWarning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discarded
Warning in data.frame(prop = as.vector(prop_mat), N_BC =
filter_to_N[as.character(filter_level)]): row names were found from a short
variable and have been discardedBC_set_prop_filter_raw <- do.call(rbind, BC_set_prop_filter_raw)
ggplot(BC_set_prop_filter_raw) +
geom_density(aes(x = prop, color = as.factor(N_BC), group = as.factor(N_BC))) +
scale_x_log10(breaks = 10^seq(-6, -1, by = 1), labels = strip_zeros) +
coord_cartesian(xlim = c(1e-6, 3e-1)) +
theme_linedraw() +
labs(color = "Num. BC", x = "Barcode Proportion", y = "Density", subtitle = "with raw proportion") +
theme_barbieQ() +
theme(aspect.ratio = 0.9) -> f3g2rbind(
data.frame(BC_set_prop_filter_raw, prop_type = "Raw proportion"),
data.frame(BC_set_prop_filter, prop_type = "Updated proportion")
) %>%
mutate(prop_type = factor(prop_type, levels = c("Updated proportion", "Raw proportion"))) %>%
## pass to gplot
ggplot() +
geom_density(aes(x = prop, color = as.factor(N_BC), group = as.factor(N_BC))) +
scale_x_log10(breaks = 10^seq(-6, -1, by = 1), labels = strip_zeros) +
coord_cartesian(xlim = c(1e-6, 3e-1)) +
facet_grid(prop_type~1, switch = "y") +
labs(color = "Num. BC", x = "Barcode Proportion", y = "Density", title = "Barcode distribution in datasets") +
theme_bw() +
theme_barbieQ_size() +
theme(panel.grid.minor = element_blank(), axis.text.x = element_text(angle = 15, vjust = 0.5, hjust = 0.5),
legend.position = "top", legend.key.size = unit(0.3, "cm"),
legend.title = element_text(size = 11), legend.text = element_text(size = 10),
strip.placement = "outside", strip.text.y = element_text(size = 12),
strip.text.x = element_blank(), strip.background.x = element_blank()) -> ## disable column label
f3g
f3g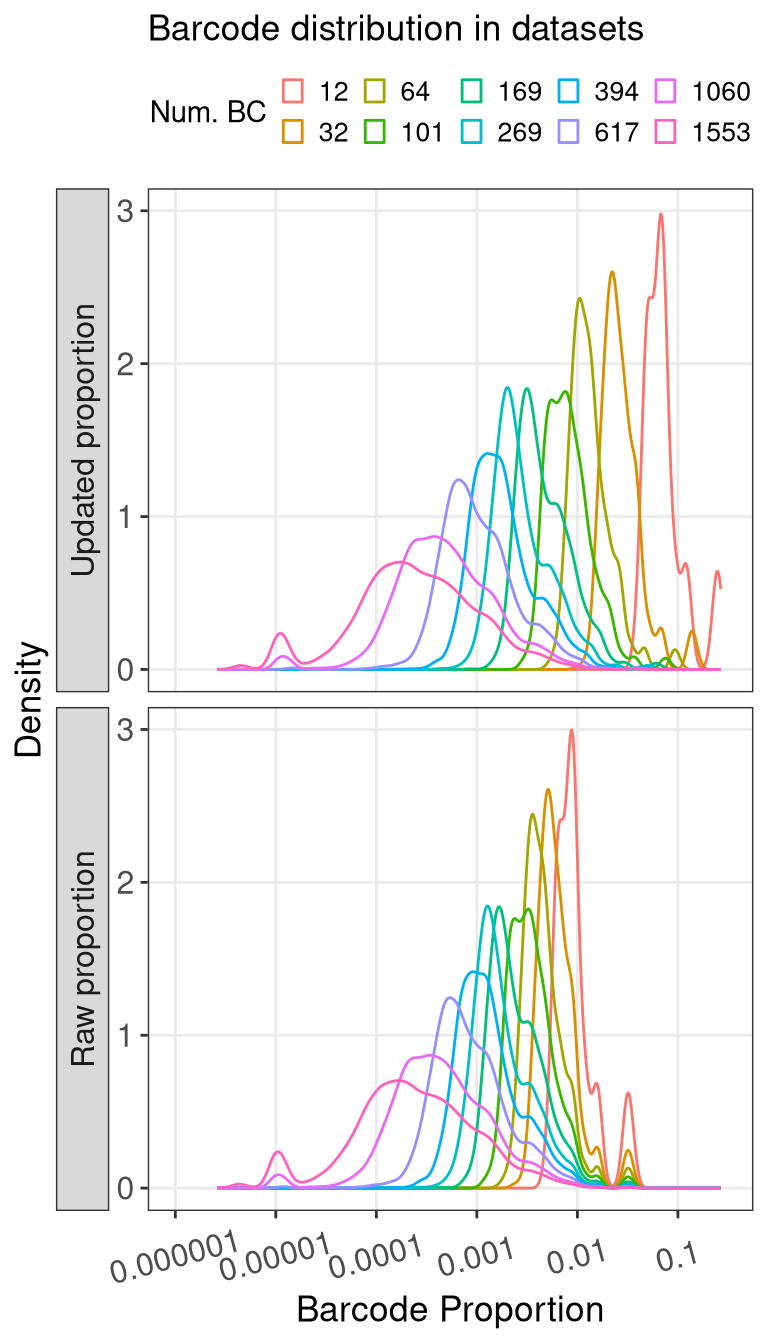
| Version | Author | Date |
|---|---|---|
| 34f5894 | FeiLiyang | 2026-01-01 |
7.5 F3D: Quantile distribution
data.frame(x = (stats_barcodes_all_levels %>% filter(filter_level==0.95))$Amean_prop) %>%
mutate(quantile_bin = ntile(x, 10)) %>%
ggplot(aes(x = as.factor(quantile_bin), y = x, group = quantile_bin)) +
geom_jitter(color = "grey", shape = ".") +
geom_boxplot() +
scale_y_log10(breaks = 10^seq(-5, -1, by = 1), labels = strip_zeros) +
theme_barbieQ() +
labs(x = "Quantiles of barcode mean proportion", y = "Barcode mean proportion",
title = "Barcode proportion distribution", subtitle = " ") +
theme(axis.text.y = element_text(angle = 90, vjust = 0.5, hjust = 0.5)) -> f3d
f3d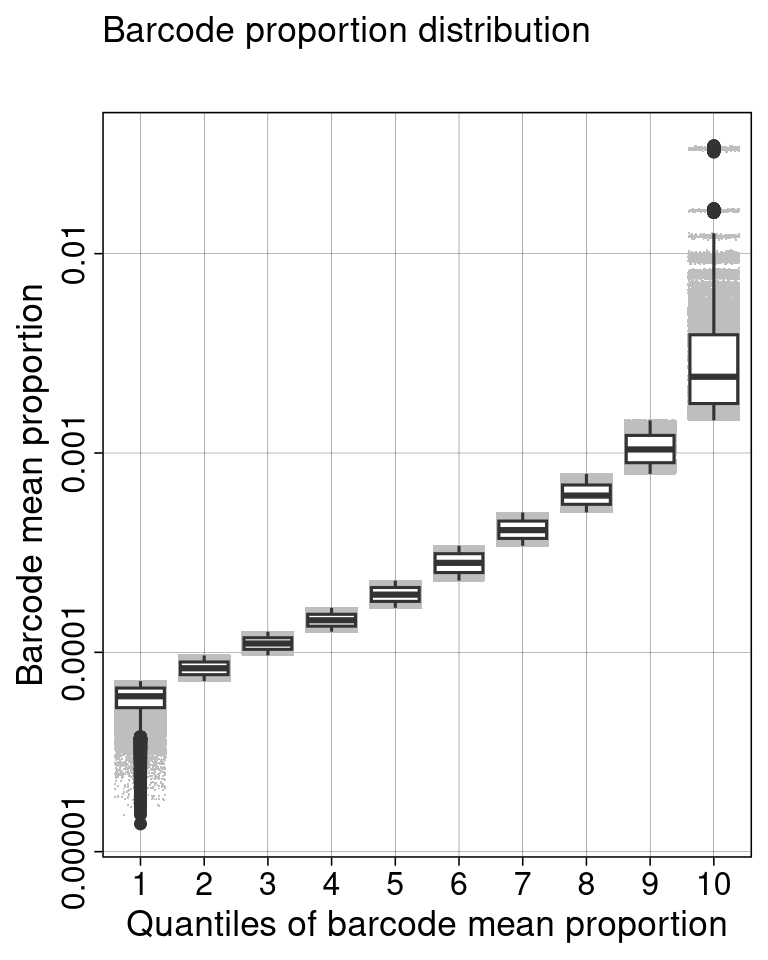
7.6 F3E: False positives in quantiles
- For diff_prop
# plot_fdr_on_quantile(stats_barcodes_all_levels %>% filter(filter_level==0.1), num_quantiles = 4)
# plot_fdr_on_quantile(stats_barcodes_all_levels %>% filter(filter_level==0.2), num_quantiles = 10)
# plot_fdr_on_quantile(stats_barcodes_all_levels %>% filter(filter_level==0.3), num_quantiles = 10)
# plot_fdr_on_quantile(stats_barcodes_all_levels %>% filter(filter_level==0.4), num_quantiles = 10)
# plot_fdr_on_quantile(stats_barcodes_all_levels %>% filter(filter_level==0.5), num_quantiles = 10)
# plot_fdr_on_quantile(stats_barcodes_all_levels %>% filter(filter_level==0.6), num_quantiles = 10)
# plot_fdr_on_quantile(stats_barcodes_all_levels %>% filter(filter_level==0.7), num_quantiles = 10)
# plot_fdr_on_quantile(stats_barcodes_all_levels %>% filter(filter_level==0.8), num_quantiles = 10)
# plot_fdr_on_quantile(stats_barcodes_all_levels %>% filter(filter_level==0.9), num_quantiles = 10)
plot_fdr_on_quantile(stats_barcodes_all_levels %>% filter(filter_level==0.95), num_quantiles = 10) +
theme_barbieQ() +
labs(title = "Type I error rate in quanties") -> f3e8 TPR
8.1 Check premix
- confirming an equal mixture of pool1 and pool2 in baseline samples (the “null” samples).
colnames(mixture_plus1_top) [1] "P1" "P2" "null_660.1" "null_660.2"
[5] "null_330.1" "null_330.2" "null_160.1" "null_160.2"
[9] "null_80.1" "null_80.2" "null_40.1" "null_40.2"
[13] "null_20.1" "null_20.2" "m_null_160.p35.1" "m_null_160.p27.1"
[17] "m_null_160.p18.1" "m_null_160.p35.2" "m_null_160.p27.2" "m_null_160.p18.2"
[21] "m_null_80.p35.1" "m_null_80.p27.1" "m_null_80.p18.1" "m_null_80.p35.2"
[25] "m_null_80.p27.2" "m_null_80.p18.2" "m_null_40.p35.1" "m_null_40.p27.1"
[29] "m_null_40.p18.1" "m_null_40.p35.2" "m_null_40.p27.2" "m_null_40.p18.2"
[33] "m_null_20.p35.1" "m_null_20.p27.1" "m_null_20.p18.1" "m_null_20.p35.2"
[37] "m_null_20.p27.2" "m_null_20.p18.2" prop_raw <- mixture_plus1_top@assays@data$proportion
pool1_vec <- prop_raw[,1]
pool2_vec <- prop_raw[,2]
## if equal mixture of pool1 and pool2; designed prop should be 1/2 of total props in original pool1 and pool2
check_mix_df <- data.frame(
ps = rowMeans(prop_raw[,1:2]),
mixture_660 = prop_raw[,3]
)
ggplot(check_mix_df, aes(x = ps, y = mixture_660)) +
geom_point() +
coord_fixed() +
geom_smooth(method = "lm")`geom_smooth()` using formula = 'y ~ x'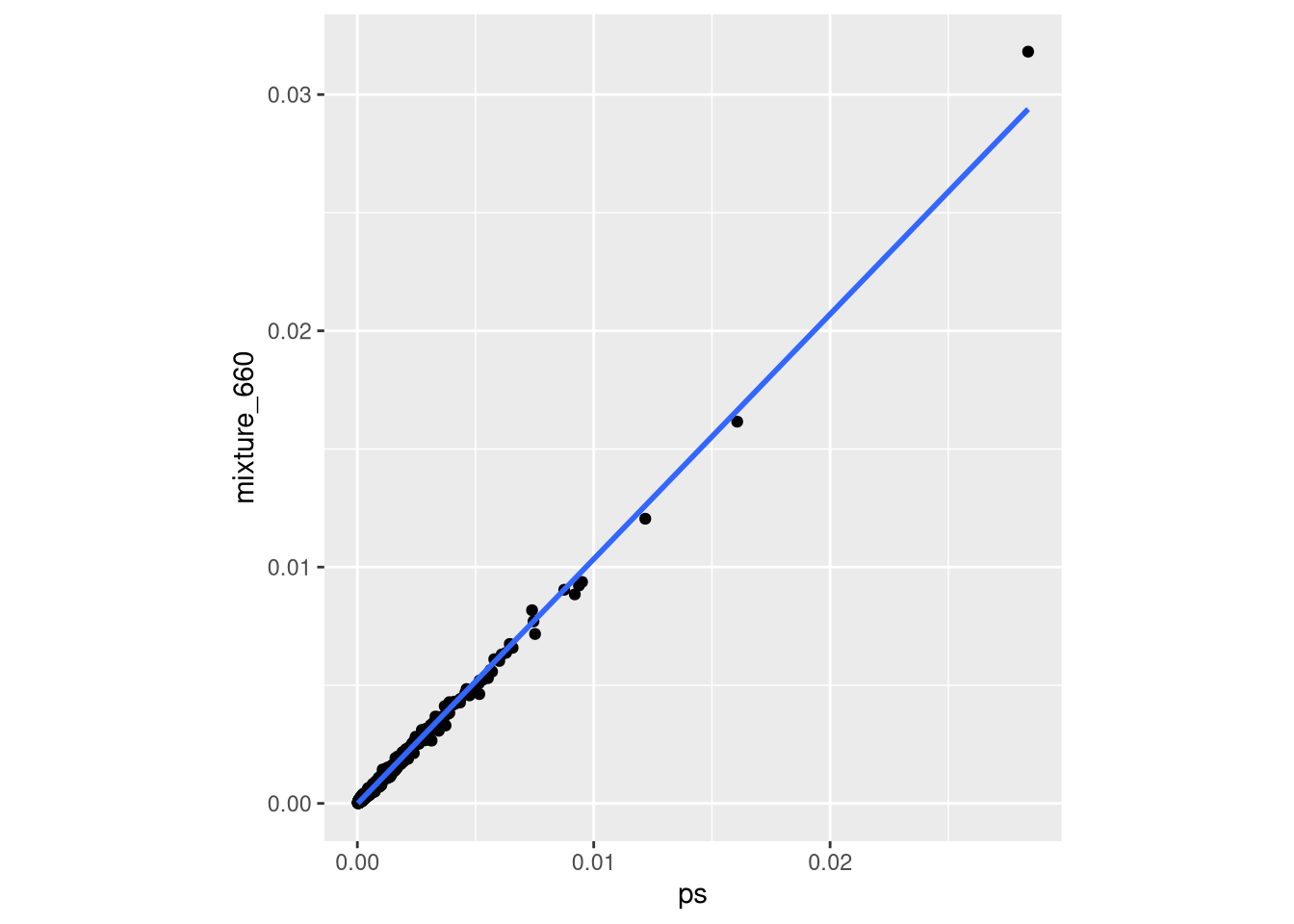
- Confirming orthogonal between premix1 and premix2: barcodes mostly not overlapping between the two premix samples, but a few noise are observed in premix2.
ggplot(data.frame(prop_raw) %>% sqrt() %>% asin()) +
geom_point(aes(x = P1, y = P2))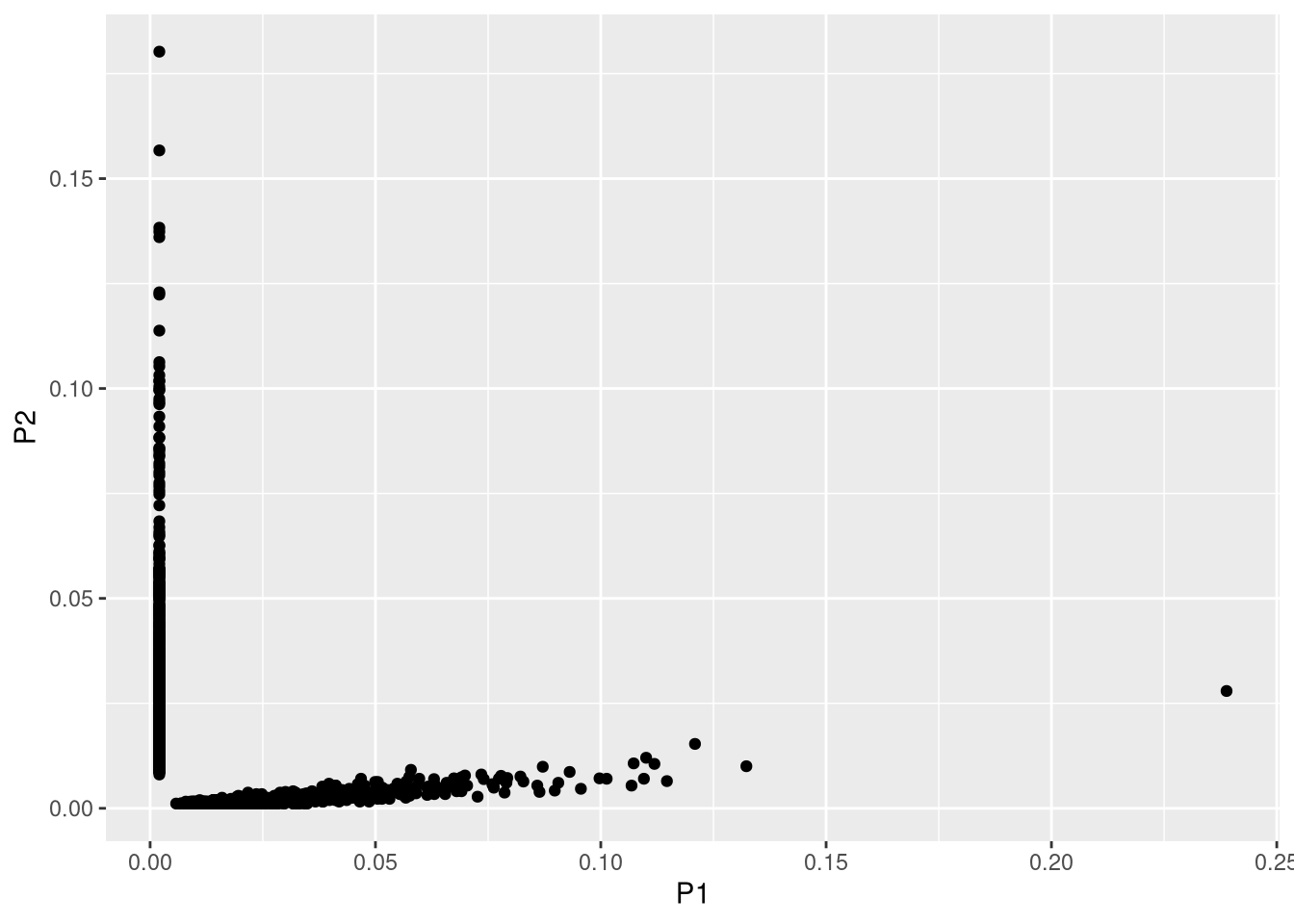
ggplot(data.frame(prop_raw) %>% sqrt() %>% asin(), aes(x = P1+P2, y = abs(P1-P2))) +
geom_point() +
coord_fixed() +
geom_smooth(method = "lm")`geom_smooth()` using formula = 'y ~ x'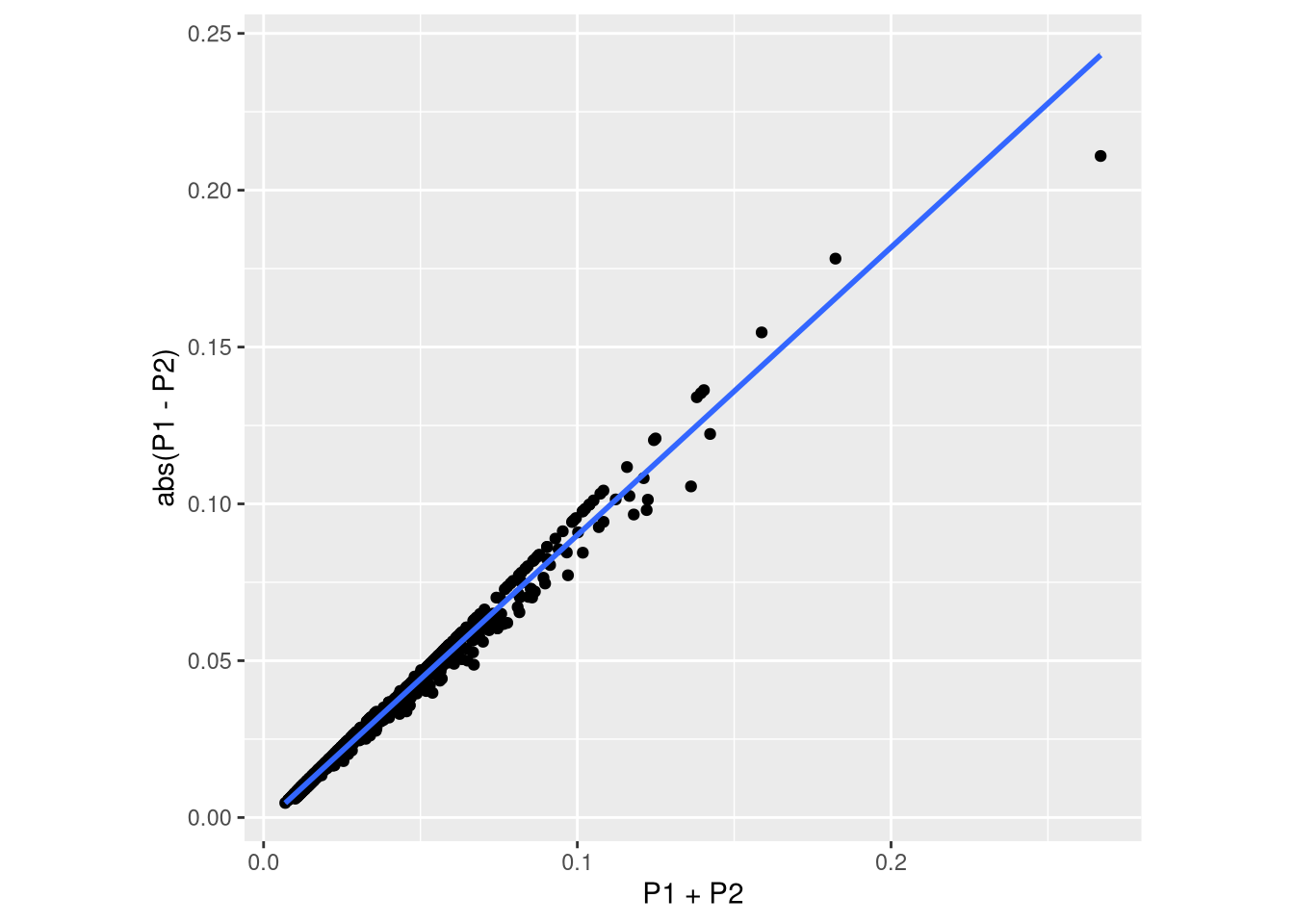
| Version | Author | Date |
|---|---|---|
| 34f5894 | FeiLiyang | 2026-01-01 |
ggplot(data.frame(prop_raw), aes(x = P1+P2, y = abs(P1-P2))) +
geom_point() +
coord_fixed() +
geom_smooth(method = "lm")`geom_smooth()` using formula = 'y ~ x'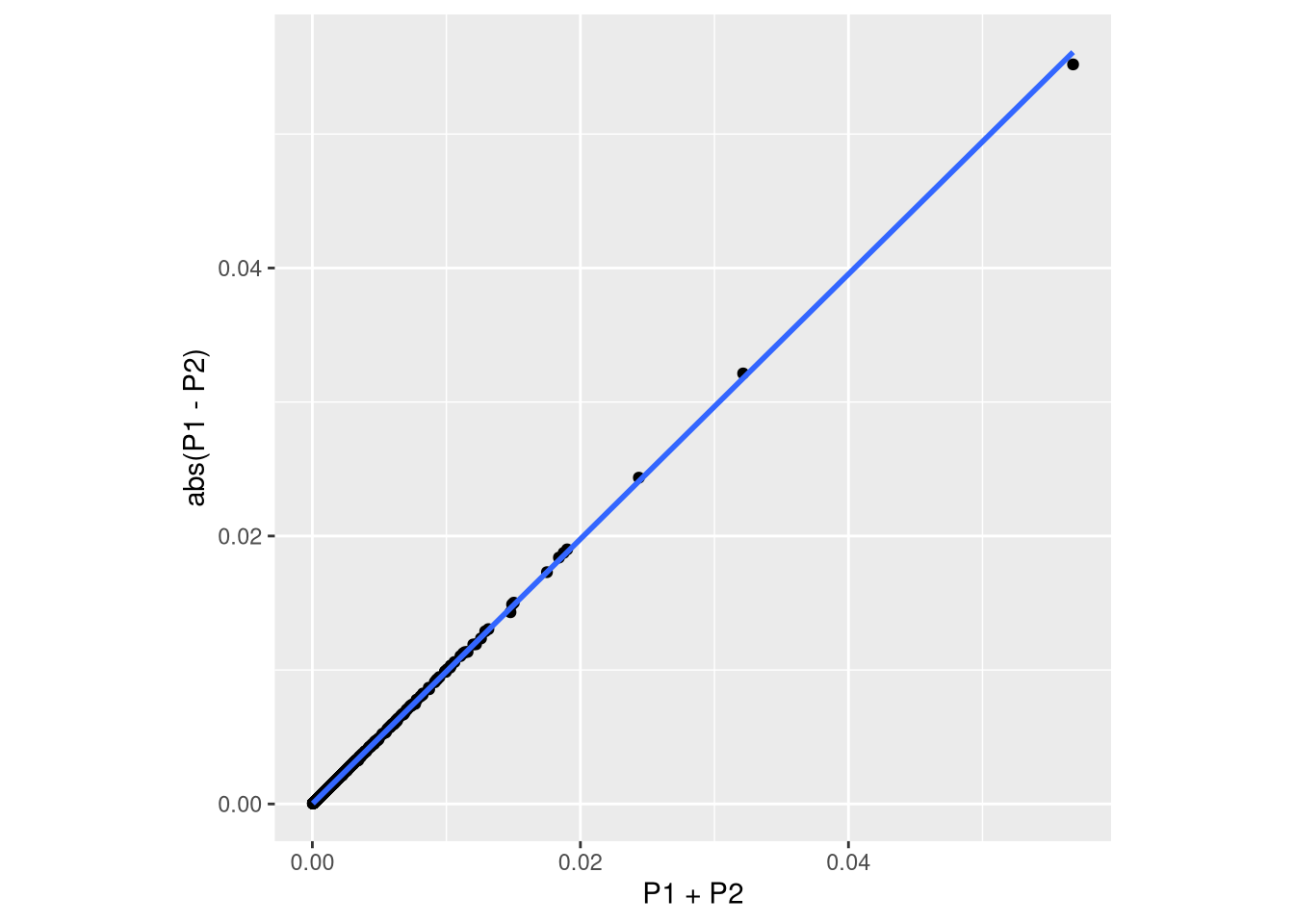
| Version | Author | Date |
|---|---|---|
| 34f5894 | FeiLiyang | 2026-01-01 |
8.2 Designed differences
## theory
a = 0.35
((1/2 + a) / (1+a)) - (1/2)[1] 0.1296296a = 0.27
(1/2 + a) / (1+a) - 1/2[1] 0.1062992a = 0.18
(1/2 + a) / (1+a) - 1/2[1] 0.076271198.3 Model perturbations
mixture_plus1_top$sampleMetadata %>% as.data.frame() %>% summary() Subset BarcodeSize Perturbation TechRep
premix1: 1 Min. : 20000 Min. :0.0000 1:20
premix2: 1 1st Qu.: 40000 1st Qu.:0.0000 2:18
null :12 Median : 80000 Median :0.1800
perturb:24 Mean :121667 Mean :0.1778
3rd Qu.:160000 3rd Qu.:0.2700
Max. :660000 Max. :0.3500
NA's :2 NA's :2 mixed <- mixture_plus1_top[, mixture_plus1_top$sampleMetadata$Subset %in% c("perturb", "null")]
mixed$sampleMetadata %>% as.data.frame() %>% data.table()mixed_target <- mixed$sampleMetadata %>% as.data.frame()
## factorize perturbation levels and barcodeSize levels
mixed_target$Perturbation <- as.factor(mixed_target$Perturbation)
levels(mixed_target$Perturbation)[1] "0" "0.18" "0.27" "0.35"mixed_target$BarcodeSize <- as.factor(mixed_target$BarcodeSize)
levels(mixed_target$BarcodeSize)[1] "20000" "40000" "80000" "160000" "330000" "660000"mixed_design <- mixed_target %>%
with(model.matrix(~0 + Perturbation + BarcodeSize))
myblock <- mixed_target %>% with(paste(BarcodeSize, Perturbation))
table(myblock)myblock
160000 0 160000 0.18 160000 0.27 160000 0.35 20000 0 20000 0.18
2 2 2 2 2 2
20000 0.27 20000 0.35 330000 0 40000 0 40000 0.18 40000 0.27
2 2 2 2 2 2
40000 0.35 660000 0 80000 0 80000 0.18 80000 0.27 80000 0.35
2 2 2 2 2 2 8.4 Apply tests: extract estimates
myblock <- NULL
es_de_none <- lapply(c(0.18, 0.27, 0.35), function(i) apply_model(mixed, mixed_design, myblock, a = i))setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.es_de_none_df <- do.call(rbind, es_de_none)
es_de_none_df <- es_de_none_df %>%
mutate(Signif. = ifelse(P.Value<0.05, "signif.", "n.s.")) %>%
mutate(Signif. = factor(Signif., levels = c("signif.", "n.s.")))
es_de_asin <- lapply(c(0.18, 0.27, 0.35), function(i) apply_model(mixed, mixed_design, myblock, a = i, trans = "asin-sqrt"))setting Perturbation as the primary factor in `sampleMetadata`.
no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.es_de_asin_df <- do.call(rbind, es_de_asin)
es_de_asin_df <- es_de_asin_df %>%
mutate(Signif. = ifelse(P.Value<0.05, "signif.", "n.s.")) %>%
mutate(Signif. = factor(Signif., levels = c("signif.", "n.s.")))
es_de_logit <- lapply(c(0.18, 0.27, 0.35), function(i) apply_model(mixed, mixed_design, myblock, a = i, trans = "logit"))setting Perturbation as the primary factor in `sampleMetadata`.
no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.es_de_logit_df <- do.call(rbind, es_de_logit)
es_de_logit_df <- es_de_logit_df %>%
mutate(Signif. = ifelse(P.Value<0.05, "signif.", "n.s.")) %>%
mutate(Signif. = factor(Signif., levels = c("signif.", "n.s.")))
es_de_df <- rbind(es_de_none_df, es_de_asin_df, es_de_logit_df)8.5 FS3A: Validate estimates
regression_stats_none <- es_de_none_df %>%
group_by(Contrast) %>%
summarise(
cor = cor(diff, meanDiff),
slope = coef(lm(meanDiff ~ I(diff)))[2],
interc = coef(lm(meanDiff ~ I(diff)))[1],
# res = resid(mod <- lm(meanDiff ~ I(diff))),
# meanDiff = meanDiff,
.groups = "drop"
)
regression_stats_none$Method = "none"
regression_stats_none$x = -0.01
regression_stats_none$y = 0.01
regression_stats_asin <- es_de_asin_df %>%
group_by(Contrast) %>%
summarise(
cor = cor(diff, meanDiff),
slope = coef(lm((meanDiff) ~ I(diff)))[2],
interc = coef(lm((meanDiff) ~ I(diff)))[1],
.groups = "drop"
)
regression_stats_asin$Method = "asin-sqrt"
regression_stats_asin$x = -0.001
regression_stats_asin$y = 0.001
regression_stats_logit <- es_de_logit_df %>%
group_by(Contrast) %>%
summarise(
cor = cor(diff, (meanDiff)),
slope = coef(lm((meanDiff) ~ I(diff)))[2],
interc = coef(lm((meanDiff) ~ I(diff)))[1],
.groups = "drop"
)
regression_stats_logit$Method = "logit"
regression_stats_logit$x = -0.05
regression_stats_logit$y = 1
regression_stats <- rbind(regression_stats_asin, regression_stats_none, regression_stats_logit)(facet_grid doesn’t allow free x sclae for the same column. so plot three methods seperately.)
p_es_de_asin <- ggplot(es_de_asin_df, aes(x = diff, y = (meanDiff))) +
geom_point(aes(color = Signif.), alpha = 0.5) +
# stat_smooth(method = "lm", linetype = "dashed", colour = "black") +
geom_abline(data = regression_stats_none, aes(intercept = interc, slope = slope), linetype = "dashed") +
theme_bw() +
# coord_fixed() +
facet_wrap(~paste0("Contrast: ", Contrast)) +
geom_text(data = regression_stats_asin, x = -0.01, y = 0.01,
aes(label = paste0("r = ", round(cor, digits = 2), "\n", "slope = ", round(slope, digits = 2)))) +
theme(axis.title.x = element_blank(), legend.position = "none") +
labs(color = "Barcode", y = " \n \n (asin-sqrt)")
p_es_de_none <- ggplot(es_de_none_df, aes(x = diff, y = (meanDiff))) +
geom_point(aes(color = Signif.), alpha = 0.5) +
# stat_smooth(method = "lm", linetype = "dashed", colour = "black") +
geom_abline(data = regression_stats_none, aes(intercept = interc, slope = slope), linetype = "dashed") +
theme_linedraw()+
# coord_fixed() +
facet_wrap(~paste0("Contrast: ", Contrast)) +
geom_text(data = regression_stats_none, x = -0.0015, y = 0.001,
aes(label = paste0("r = ", round(cor, digits = 2), "\n", "slope = ", round(slope, digits = 2)))) +
labs(y = " \n \n (no transformation)", x = " \n Designed differences")+
## zoom in, removing a few barcodes with large proportion.
scale_x_continuous(limits = c(-0.003, 0.003)) +
scale_y_continuous(limits = c(-0.003, 0.003)) +
theme(legend.position = "none", strip.text = element_blank()) + # remove facet titles
labs(color = "Barcode")
p_es_de_logit <- ggplot(es_de_logit_df, aes(x = diff, y = (meanDiff))) +
geom_point(aes(color = Signif.), alpha = 0.5) +
# stat_smooth(method = "lm", linetype = "dashed", colour = "black") +
geom_abline(data = regression_stats_none, aes(intercept = interc, slope = slope), linetype = "dashed") +
theme_linedraw()+
facet_wrap(~paste0("Contrast: ", Contrast)) +
geom_text(data = regression_stats_logit, x = -0.15, y = 1,
aes(label = paste0("r = ", round(cor, digits = 2), "\n", "slope = ", round(slope, digits = 2)))) +
labs(y = "Estimated differences \n \n (logit)", color = "Barcode") +
## zoom in, removing one barcode with large proportion.
# scale_x_continuous(limits = c(-0.3, 0.3)) +
# scale_y_continuous(limits = c(-0.3, 0.3)) +
theme(axis.title.x = element_blank(), legend.position = "right", strip.text = element_blank()) # remove facet titlesfs3a <- (p_es_de_asin + theme(legend.title = element_text(size = 12), legend.text = element_text(size = 11),
axis.title = element_text(size = 13), axis.text = element_text(size = 12)) ) /
(p_es_de_logit + theme(legend.title = element_text(size = 12), legend.text = element_text(size = 11),
axis.title = element_text(size = 13), axis.text = element_text(size = 12))) /
(p_es_de_none + theme(legend.title = element_text(size = 12), legend.text = element_text(size = 11),
axis.title = element_text(size = 13), axis.text = element_text(size = 12)))
fs3aWarning: Removed 7 rows containing missing values or values outside the scale range
(`geom_point()`).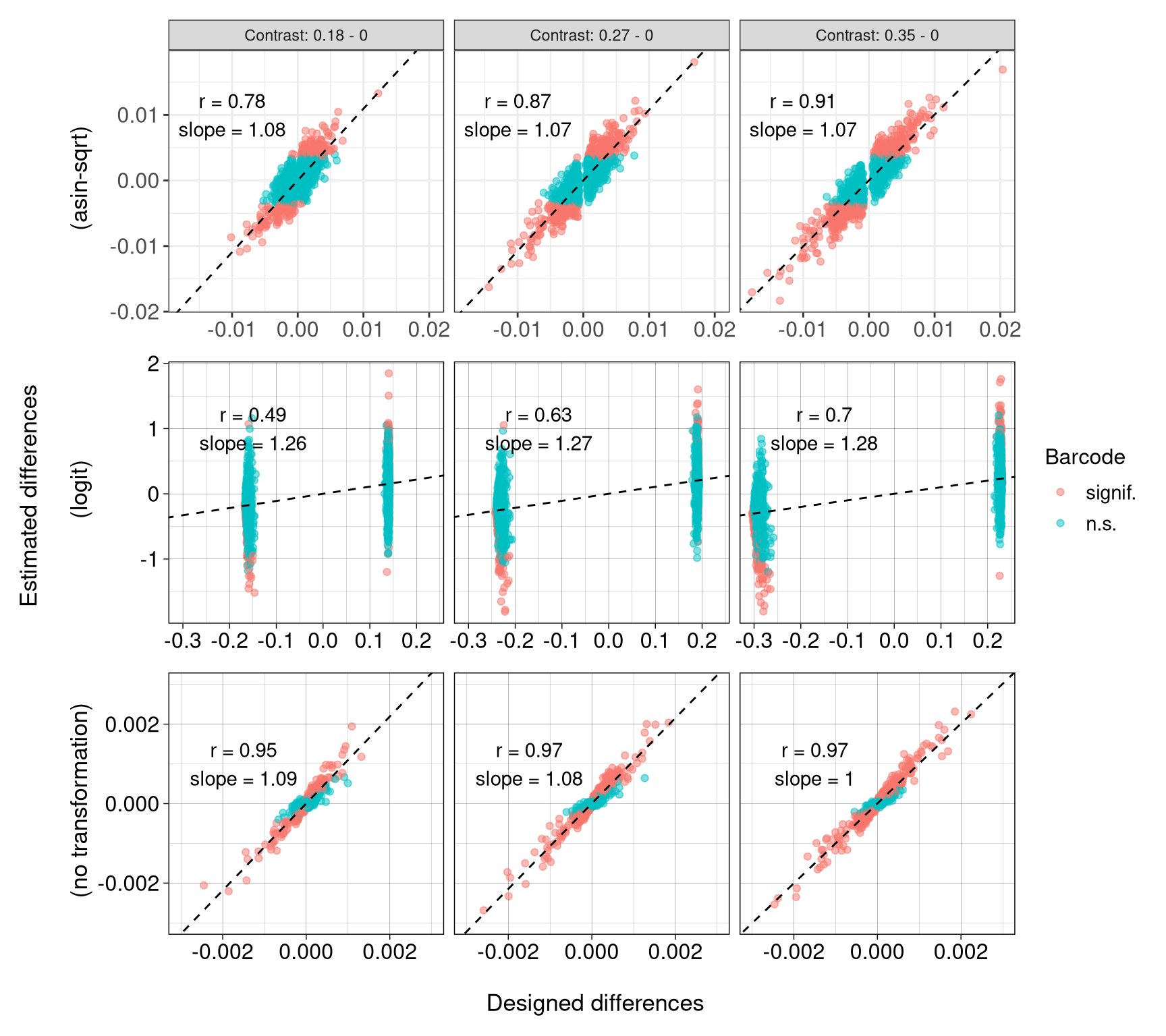
8.6 F3I: TPR as a function to N_BC
8.6.1 Updated proportion
test_perturb <- function(mixed, mixed_target, mixed_design, myblock, trans = "none", a = 0.18) {
tester <- testBarcodeSignif(
barbieQ = mixed, sampleMetadata = mixed_target, sampleGroup = "Perturbation",
contrastFormula = paste0("Perturbation", a, " - Perturbation0"), transformation = trans,
designMatrix = mixed_design, block = myblock)
veri_diff <- as.data.frame(tester@elementMetadata$testingBarcode)
veri_diff <- rownames_to_column(veri_diff, var = "BC")
veri_diff$Contrast = paste0(a, " - 0")
veri_diff$Method = trans
return(veri_diff)
}test_all_levels <- lapply(
c(1:9/10, 0.95),
function(i) {get_mixted_by_filter(barbieQ = mixture_plus1, filter_level = i)}
)setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.test_all_levels <- do.call(rbind, test_all_levels)
test_TPR <- test_all_levels %>%
group_by(Method, Contrast, N_BC) %>%
summarize(Frac_signif = sum(P.Value < 0.05) / n(), .groups = "drop")ggplot(test_TPR, aes(x = as.factor(N_BC), y = Frac_signif, color = Method, linetype = Contrast, shape = Contrast)) +
# geom_boxplot(alpha = 0.6, position = position_dodge(), outliers = T) +
geom_line(aes(group = paste0(Contrast, Method))) +
geom_point() +
labs(
x = paste0("Number of total barcodes after filtering"),
y = "Fraction of P-Value < 0.05",
title = "True postive rate as a function to total barcode number"
) +
geom_hline(yintercept = 0.05, linetype = "dashed") +
theme_linedraw()+
theme(axis.text.x = element_text(angle = 0, vjust = 0.5, size = 10, hjust = 0.5), aspect.ratio = 0.5) +
theme(
legend.title = element_text(size = 12), legend.text = element_text(size = 11),
axis.title = element_text(size = 13), axis.text = element_text(size = 12))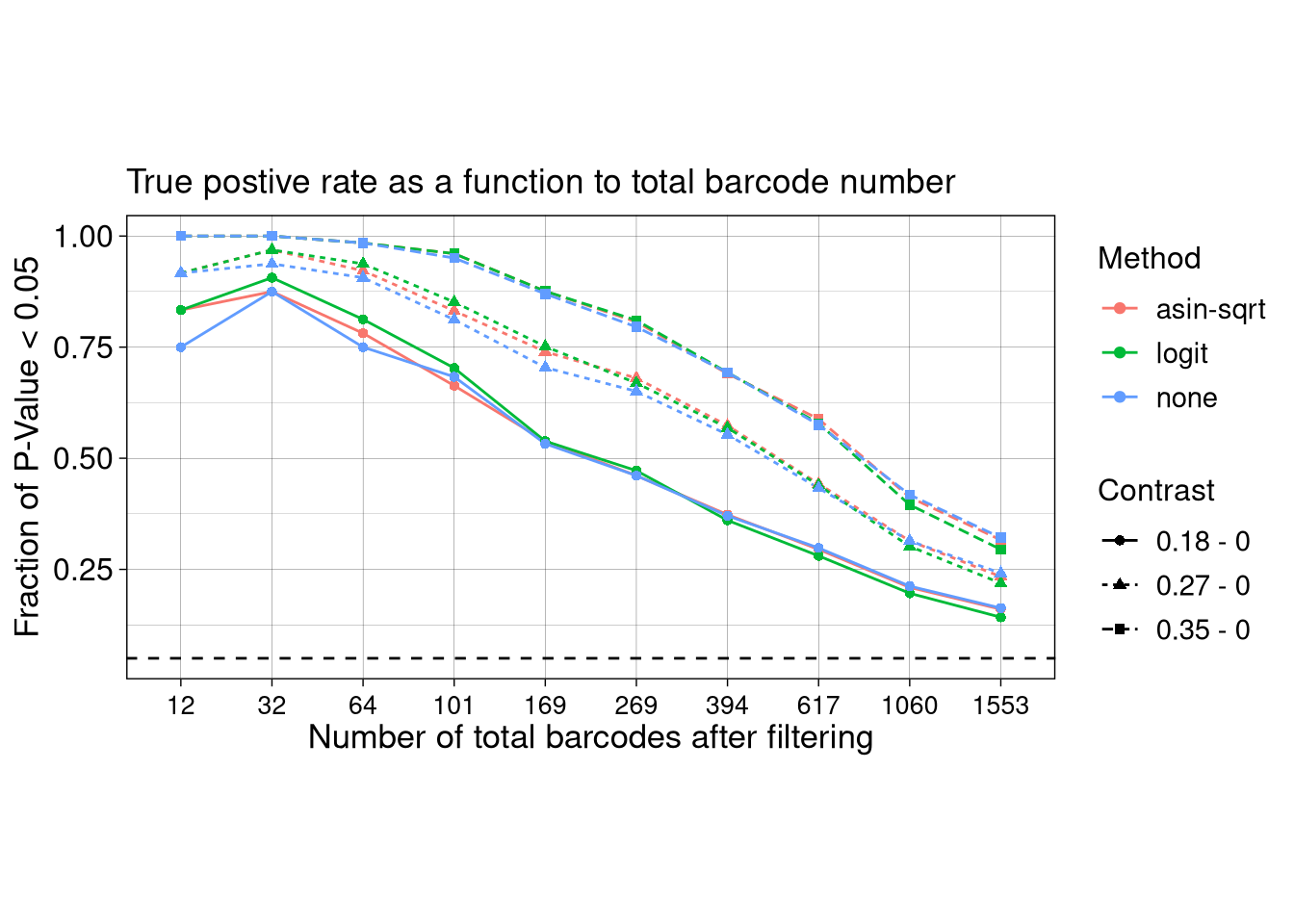
| Version | Author | Date |
|---|---|---|
| 34f5894 | FeiLiyang | 2026-01-01 |
8.6.2 Raw proportion
## test on raw proportion
test_0.95_raw <- get_mixted_by_filter(barbieQ = mixture_plus1, filter_level = 0.95, update_prop = FALSE)setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.setting Perturbation as the primary factor in `sampleMetadata`.no block specified, so there are no duplicate measurements.test_all_levels_raw <- cbind(
test_0.95_raw,
mean_prop,
BC_set_by_filter[
rep(
seq_len(nrow(BC_set_by_filter)),
length.out = nrow(test_0.95_raw)
),
]
)Warning in data.frame(..., check.names = FALSE): row names were found from a
short variable and have been discardedtest_TPR_raw <- test_all_levels_raw %>%
select(-N_BC) %>%
pivot_longer(
cols = as.character(filter_to_N[c(1:9/10, 0.95) %>% as.character()]),
names_to = "N_BC",
values_to = "selectedBy"
) %>%
group_by(Method, Contrast, N_BC) %>%
summarize(Frac_signif = sum(P.Value < 0.05 & selectedBy) / sum(selectedBy), .groups = "drop")
test_TPR_raw$N_BC <- as.numeric(test_TPR_raw$N_BC)
ggplot(test_TPR_raw, aes(x = as.factor(N_BC), y = Frac_signif, color = Method, group = paste0(Method, Contrast))) +
# geom_boxplot(alpha = 0.6, position = position_dodge(), outliers = T) +
geom_point(aes(shape = Contrast)) +
geom_line(aes(linetype = Contrast)) +
labs(
x = paste0("Number of total barcodes after filtering"),
y = "Fraction of P-Value < 0.05"
) +
geom_hline(yintercept = 0.05, linetype = "dashed") +
# facet_wrap(~Contrast, ncol =3) +
theme_linedraw()+
theme(axis.text.x = element_text(angle = 0, vjust = 0.5, size = 10, hjust = 0.5), aspect.ratio = 0.5) +
theme(
legend.title = element_text(size = 12), legend.text = element_text(size = 11),
axis.title = element_text(size = 13), axis.text = element_text(size = 12))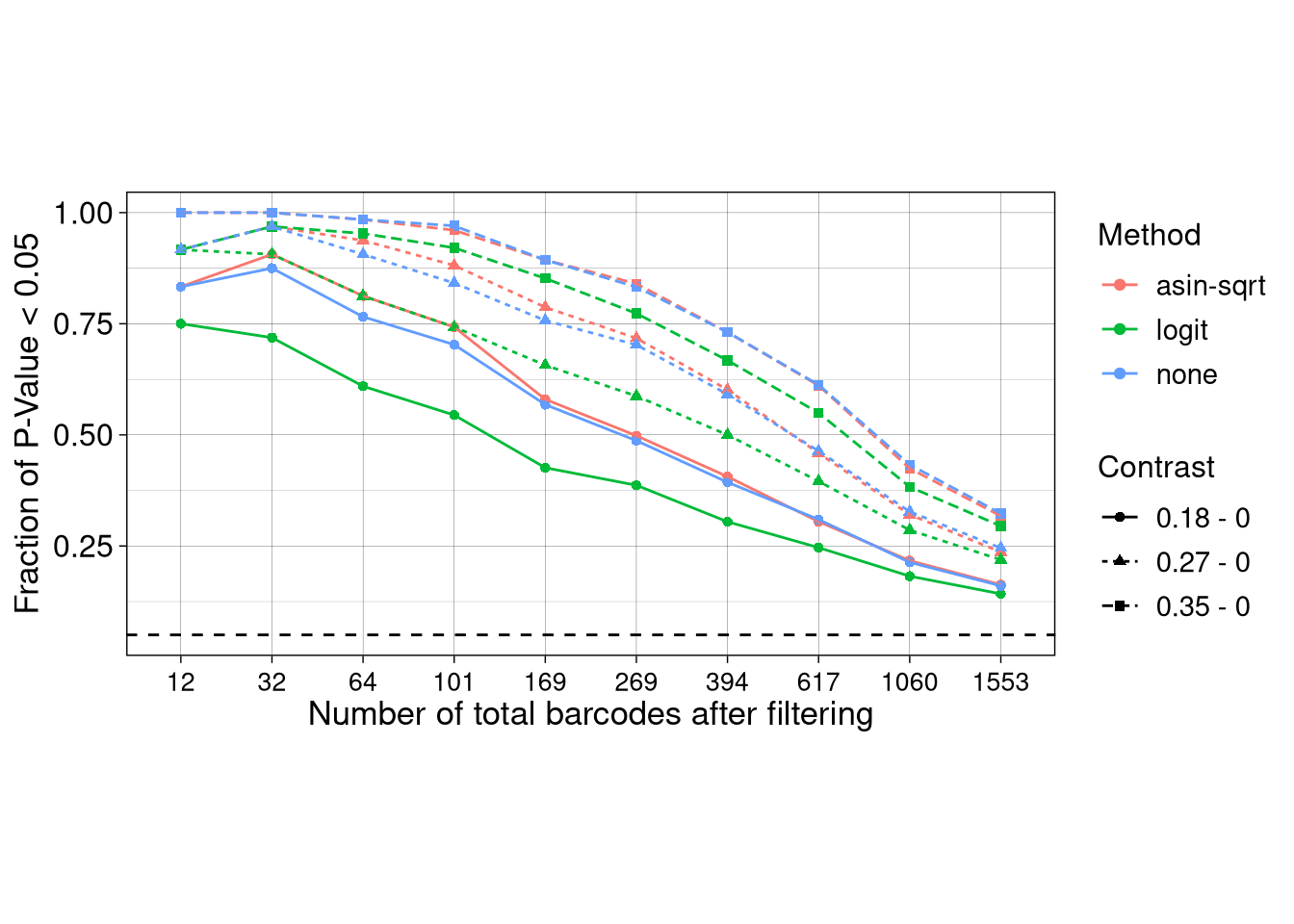
rbind(
data.frame(test_TPR_raw, prop_type = "Raw proportion"),
data.frame(test_TPR, prop_type = "Updated proportion")
) %>%
mutate(prop_type = factor(prop_type, levels = c("Updated proportion", "Raw proportion"))) %>%
## plot
ggplot(aes(x = as.factor(N_BC), y = Frac_signif, color = Method, group = paste0(Method, Contrast))) +
# geom_boxplot(alpha = 0.6, position = position_dodge(), outliers = T) +
geom_point(aes(shape = Contrast)) +
geom_line(aes(linetype = Contrast)) +
labs(
x = paste0("Number of total barcodes after filtering"),
y = "Fraction of P-Value < 0.05", color = "Method", shape = "Contrast", linetype = "Contrast",
title = "True positive rate to number of barcodes"
) +
geom_hline(yintercept = 0.05, linetype = "dashed") +
facet_grid(prop_type~1, switch = "y") +
theme_bw()+
theme_barbieQ_size() +
theme(panel.grid.minor = element_blank(), axis.text.x = element_text(angle = 15, vjust = 0.5, hjust = 0.5),
legend.position = "top",
strip.placement = "outside", strip.text.y = element_text(size = 12),
strip.text.x = element_blank(), strip.background.x = element_blank()) + guides(color = "none") -> f3i
f3i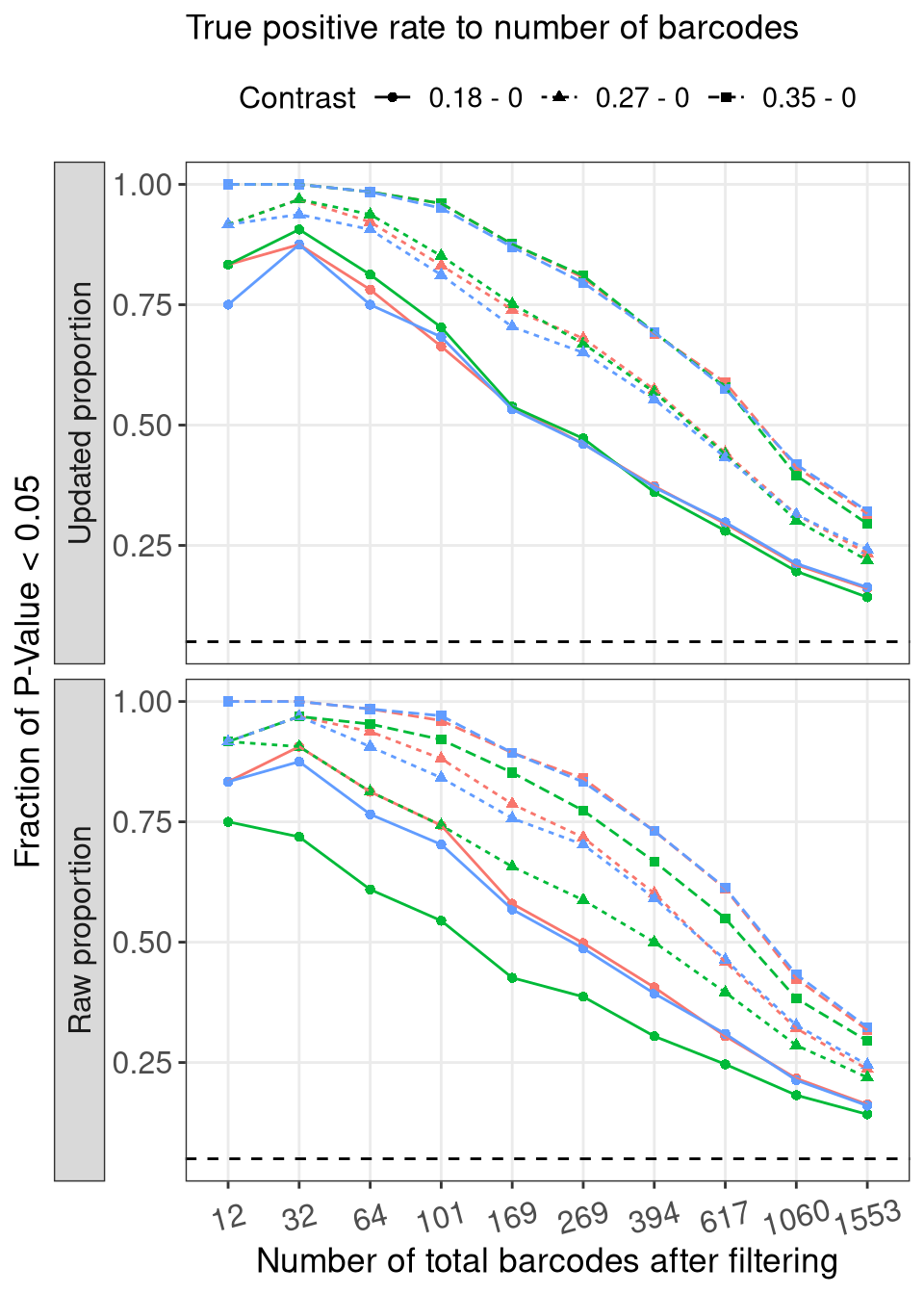
8.7 F3F: TPR in quantiles
# plot_tpr_on_quantile(test_all_levels %>% filter(filter_level==0.1), num_quantiles = 4)
# plot_tpr_on_quantile(test_all_levels %>% filter(filter_level==0.2), num_quantiles = 10)
# plot_tpr_on_quantile(test_all_levels %>% filter(filter_level==0.3), num_quantiles = 10)
# plot_tpr_on_quantile(test_all_levels %>% filter(filter_level==0.4), num_quantiles = 10)
# plot_tpr_on_quantile(test_all_levels %>% filter(filter_level==0.5), num_quantiles = 10)
# plot_tpr_on_quantile(test_all_levels %>% filter(filter_level==0.6), num_quantiles = 10)
# plot_tpr_on_quantile(test_all_levels %>% filter(filter_level==0.7), num_quantiles = 10)
# plot_tpr_on_quantile(test_all_levels %>% filter(filter_level==0.8), num_quantiles = 10)
# plot_tpr_on_quantile(test_all_levels %>% filter(filter_level==0.9), num_quantiles = 10)
plot_tpr_on_quantile(test_all_levels %>% filter(filter_level==0.95), num_quantiles = 10) +
theme_barbieQ() +
labs(title = "True positive rate in quanties") -> f3f8.8 FS3B: Residules
resi_none <- lapply(c(0.18, 0.27, 0.35), function(i) get_residules(mixed, mixed_design, myblock, a = i, trans = "none"))
resi_asin <- lapply(c(0.18, 0.27, 0.35), function(i) get_residules(mixed, mixed_design, myblock, a = i, trans = "asin-sqrt"))
resi_logit <- lapply(c(0.18, 0.27, 0.35), function(i) get_residules(mixed, mixed_design, myblock, a = i, trans = "logit"))
resi_df <- rbind(do.call(rbind, resi_none), do.call(rbind, resi_asin), do.call(rbind, resi_logit))- Distribution and QQ plot.
p_resid_distr_35 <- ggplot(resi_df %>% filter(Contrast == "0.35 - 0"),
aes(x = Residual, color = Method)) +
geom_density() +
labs(x = "Residual", y = "Density") +
facet_wrap(~Method, scales = "free", ncol = 1, strip.position = "left") +
theme_bw() +
theme(legend.position = "none", plot.title.position = "panel")
p_resid_qq_35 <- ggplot(resi_df %>% filter(Contrast == "0.35 - 0"),
aes(sample = Residual, color = Method)) +
stat_qq() +
stat_qq_line(color = "black", linetype = "dashed") +
labs(x = "Normal Quantiles", y = "\n Residual Quuantiles") +
facet_wrap(~Method, scales = "free", ncol = 1) +
theme_bw() +
theme(legend.position = "none", strip.text = element_blank(), plot.title.position = "panel")fs3b <- (p_resid_distr_35 +
theme(legend.title = element_text(size = 12), legend.text = element_text(size = 11), strip.text = element_text(size = 12),
axis.title = element_text(size = 13), axis.text = element_text(size = 10))
) +
(p_resid_qq_35 +
theme(legend.title = element_text(size = 12), legend.text = element_text(size = 11),
axis.title = element_text(size = 13), axis.text = element_text(size = 10)))
fs3b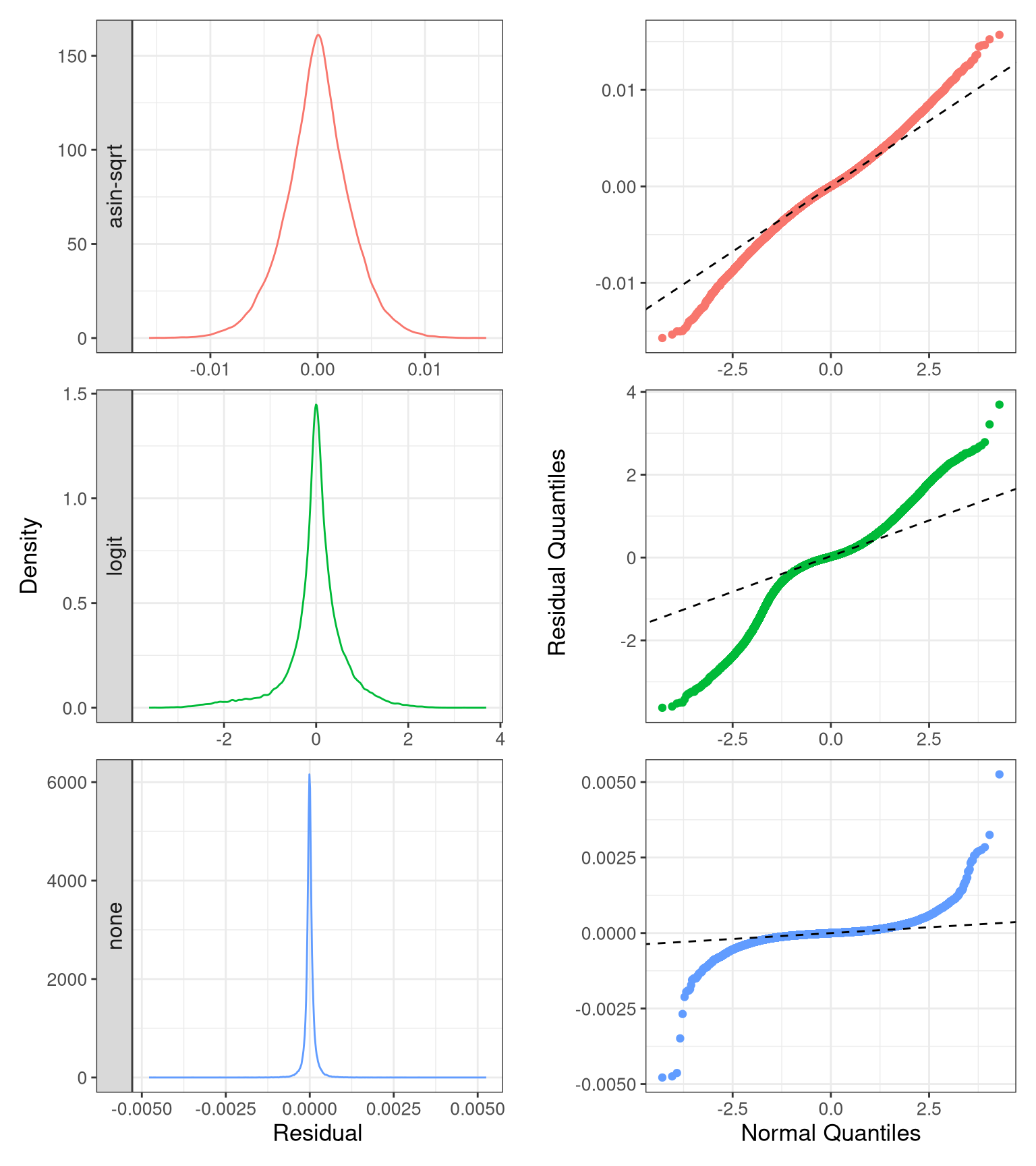
| Version | Author | Date |
|---|---|---|
| 34f5894 | FeiLiyang | 2026-01-01 |
p_resid_distr_27 <- ggplot(resi_df %>% filter(Contrast == "0.27 - 0"), aes(x = Residual, color = Method)) +
geom_density() +
labs(x = "Residual", y = "Density") +
facet_wrap(~Method, scales = "free", ncol = 1) +
theme_bw() +
theme(legend.position = "none", strip.text = element_blank())
p_resid_qq_27 <- ggplot(resi_df %>% filter(Contrast == "0.27 - 0"), aes(sample = Residual, color = Method)) +
stat_qq() +
stat_qq_line(color = "black", linetype = "dashed") +
labs(x = "Normal Quantiles", y = "Residule Quantiles") +
facet_wrap(~Method, scales = "free", ncol = 1) +
theme_bw() +
theme(legend.position = "none", strip.text = element_blank())
p_resid_distr_18 <- ggplot(resi_df %>% filter(Contrast == "0.18 - 0"), aes(x = Residual, color = Method)) +
geom_density() +
labs(x = "Residual", y = "Density") +
facet_wrap(~Method, scales = "free", ncol = 1) +
theme_bw() +
theme(legend.position = "none", strip.text = element_blank())
p_resid_qq_18 <- ggplot(resi_df %>% filter(Contrast == "0.18 - 0"), aes(sample = Residual, color = Method)) +
stat_qq() +
stat_qq_line(color = "black", linetype = "dashed") +
labs(x = "Normal Quantiles", y = "Residule Quantiles") +
facet_wrap(~Method, scales = "free", ncol = 1) +
theme_bw() +
theme(legend.position = "none", strip.text = element_blank())p_resid_distr_35 + p_resid_distr_27 + p_resid_distr_18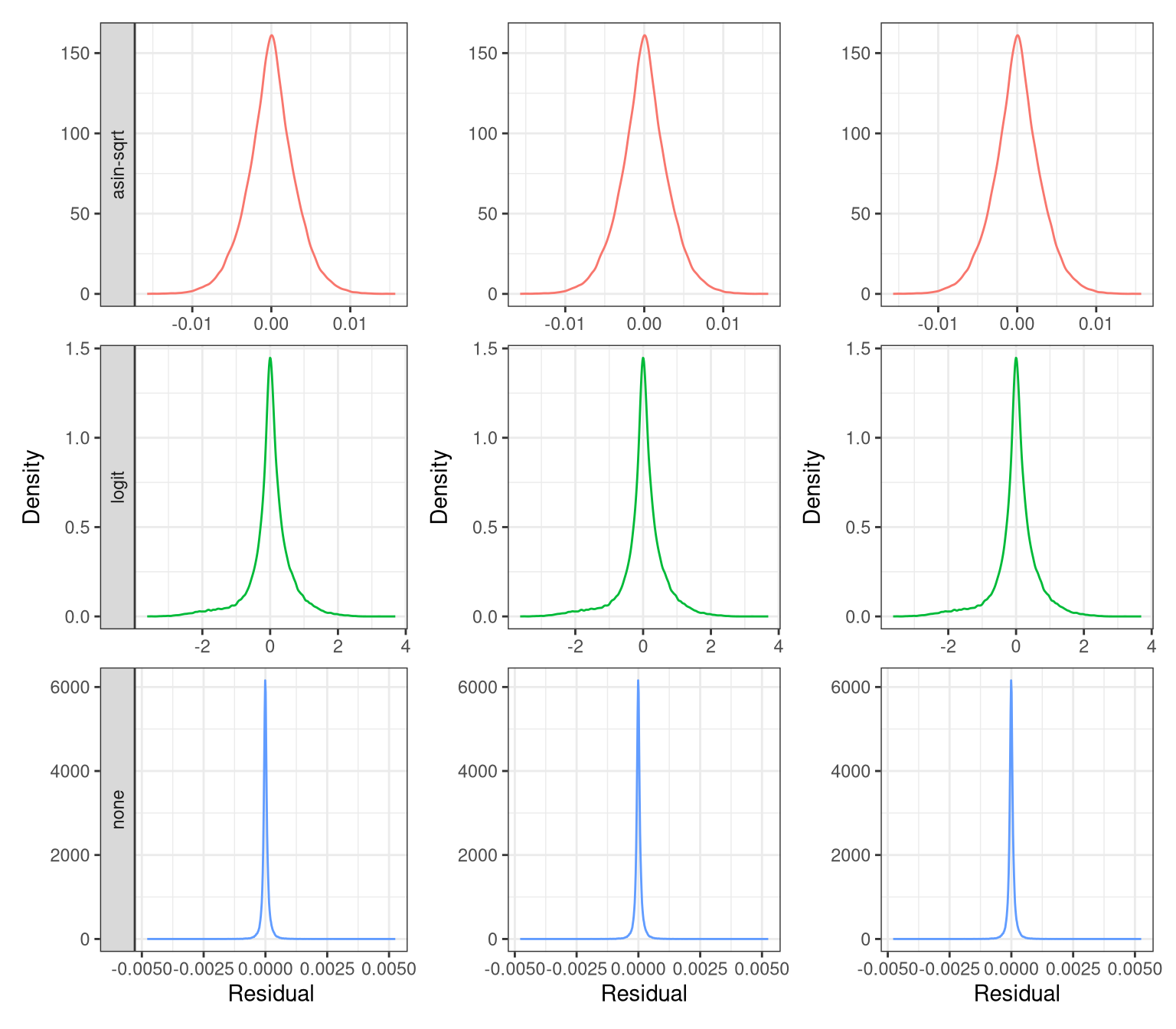
p_resid_qq_35 + p_resid_qq_27 + p_resid_qq_18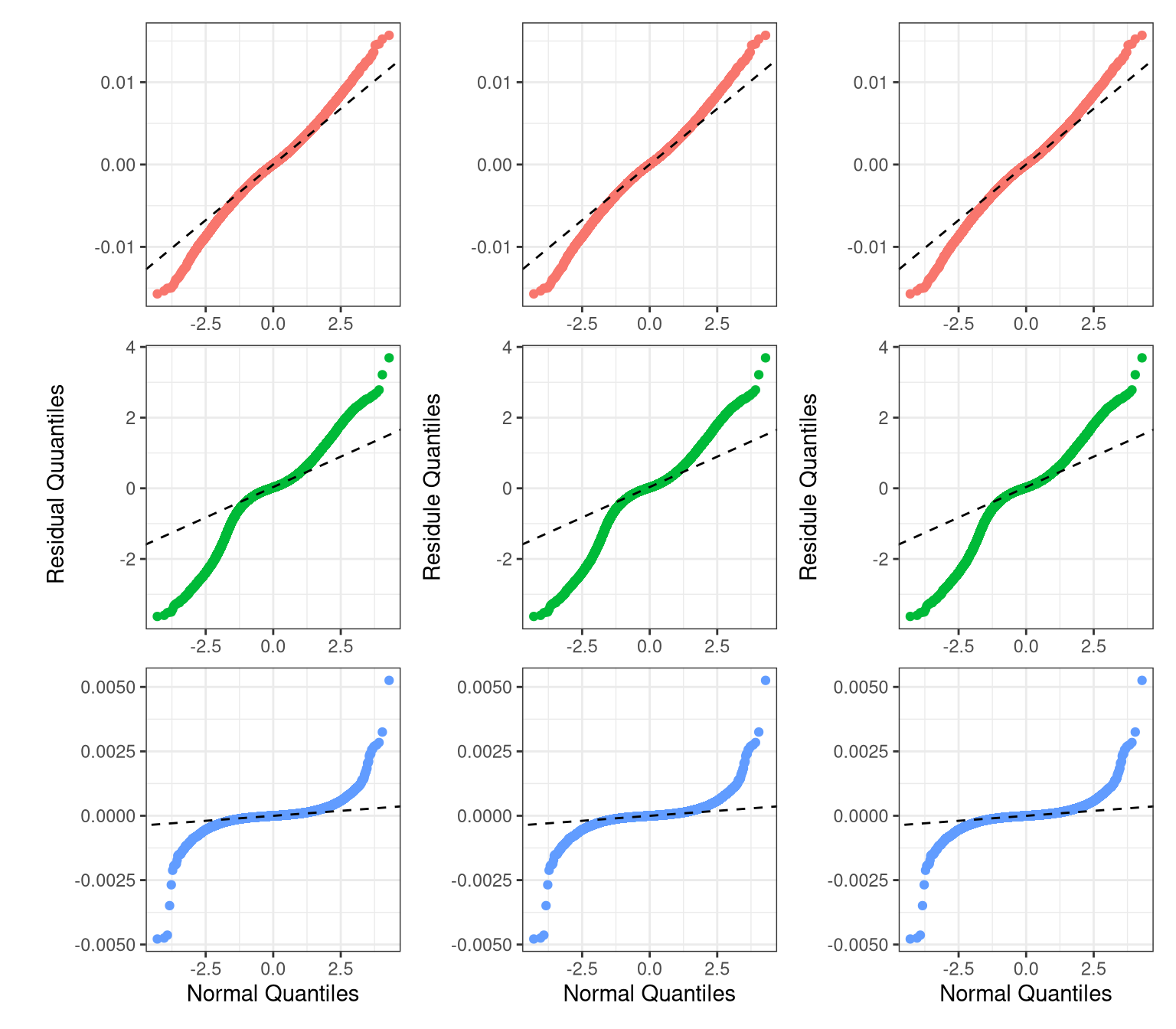
| Version | Author | Date |
|---|---|---|
| 34f5894 | FeiLiyang | 2026-01-01 |
8.9 Consistency
asin_adjP_3con <- es_de_asin_df %>%
dplyr::select(BC, Contrast, P.Value) %>%
tidyr::pivot_wider(
id_cols = BC,
names_from = Contrast,
values_from = P.Value
)
none_adjP_3con <- es_de_none_df %>%
dplyr::select(BC, Contrast, P.Value) %>%
tidyr::pivot_wider(
id_cols = BC,
names_from = Contrast,
values_from = P.Value
)eu <- eulerr::euler(list(
asin_C0.18 = rownames(asin_adjP_3con)[asin_adjP_3con$`0.18 - 0` < 0.05],
asin_C0.27 = rownames(asin_adjP_3con)[asin_adjP_3con$`0.27 - 0` < 0.05],
asin_C0.35 = rownames(asin_adjP_3con)[asin_adjP_3con$`0.35 - 0` < 0.05],
none_C0.18 = rownames(none_adjP_3con)[none_adjP_3con$`0.18 - 0` < 0.05],
none_C0.27 = rownames(none_adjP_3con)[none_adjP_3con$`0.27 - 0` < 0.05],
none_C0.35 = rownames(none_adjP_3con)[none_adjP_3con$`0.35 - 0` < 0.05]
))
plot(eu, quantities = TRUE)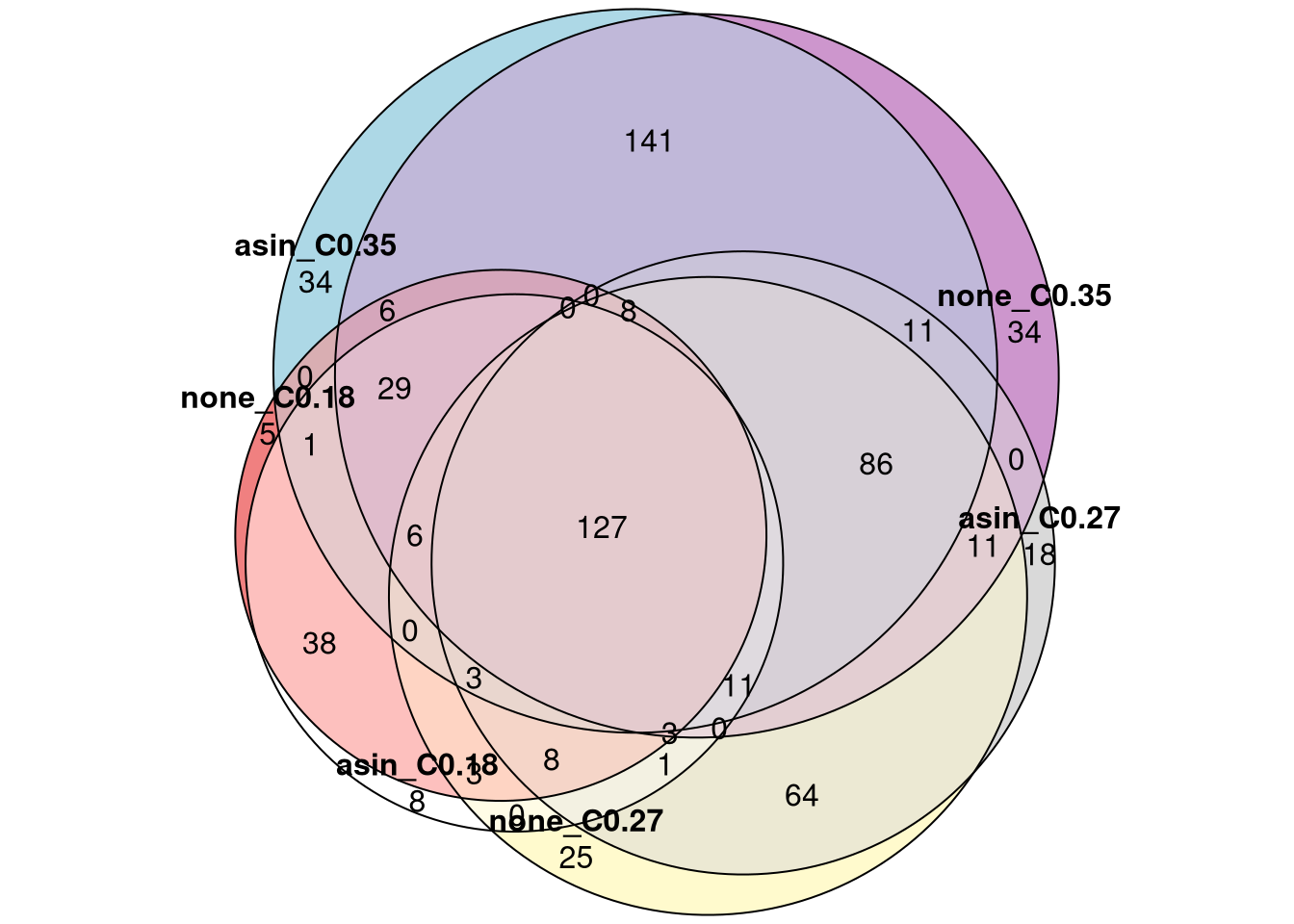
9 Figure 3
layout = "
AAAAAAAAAAAAA##
AAAAAAAAAAAAA##
AAAAAAAAAAAAA##
BBBBBBBCCCCCCCC
BBBBBBBCCCCCCCC
BBBBBBBCCCCCCCC
BBBBBBBCCCCCCCC
DDDDEEEEEFFFFFF
DDDDEEEEEFFFFFF
DDDDEEEEEFFFFFF
DDDDEEEEEFFFFFF
DDDDEEEEEFFFFFF
GGGGHHHHHIIIIII
GGGGHHHHHIIIIII
GGGGHHHHHIIIIII
GGGGHHHHHIIIIII
GGGGHHHHHIIIIII
GGGGHHHHHIIIIII
GGGGHHHHHIIIIII
"
f3 <- ((wrap_elements(full = f3a) + theme(plot.margin = unit(c(0,0,0,0), "cm"))) +
wrap_elements(f3b + theme(plot.margin = unit(rep(0,4), "cm"))) +
wrap_elements(f3c + theme(plot.margin = unit(c(0,0,0,0), "cm"))) +
wrap_elements(f3d + theme(legend.position = "top", plot.margin = unit(rep(0,4), "cm"))) +
wrap_elements(f3e + theme(legend.position = "top", plot.margin = unit(rep(0,4), "cm"))) +
wrap_elements(f3f + theme(legend.position = "top", plot.margin = unit(rep(0,4), "cm")) + guides(color = "none")) +
wrap_elements(f3g + theme(legend.position = "top", plot.margin = unit(c(0,0,0,0), "cm"))) +
wrap_elements(f3h + theme(legend.position = "top",plot.margin = unit(rep(0,4), "cm"))) +
wrap_elements(f3i + theme(legend.position = "top",plot.margin = unit(rep(0,4), "cm")))
)+
plot_layout(design = layout) +
## space is okay but must be unique
plot_annotation(tag_levels = list(c("A", "B","C", "D", "E", "F", "G", "H", "I"))) &
theme(plot.tag = element_text(size = 24, face = "bold", family = "arial"))
f3Ignoring unknown labels:
• titlte : ""Warning: No shared levels found between `names(values)` of the manual scale and the
data's colour values.Ignoring unknown labels:
• shape : "N_sample"
• linetype : "N_sample"
Ignoring unknown labels:
• shape : "N_sample"
• linetype : "N_sample"Warning: Removed 165 rows containing non-finite outside the scale range
(`stat_boxplot()`).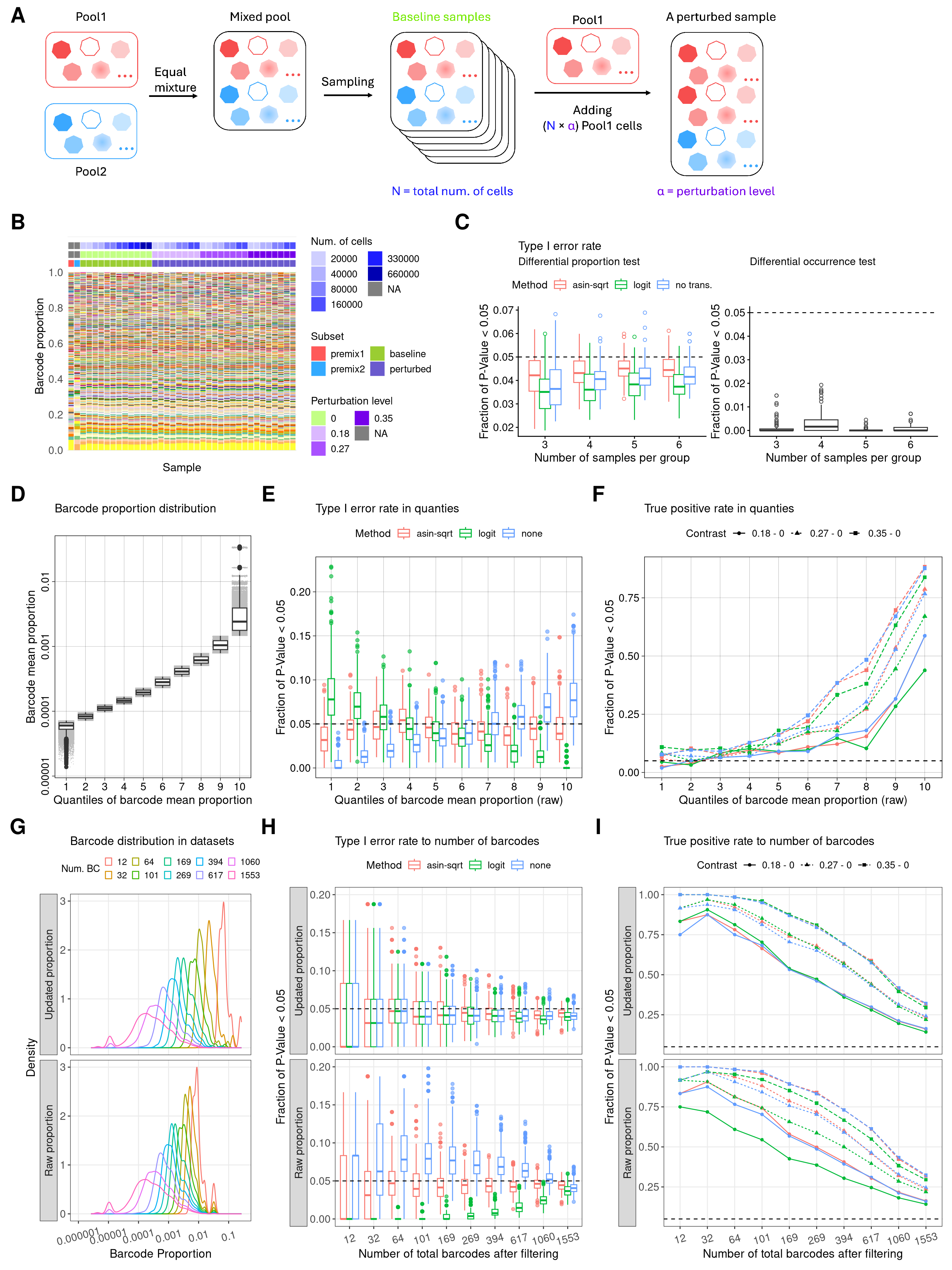
ggsave(
filename = "output/f3.png",
plot = f3,
width = 15,
height = 20,
units = "in", # for Rmd r chunk fig size, unit default to inch
dpi = 350
)Ignoring unknown labels:
• titlte : ""Warning: No shared levels found between `names(values)` of the manual scale and the
data's colour values.Ignoring unknown labels:
• shape : "N_sample"
• linetype : "N_sample"
Ignoring unknown labels:
• shape : "N_sample"
• linetype : "N_sample"Warning: Removed 165 rows containing non-finite outside the scale range
(`stat_boxplot()`).Saving this figure in f3
10 Figure S3
layoutS4 = "
AAABB
AAABB
AAABB
"
fs3 <- (
wrap_elements(fs3a + theme(plot.margin = unit(rep(0,4), "cm"))) +
wrap_elements(fs3b + theme(plot.margin = unit(c(0,0,0,0), "cm")))
)+
plot_layout(design = layoutS4) +
## space is okay but must be unique
plot_annotation(tag_levels = list(c("A", "B"))) &
theme(plot.tag = element_text(size = 24, face = "bold", family = "arial"))
fs3Warning: Removed 7 rows containing missing values or values outside the scale range
(`geom_point()`).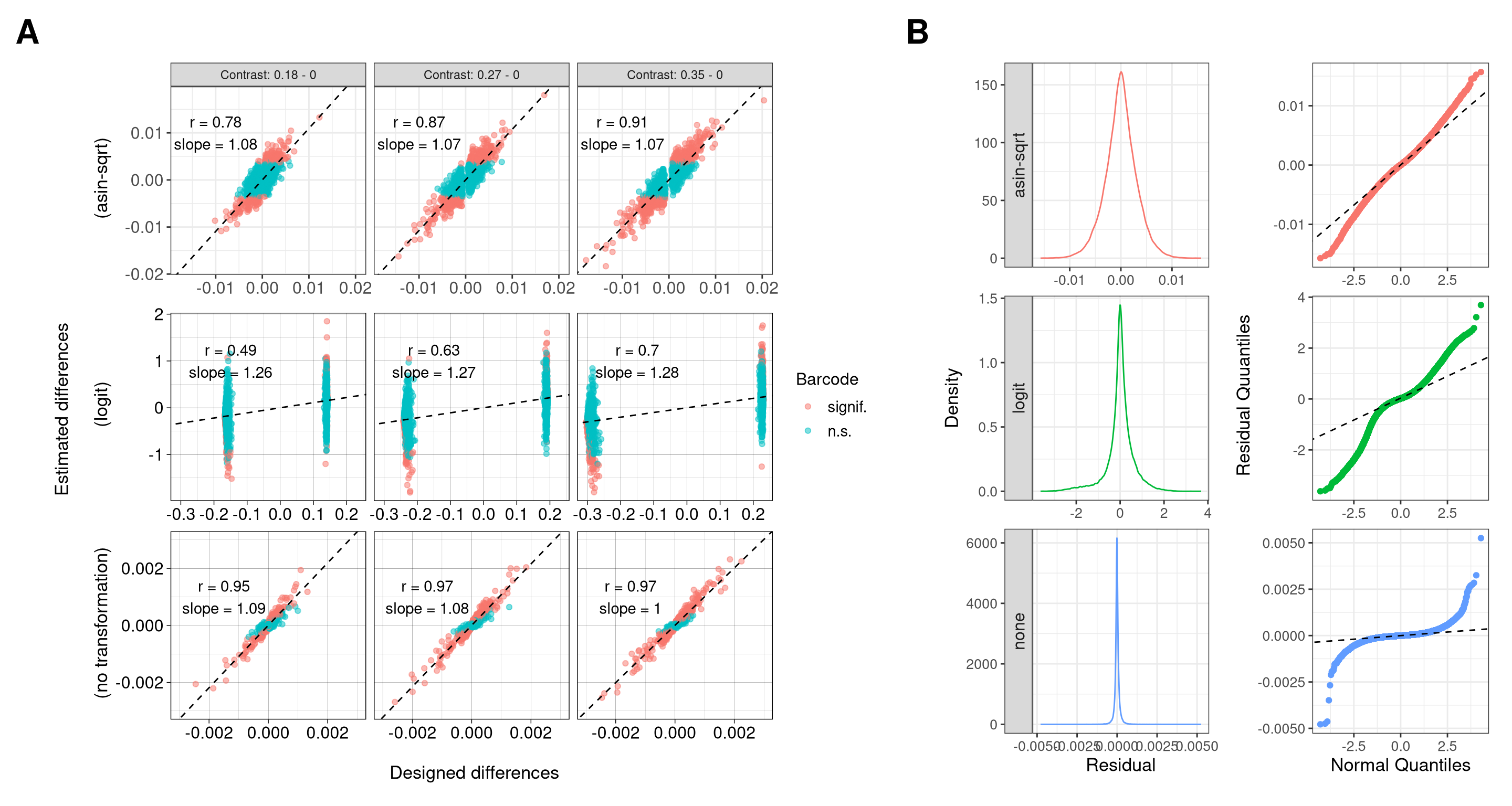
ggsave(
filename = "output/fs3.png",
plot = fs3,
width = 15,
height = 8,
units = "in", # for Rmd r chunk fig size, unit default to inch
dpi = 350
)Warning: Removed 7 rows containing missing values or values outside the scale range
(`geom_point()`).Saving this figure in fs3
sessionInfo()R version 4.5.0 (2025-04-11)
Platform: x86_64-pc-linux-gnu
Running under: Red Hat Enterprise Linux 9.6 (Plow)
Matrix products: default
BLAS/LAPACK: FlexiBLAS OPENBLAS-OPENMP; LAPACK version 3.9.0
locale:
[1] LC_CTYPE=en_AU.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_AU.UTF-8 LC_COLLATE=en_AU.UTF-8
[5] LC_MONETARY=en_AU.UTF-8 LC_MESSAGES=en_AU.UTF-8
[7] LC_PAPER=en_AU.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_AU.UTF-8 LC_IDENTIFICATION=C
time zone: Australia/Melbourne
tzcode source: system (glibc)
attached base packages:
[1] grid stats4 stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] barbieQ_1.1.3 edgeR_4.6.3
[3] limma_3.64.3 eulerr_7.0.4
[5] magick_2.9.0 ComplexHeatmap_2.24.1
[7] ggVennDiagram_1.5.4 scales_1.4.0
[9] patchwork_1.3.2 ggnewscale_0.5.2
[11] ggbreak_0.1.6 ggplot2_4.0.0
[13] SummarizedExperiment_1.38.1 Biobase_2.68.0
[15] GenomicRanges_1.60.0 GenomeInfoDb_1.44.3
[17] IRanges_2.42.0 S4Vectors_0.48.0
[19] BiocGenerics_0.54.0 generics_0.1.4
[21] MatrixGenerics_1.20.0 matrixStats_1.5.0
[23] data.table_1.17.8 knitr_1.50
[25] tibble_3.3.0 tidyr_1.3.1
[27] dplyr_1.1.4 magrittr_2.0.4
[29] readxl_1.4.5 workflowr_1.7.2
loaded via a namespace (and not attached):
[1] RColorBrewer_1.1-3 rstudioapi_0.17.1 jsonlite_2.0.0
[4] shape_1.4.6.1 jomo_2.7-6 nloptr_2.2.1
[7] farver_2.1.2 logistf_1.26.1 rmarkdown_2.30
[10] ragg_1.5.0 GlobalOptions_0.1.2 fs_1.6.6
[13] vctrs_0.6.5 minqa_1.2.8 memoise_2.0.1
[16] htmltools_0.5.8.1 S4Arrays_1.8.1 broom_1.0.10
[19] cellranger_1.1.0 SparseArray_1.8.1 gridGraphics_0.5-1
[22] mitml_0.4-5 sass_0.4.10 bslib_0.9.0
[25] cachem_1.1.0 whisker_0.4.1 igraph_2.1.4
[28] lifecycle_1.0.4 iterators_1.0.14 pkgconfig_2.0.3
[31] Matrix_1.7-3 R6_2.6.1 fastmap_1.2.0
[34] rbibutils_2.3 GenomeInfoDbData_1.2.14 clue_0.3-66
[37] digest_0.6.37 aplot_0.2.9 colorspace_2.1-2
[40] ps_1.9.1 rprojroot_2.1.1 textshaping_1.0.3
[43] labeling_0.4.3 httr_1.4.7 polyclip_1.10-7
[46] abind_1.4-8 mgcv_1.9-1 compiler_4.5.0
[49] withr_3.0.2 doParallel_1.0.17 backports_1.5.0
[52] S7_0.2.0 viridis_0.6.5 ggforce_0.5.0
[55] pan_1.9 MASS_7.3-65 rappdirs_0.3.3
[58] DelayedArray_0.34.1 rjson_0.2.23 tools_4.5.0
[61] httpuv_1.6.16 nnet_7.3-20 glue_1.8.0
[64] callr_3.7.6 nlme_3.1-168 promises_1.3.3
[67] polylabelr_0.3.0 getPass_0.2-4 cluster_2.1.8.1
[70] operator.tools_1.6.3 gtable_0.3.6 formula.tools_1.7.1
[73] tidygraph_1.3.1 XVector_0.48.0 ggrepel_0.9.6
[76] foreach_1.5.2 pillar_1.11.1 stringr_1.5.2
[79] yulab.utils_0.2.1 later_1.4.4 circlize_0.4.16
[82] splines_4.5.0 tweenr_2.0.3 lattice_0.22-6
[85] survival_3.8-3 tidyselect_1.2.1 locfit_1.5-9.12
[88] git2r_0.36.2 reformulas_0.4.1 gridExtra_2.3
[91] xfun_0.53 graphlayouts_1.2.2 statmod_1.5.0
[94] stringi_1.8.7 UCSC.utils_1.4.0 boot_1.3-31
[97] ggfun_0.2.0 yaml_2.3.10 evaluate_1.0.5
[100] codetools_0.2-20 ggraph_2.2.2 ggplotify_0.1.3
[103] cli_3.6.5 rpart_4.1.24 systemfonts_1.3.1
[106] Rdpack_2.6.4 processx_3.8.6 jquerylib_0.1.4
[109] Rcpp_1.1.0 png_0.1-8 parallel_4.5.0
[112] lme4_1.1-37 glmnet_4.1-10 viridisLite_0.4.2
[115] purrr_1.1.0 crayon_1.5.3 GetoptLong_1.0.5
[118] rlang_1.1.6 mice_3.18.0