barbieQ_paper_Figure4: Case study of
monkey HSPC
Liyang Fei
Initiate: 2025-09-07
Last update: 2026-01-11
Last updated: 2026-01-11
Checks: 5 2
Knit directory: public_barcode_count/
This reproducible R Markdown analysis was created with workflowr (version 1.7.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown file has unstaged changes. To know which version of
the R Markdown file created these results, you’ll want to first commit
it to the Git repo. If you’re still working on the analysis, you can
ignore this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
The global environment had objects present when the code in the R
Markdown file was run. These objects can affect the analysis in your R
Markdown file in unknown ways. For reproduciblity it’s best to always
run the code in an empty environment. Use wflow_publish or
wflow_build to ensure that the code is always run in an
empty environment.
The following objects were defined in the global environment when these results were created:
| Name | Class | Size |
|---|---|---|
| module | function | 5.6 Kb |
The command set.seed(20250112) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 506c314. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .RData
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: public_barcode_count.Rproj
Unstaged changes:
Modified: analysis/barbieQ_paper_Figure4.Rmd
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
These are the previous versions of the repository in which changes were
made to the R Markdown (analysis/barbieQ_paper_Figure4.Rmd)
and HTML (docs/barbieQ_paper_Figure4.html) files. If you’ve
configured a remote Git repository (see ?wflow_git_remote),
click on the hyperlinks in the table below to view the files as they
were in that past version.
| File | Version | Author | Date | Message |
|---|---|---|---|---|
| Rmd | f48add2 | FeiLiyang | 2026-01-07 | supple analyses |
| Rmd | 34f5894 | FeiLiyang | 2026-01-01 | reorder f3 |
| Rmd | 7b7813d | FeiLiyang | 2025-12-17 | update f3 fs4 |
| html | 7b7813d | FeiLiyang | 2025-12-17 | update f3 fs4 |
| Rmd | c2766f9 | Liyang Fei | 2025-10-07 | remake figures |
1 Load Dependencies
library(readxl)
library(magrittr)
library(dplyr)
library(tidyr) # for pivot_longer
library(tibble) # for rownames_to _column
library(knitr) # for kable()
library(data.table) # for data.table()
library(SummarizedExperiment)
library(S4Vectors)
library(grid)
library(ggplot2)
library(ggbreak)
library(ggnewscale) # ggnewscale::new_scale_fill()
library(patchwork)
library(scales)
library(ggVennDiagram)
library(ComplexHeatmap)
library(magick)
library(eulerr)
library(edgeR)
source("analysis/plotBarcodeHistogram.R") ## accommodated from bartools::plotBarcodehistogram
source("analysis/F4_validation.R") ## functions for Figure4 validations
source("analysis/ggplot_theme.R") ## setting ggplot theme2 Install
barbieQ
Installing the latest devel version of barbieQ from
GitHub.
if (!requireNamespace("barbieQ", quietly = TRUE)) {
remotes::install_github("Oshlack/barbieQ")
}Warning: replacing previous import 'data.table::first' by 'dplyr::first' when
loading 'barbieQ'Warning: replacing previous import 'data.table::last' by 'dplyr::last' when
loading 'barbieQ'Warning: replacing previous import 'data.table::between' by 'dplyr::between'
when loading 'barbieQ'Warning: replacing previous import 'dplyr::as_data_frame' by
'igraph::as_data_frame' when loading 'barbieQ'Warning: replacing previous import 'dplyr::groups' by 'igraph::groups' when
loading 'barbieQ'Warning: replacing previous import 'dplyr::union' by 'igraph::union' when
loading 'barbieQ'library(barbieQ)Check the version of barbieQ.
packageVersion("barbieQ")[1] '1.1.3'3 Set seeds
set.seed(2025)4 Load preprocessed data
Loading barbieQ object preprocessed from barbieQ_paper_Figure2
load("output/monkeyHSPC_barbieQ.rda")
# load("output/monkeyHSPC_raw_barbieQ.rda")5 F4A: Experimental schematic
schematic <- image_read("output/f4a_schematic.jpeg")
f4a <- rasterGrob(schematic, interpolate = FALSE)
wrap_elements(full = f4a)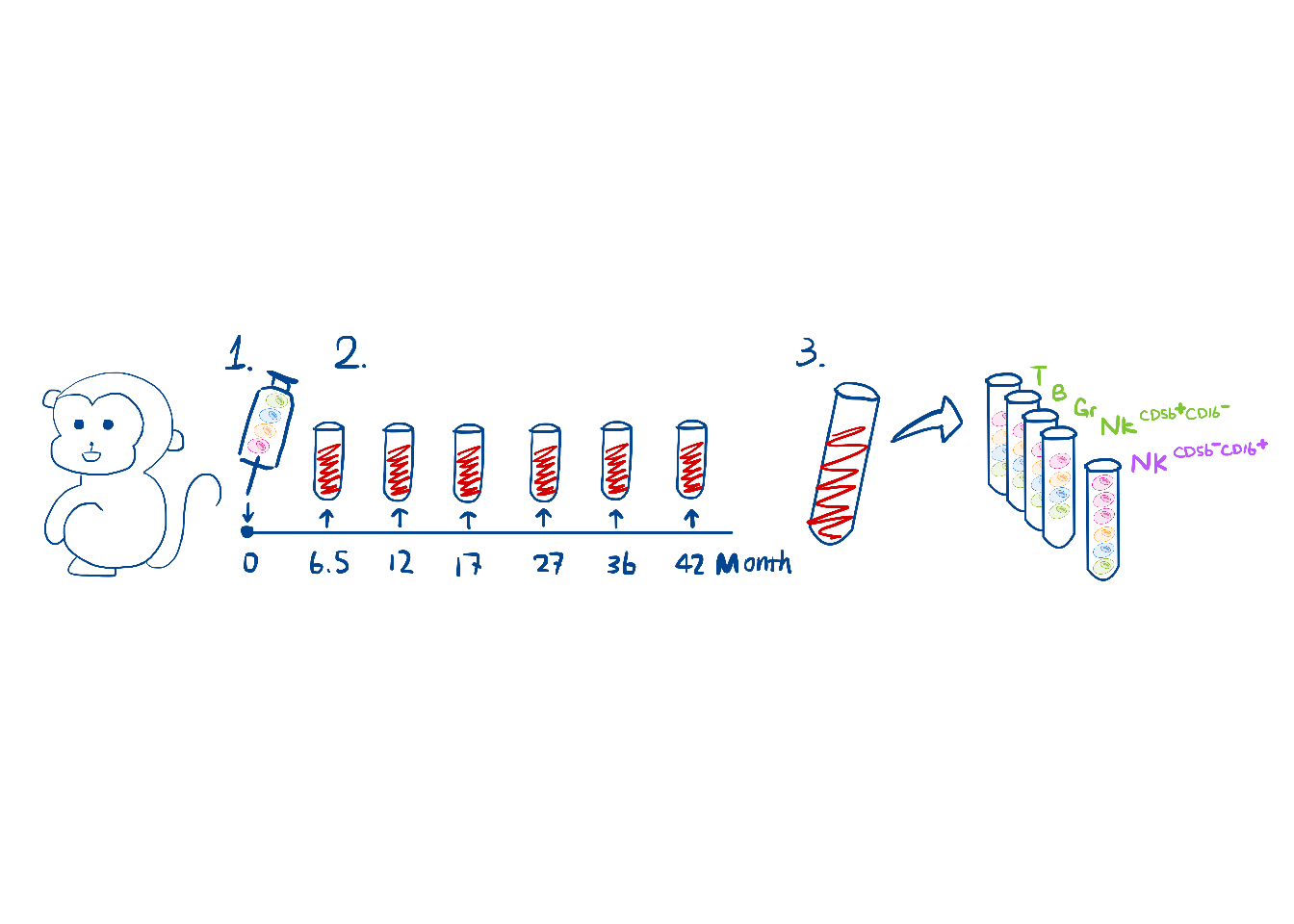
| Version | Author | Date |
|---|---|---|
| dfa737a | FeiLiyang | 2025-12-17 |
6 F4B: Plot barcode proportion and sample annotation
plotBarcodeHistogram is customized from the
bartools package.
## get sample conditions and color mapping
metadata <- monkeyHSPC_merged_top$sampleMetadata %>% as.data.frame()
metadata$sample <- rownames(metadata)
metadata$y = 1.05 ## for ggplot tile position
color_mapping <- monkeyHSPC_merged_top@metadata$factorColors
## order samples
order_sample <- metadata %>% with(order(Celltype, Months))
monkey_dge <- DGEList(
counts = assay(monkeyHSPC_merged_top),
group = metadata$Celltype)
## pull out ggplot data
p_stacked_prop <- plotBarcodeHistogram(monkey_dge, orderSamples = colnames(monkeyHSPC_merged_top)[order_sample])Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"barcode"` instead of `.data$barcode`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"value"` instead of `.data$value`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"name"` instead of `.data$name`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"freq"` instead of `.data$freq`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.f4b <- p_stacked_prop +
## label perturbations
ggnewscale::new_scale_fill() +
geom_tile(data = metadata, aes(x = sample,y = y, fill = Celltype), height = 0.04, inherit.aes = FALSE, color = "white") +
scale_fill_manual(values = color_mapping$Celltype) +
labs(fill = "Cell type") +
## label libsize
ggnewscale::new_scale_fill() +
geom_tile(data = metadata, aes(x = sample,y = y+0.05, fill = as.factor(Months)), height = 0.04, inherit.aes = FALSE, color = "white") +
scale_fill_manual(values = setNames(attr(color_mapping$Months, "colors"), attr(color_mapping$Months, "breaks"))) +
labs(fill = "Month") +
labs(x = "Sample", y = "Barcode proportion") +
theme_minimal() +
scale_y_continuous(breaks = seq(0, 1, by = 0.2)) +
theme(axis.text.x = element_blank(), panel.grid.major.x = element_blank())
f4bWarning: The `guide` argument in `scale_*()` cannot be `FALSE`. This was deprecated in
ggplot2 3.3.4.
ℹ Please use "none" instead.
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.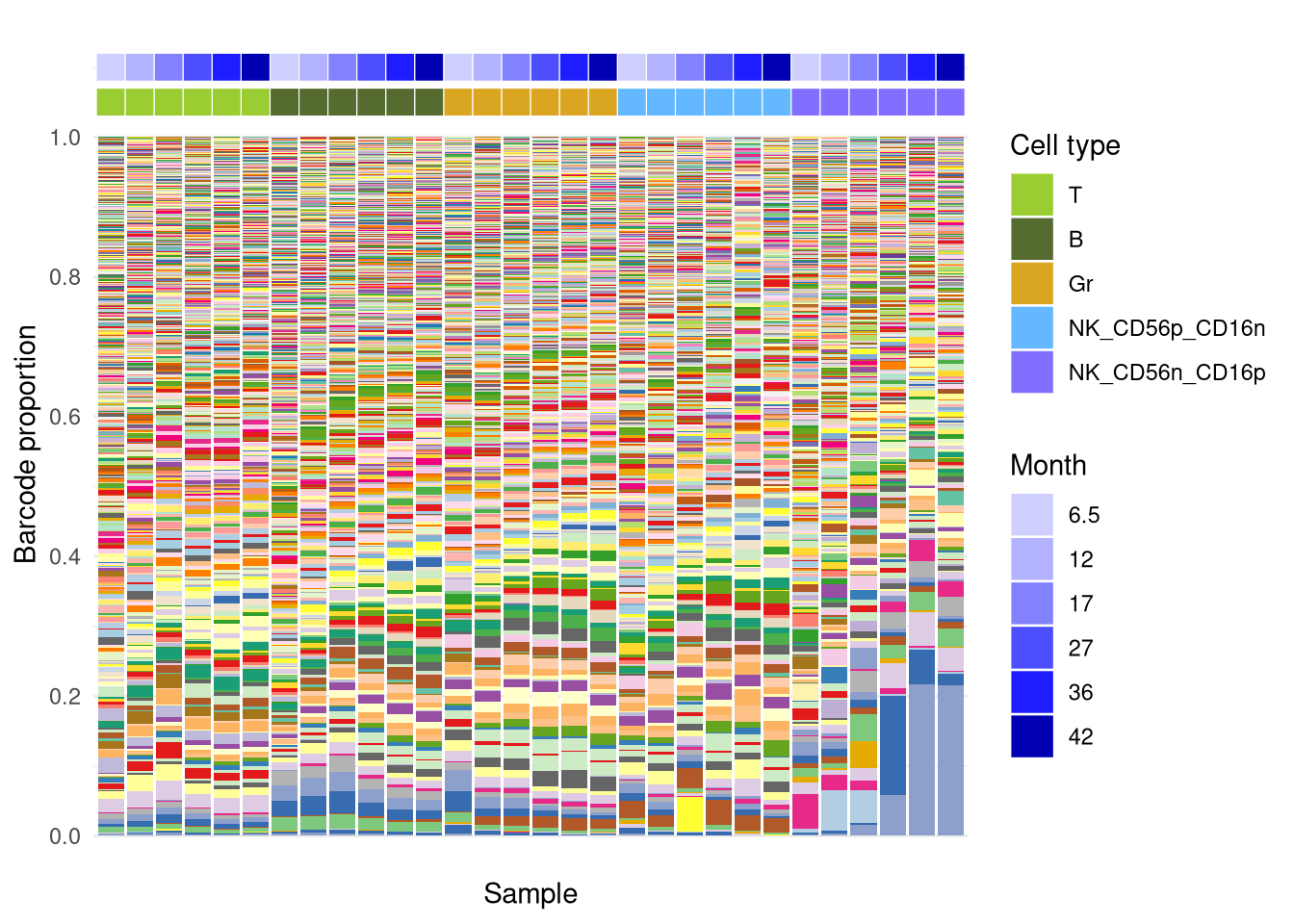
| Version | Author | Date |
|---|---|---|
| dfa737a | FeiLiyang | 2025-12-17 |
7 Compare filtering
7.1 F4D: top10each vs. barbieQ filter
The original paper filter barcodes by tagging top 10 barcodes from
each sample, and collectively keeping the tagged barcodes across all
samples. Here we compare this “top10each” (and “top20each”) strategy
with barbieQ filtering.
The tagTopN function is sourced from
“F4_validation.R”.
## tagging top 10 in all samples
colTags <- lapply(as.data.frame(assays(monkeyHSPC_merged)$proportion), function(col) tagTopN(col, N = 10))
colTagMat <- do.call(cbind, colTags) %>% as.matrix()
colnames(colTagMat) <- colnames(monkeyHSPC_merged)
rownames(colTagMat) <- rownames(monkeyHSPC_merged)
## filtering by keeping barcodes tagged for at least once
colTagVec <- rowSums(colTagMat) >= 1
## ensuring we've tagged 10 barcodes within each sample
all(colSums(colTagMat) == 10)[1] TRUE## copying the barbieQ object and manually labeling the filtered barcodes based on the above strategy
monkeyHSPC_top10each <- monkeyHSPC_merged
rowData(monkeyHSPC_top10each)$isTopBarcode$isTop <- colTagVec
assays(monkeyHSPC_top10each)$isTopAssay <- colTagMat
p_sankey_top10 <- barbieQ::plotBarcodeSankey(monkeyHSPC_top10each)
p_sankey_barbieQ <- barbieQ::plotBarcodeSankey(monkeyHSPC_merged)
## repeat the process for top20 each
colTags2 <- lapply(as.data.frame(assays(monkeyHSPC_merged)$proportion), function(col) tagTopN(col, N = 20))
colTagMat2 <- do.call(cbind, colTags2) %>% as.matrix()
colnames(colTagMat2) <- colnames(monkeyHSPC_merged)
rownames(colTagMat2) <- rownames(monkeyHSPC_merged)
colTagVec2 <- rowSums(colTagMat2) >= 1
all(colSums(colTagMat2) == 20)[1] TRUEmonkeyHSPC_top20each <- monkeyHSPC_merged
rowData(monkeyHSPC_top20each)$isTopBarcode$isTop <- colTagVec2
assays(monkeyHSPC_top20each)$isTopAssay <- colTagMat2
p_sankey_top20 <- barbieQ::plotBarcodeSankey(monkeyHSPC_top20each)
## plot sankeys
f4d <- p_sankey_barbieQ +
theme(legend.position = "none", plot.title = element_text(hjust = 0.5, vjust = 1),
strip.text.x = element_blank(), strip.text.y = element_blank()) + labs(title = "barbieQ") +
p_sankey_top10 +
theme(legend.position = "none", plot.title = element_text(hjust = 0.5, vjust = 1),
strip.text.x = element_blank(), strip.text.y = element_blank()) + labs(title = "top10each") +
p_sankey_top20 +
theme(legend.position = "right", plot.title = element_text(hjust = 0.5, vjust = 1),
strip.text.x = element_blank(), strip.text.y = element_blank()) + labs(title = "top20each")
f4d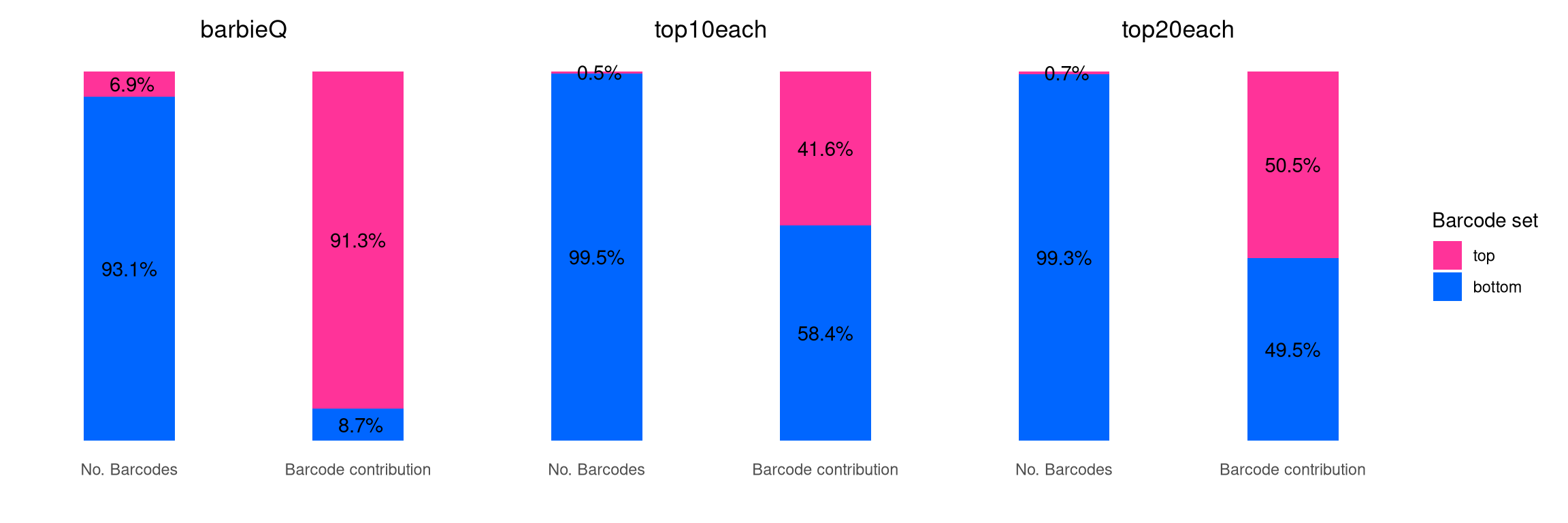
| Version | Author | Date |
|---|---|---|
| dfa737a | FeiLiyang | 2025-12-17 |
7.2 F4C: overlap
list_top_BC <- list(
barbieQ = rownames(monkeyHSPC_merged)[monkeyHSPC_merged@elementMetadata$isTopBarcode$isTop],
top10each = rownames(monkeyHSPC_merged)[colTagVec],
top20each = rownames(monkeyHSPC_merged)[colTagVec2]
)
## plot Euler
# venn_filter <- eulerr::euler(list_top_BC) %>%
# plot(fills = c("pink", "skyblue", "steelblue"),
# alpha = 0.5, labels = list(font = 4, cex = 1), quantities = list(font = 1, cex = 1))
#
# venn_filter %>% plot() %>% grid.grabExpr() %>% wrap_plots()
## plot ggVenn
# ggVennDiagram(list_top_BC, label = "count") + scale_fill_gradient(low="white", high = "skyblue")
## plot UpSet plot
f4c <- list_top_BC %>%
Venn() %>%
plot_upset(top.bar.numbers.size = 4, relative_width = 0.0) ## limit space for left bar plot
## getting theme settings from an example ggplot using barbieQ theme
tester <- ggplot(data.frame(X = 1:3, Y = 4:6)) +
geom_point(aes(x = X, y = Y)) +
theme_barbieQ_size()
tester_col <- ggplot(data.frame(X = 1:3, Y = 4:6)) +
geom_col(aes(x = X, y = Y), width = 0.2)
tester_col@layers$geom_col$aes_params$width[1] 0.2mytittle <- tester@theme$axis.title
mytext <- tester@theme$axis.text
## modifying components in UpSet plots: three gplot objects
f4c[["plotlist"]][[1]]@theme$axis.title <- mytittle
f4c[["plotlist"]][[1]]@theme$axis.text <- mytext
f4c[["plotlist"]][[2]]@theme$axis.title <- mytittle
f4c[["plotlist"]][[2]]@theme$axis.text <- mytext
f4c[["plotlist"]][[2]]@layers$geom_col$aes_params$width <- 0.4
f4c[["plotlist"]][[3]]@theme$axis.title <- mytittle
f4c[["plotlist"]][[3]]@theme$axis.text <- mytext
f4c[["plotlist"]][[3]]@theme$axis.text@size <- 11
f4c[["plotlist"]][[3]]@theme$axis.text@angle <- 15
## removing the left bar plot
f4c[["plotlist"]][[3]] <- f4c[["plotlist"]][[3]] + theme(
panel.grid = element_blank(),
axis.text = element_blank(),
axis.title = element_blank(),
axis.ticks = element_blank()
)
## plot 4c
f4c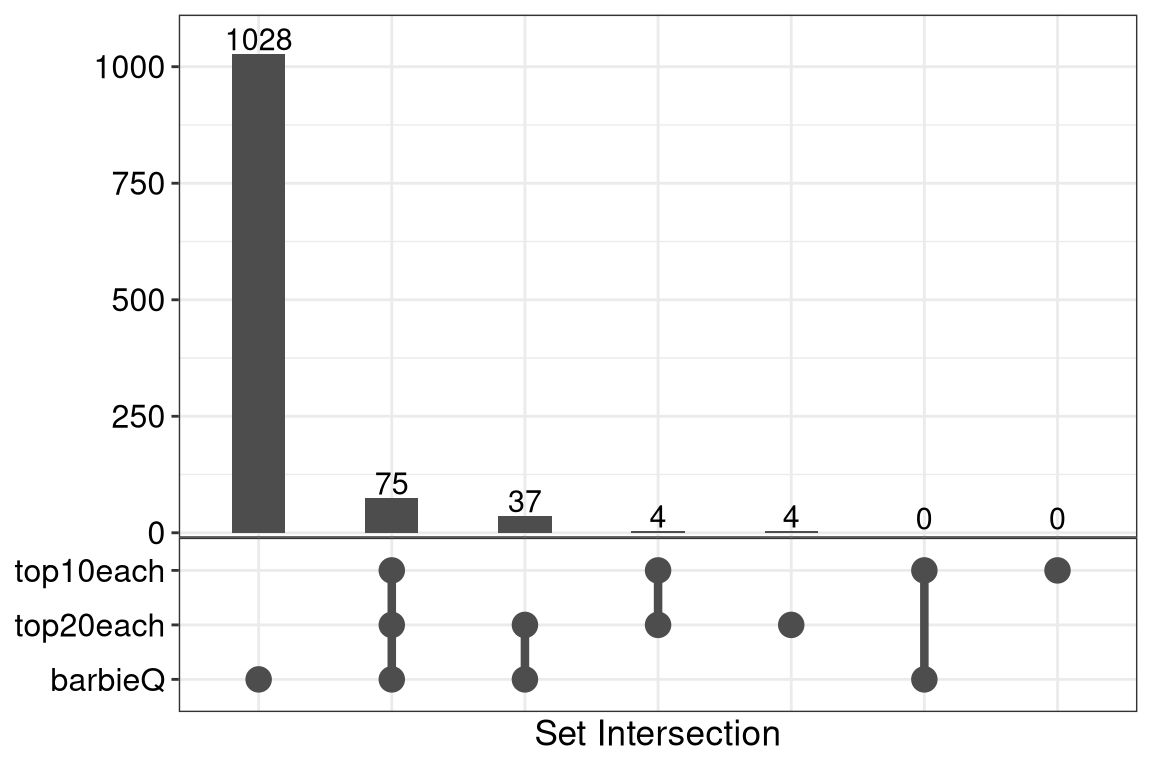
| Version | Author | Date |
|---|---|---|
| dfa737a | FeiLiyang | 2025-12-17 |
7.3 F4E: inspect two sets: heatmap
barcodeArray <- monkeyHSPC_merged@elementMetadata$isTopBarcode$isTop | colTagVec
length(colTagVec)[1] 16598length(monkeyHSPC_merged@elementMetadata$isTopBarcode$isTop)[1] 16598table(barcodeArray)barcodeArray
FALSE TRUE
15454 1144 monkeyHSPC_compare_filter <- monkeyHSPC_merged[barcodeArray,]
# Create a named vector indicating where each element belongs
region_labels <- data.frame(
barcode = rownames(monkeyHSPC_compare_filter),
region = "both")
region_labels[monkeyHSPC_merged@elementMetadata$isTopBarcode$isTop[barcodeArray] & !(colTagVec[barcodeArray]), "region"] <- "barbieQ only"
region_labels[!(monkeyHSPC_merged@elementMetadata$isTopBarcode$isTop[barcodeArray]) & colTagVec[barcodeArray], "region"] <- "top10each only"
region_labels$region %>% table().
barbieQ only both top10each only
1065 75 4 ht_filter <- plotBarcodeHeatmap(
monkeyHSPC_compare_filter,
barcodeAnnotation = rowAnnotation(
filteredBy = region_labels$region,
col = list(filteredBy = c("barbieQ only" = "hotpink", "both" = "slateblue", "top10each only" = "steelblue1")),
annotation_name_side = "top", annotation_name_rot = 45, annotation_name_gp = grid::gpar(fontsize = 10)))setting Celltype as the primary factor in `sampleMetadata`.The automatically generated colors map from the 1^st and 99^th of the
values in the matrix. There are outliers in the matrix whose patterns
might be hidden by this color mapping. You can manually set the color
to `col` argument.
Use `suppressMessages()` to turn off this message.
The automatically generated colors map from the 1^st and 99^th of the
values in the matrix. There are outliers in the matrix whose patterns
might be hidden by this color mapping. You can manually set the color
to `col` argument.
Use `suppressMessages()` to turn off this message.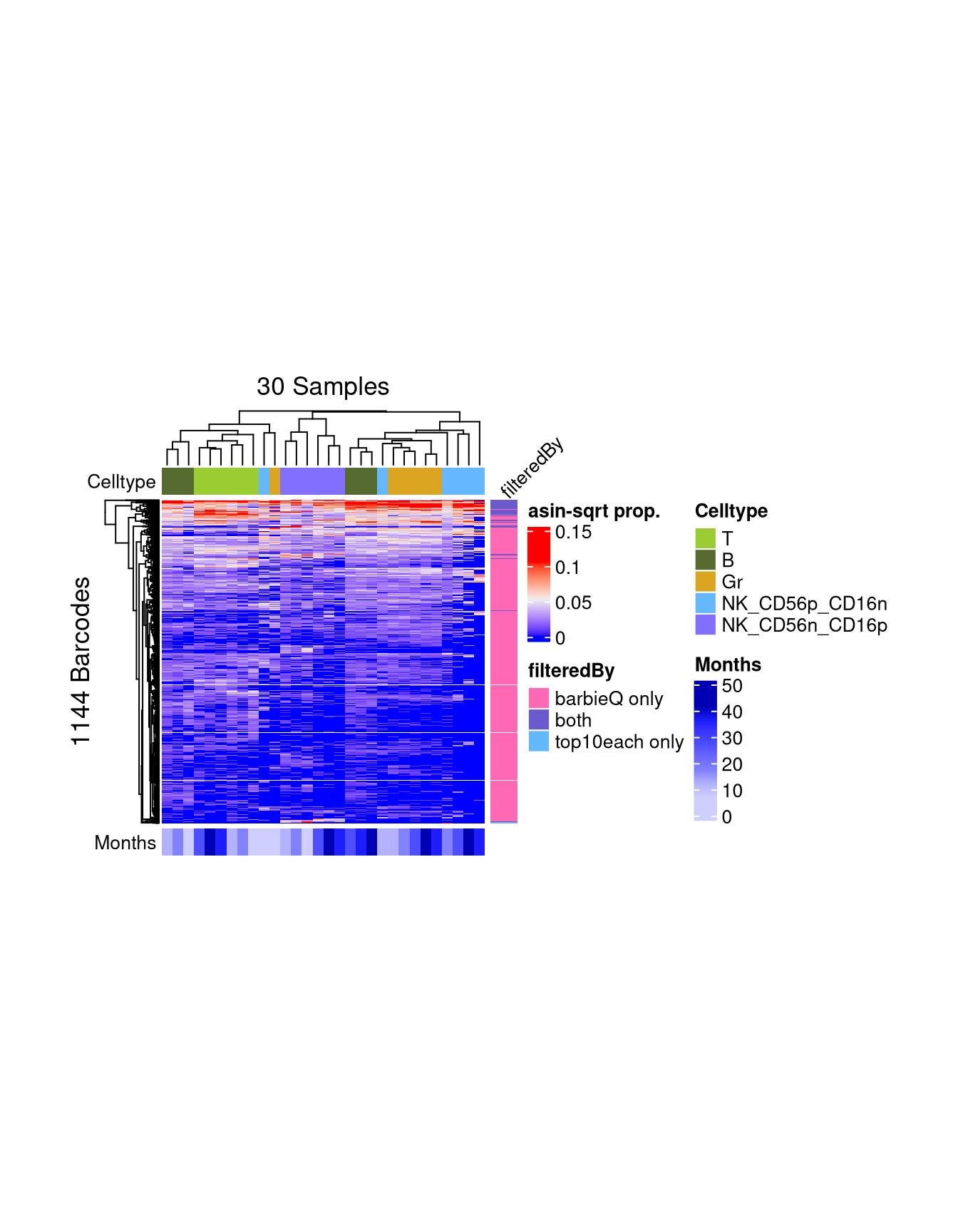
| Version | Author | Date |
|---|---|---|
| dfa737a | FeiLiyang | 2025-12-17 |
matrix color is mapped to `asin-sqrt proportion` but labeled by raw proportion.## ht_filter@heatmap_param[["calling_env"]][["sampleAnnotation"]]@anno_list[["Months"]]@show_legend <- FALSE
## ht_filter@heatmap_param[["calling_env"]][["groupAnnotation"]]@anno_list[["Celltype"]]@show_legend <- FALSE
f4e <- draw(ht_filter, row_split = region_labels$region, width = unit(6, "cm"), height = unit(15, "cm")) %>%
grid.grabExpr() %>% list() %>% wrap_plots()'width' should not be set in draw() for horizontal heatmap list (Note a
single heatmap is a horizontal heatmap list). Please directly set it in
`Heatmap()`.f4e <- Heatmap(
monkeyHSPC_compare_filter@assays@data$proportion %>% sqrt() %>% asin(), name = "asin-sqrt prop.",
cluster_rows = TRUE, cluster_columns = TRUE, show_row_names = FALSE, show_column_names = FALSE,
right_annotation = rowAnnotation(
filteredBy = region_labels$region,
col = list(filteredBy = c("barbieQ only" = "hotpink", "both" = "slateblue", "top10each only" = "steelblue1")),
annotation_name_side = "top", annotation_name_rot = 45),
row_split = region_labels$region, width = unit(6, "cm"), height = unit(15, "cm"), row_title = "Barcodes",
top_annotation = HeatmapAnnotation(
Celltype = monkeyHSPC_compare_filter$sampleMetadata$Celltype,
Months = monkeyHSPC_compare_filter$sampleMetadata$Months,
col = monkeyHSPC_compare_filter@metadata$factorColors,
show_legend = FALSE, annotation_name_side = "left"
))The automatically generated colors map from the 1^st and 99^th of the
values in the matrix. There are outliers in the matrix whose patterns
might be hidden by this color mapping. You can manually set the color
to `col` argument.
Use `suppressMessages()` to turn off this message.f4e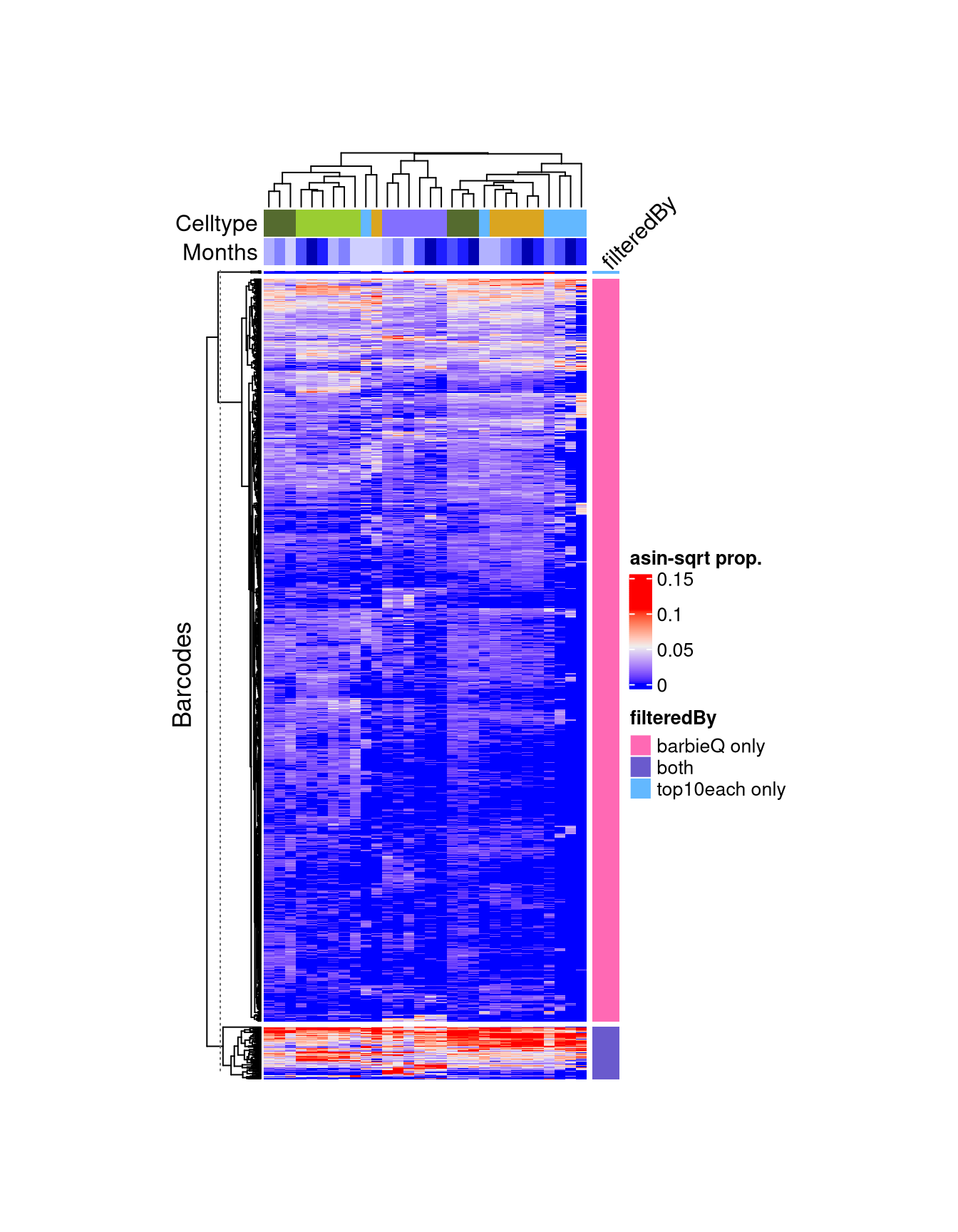
| Version | Author | Date |
|---|---|---|
| dfa737a | FeiLiyang | 2025-12-17 |
8 Compare difference identification on top10each filtering approach
8.1 barbieQ filter + barbieQ test
monkeyHSPC_top10each@elementMetadata$isTopBarcode$isTop %>% table().
FALSE TRUE
16519 79 ## subset the 79 barcodes filtered by top10each
monkeyHSPC_top10each_79 <- monkeyHSPC_top10each[monkeyHSPC_top10each@elementMetadata$isTopBarcode$isTop,]
##need to take Time into account
designs <- monkeyHSPC_top10each_79$sampleMetadata %>% as.data.frame() %>%
mutate(Months = as.factor(Months)) %>%
with(model.matrix(~0+ Celltype + Months))
monkeyHSPC_top10each_79 <- testBarcodeSignif(
barbieQ = monkeyHSPC_top10each_79, designMatrix = designs,
contrastFormula = "(CelltypeNK_CD56n_CD16p) - (CelltypeB+CelltypeGr+CelltypeT+CelltypeNK_CD56p_CD16n)/4",
method = "diffProp", transformation = "asin-sqrt"
)setting Celltype as the primary factor in `sampleMetadata`.deleting 3 nested factor(s) because `designMatrix` must be full rank.no block specified, so there are no duplicate measurements.# check design
monkeyHSPC_top10each_79@elementMetadata$testingBarcode@metadata$contrasts Contrasts
Levels (CelltypeNK_CD56n_CD16p) - (CelltypeB+CelltypeGr+CelltypeT+CelltypeNK_CD56p_CD16n)/4
CelltypeT -0.25
CelltypeB -0.25
CelltypeGr -0.25
CelltypeNK_CD56p_CD16n -0.25
CelltypeNK_CD56n_CD16p 1.00
Months12 0.00
Months17 0.00
Months27 0.00
Months36 0.00
Months42 0.00monkeyHSPC_top10each_79@elementMetadata$testingBarcode@metadata$design %>% data.table(keep.rownames = T)8.2 barbieQ filter + LFC approach
1% contribution to CD56−CD16+ cells and
10-fold abundance in contribution to CD56−CD16+ cells compared with T, B, Gr, and CD56+CD16− cells.
calc_bias_LFC function sourced from
“F4_validation.R”.
list_BC_top10each_LFC <- calc_bias_LFC(monkeyHSPC_top10each_79)
union_LFC_top10each <- unlist(list_BC_top10each_LFC) %>% unique()## check overlaps between months
venn_top10each_LFC <- eulerr::euler(list_BC_top10each_LFC)
plot(venn_top10each_LFC, fills = c("white"), alpha = 0.5, labels = T, quantities = T)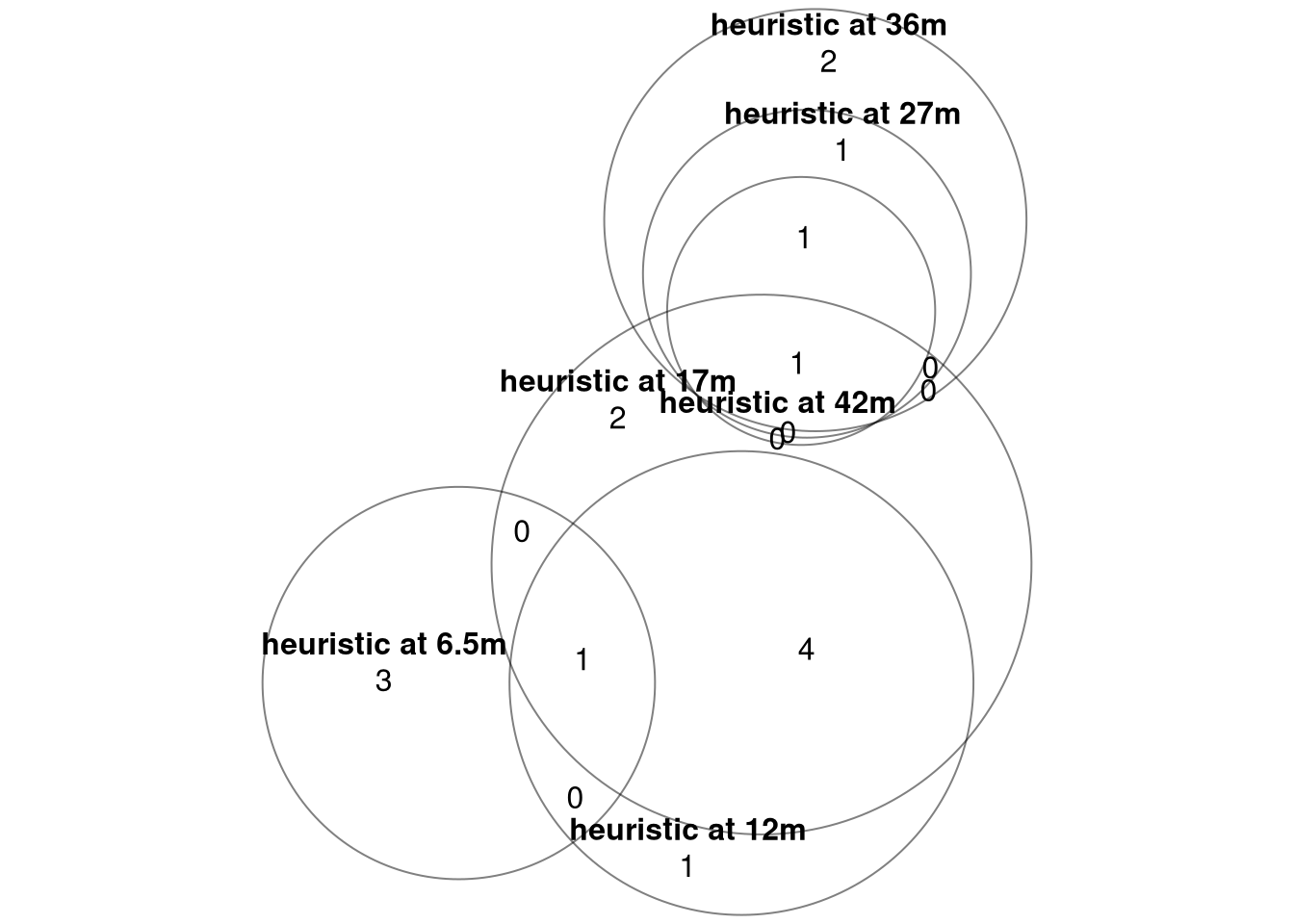
8.3 F4G: overlap
list_top10each_feature <- list(
barbieQ = rownames(monkeyHSPC_top10each_79)[monkeyHSPC_top10each_79@elementMetadata$testingBarcode$tendencyTo == "CelltypeNK_CD56n_CD16p"],
heuristics = union_LFC_top10each)
## plot Euler
# venn_top10each_feature <- eulerr::euler(list_top10each_feature) %>%
# plot(fills = c("pink","slateblue"),
# alpha = 0.5, labels = list(font = 4, cex = 1), quantities = list(font = 1, cex = 1))
#
# venn_top10each_feature %>% plot() %>% grid.grabExpr() %>% wrap_plots()
## plot UpSet plot
f4g <- list_top10each_feature %>%
Venn() %>%
plot_upset(top.bar.numbers.size = 4, relative_width = 0.0) ## limit left bar plot
## getting theme settings from an example ggplot using barbieQ theme
tester <- ggplot(data.frame(X = 1:3, Y = 4:6)) +
geom_point(aes(x = X, y = Y)) +
theme_barbieQ_size()
mytittle <- tester@theme$axis.title
mytext <- tester@theme$axis.text
## modifying components in UpSet plots: three gplot objects
f4g[["plotlist"]][[1]]@theme$axis.title <- mytittle
f4g[["plotlist"]][[1]]@theme$axis.text <- mytext
f4g[["plotlist"]][[2]]@theme$axis.title <- mytittle
f4g[["plotlist"]][[2]]@theme$axis.text <- mytext
f4g[["plotlist"]][[2]]@layers$geom_col$aes_params$width <- 0.4
f4g[["plotlist"]][[3]]@theme$axis.title <- mytittle
f4g[["plotlist"]][[3]]@theme$axis.text <- mytext
f4g[["plotlist"]][[3]]@theme$axis.text@size <- 11
f4g[["plotlist"]][[3]]@theme$axis.text@angle <- 15
## removing the left bar plot
f4g[["plotlist"]][[3]] <- f4g[["plotlist"]][[3]] + theme(
panel.grid = element_blank(),
axis.text = element_blank(),
axis.title = element_blank(),
axis.ticks = element_blank()
)
## plot 4g
f4g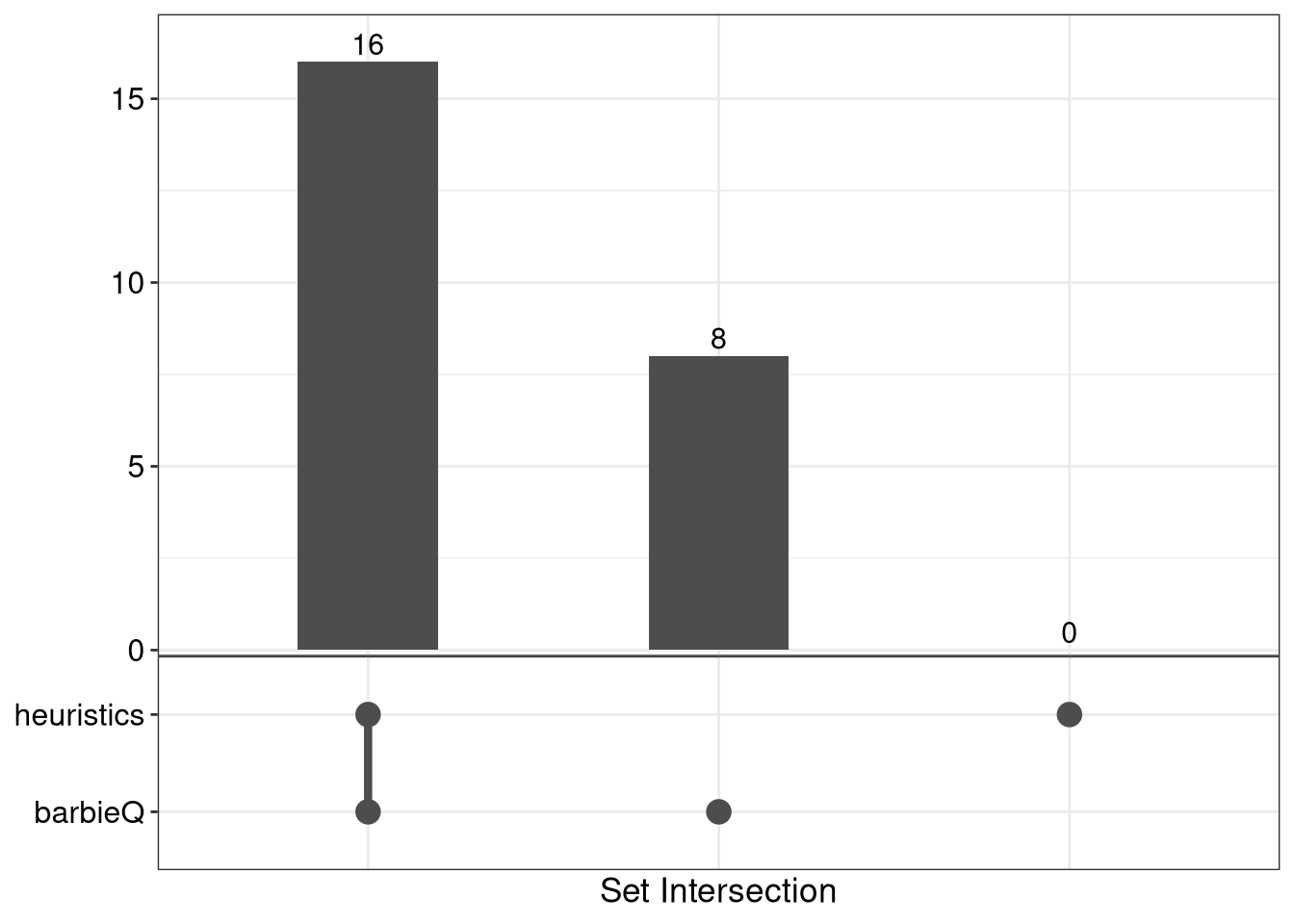
| Version | Author | Date |
|---|---|---|
| dfa737a | FeiLiyang | 2025-12-17 |
8.4 F4F: inspect two sets: boxplot
- Box plot to visualize the LFC of sets of selected barcodes.
FC_top10each <- calc_bias_LFC(barbieQ = monkeyHSPC_top10each_79, get_FC = T) %>% as.data.frame()
rownames(FC_top10each) <- rownames(monkeyHSPC_top10each_79)
## annotating BC by barbieQ seletion
FC_top10each$bbq <- monkeyHSPC_top10each_79@elementMetadata$testingBarcode$direction
## annotating BC by heuristic selection
FC_top10each$heu = 0
FC_top10each[union_LFC_top10each, "heu"] = 1
## adding BC name
FC_top10each <- FC_top10each %>%
mutate(Barcode = rownames(monkeyHSPC_top10each_79)) %>%
## summarizing two selecting methods overlaps
mutate(identifiedBy = ifelse(bbq == 1,
ifelse(heu == 1, "both", "barbieQ only"),
ifelse(heu == 1, "heuristics only", "neither"))) %>%
dplyr::select(-c(heu, bbq))
## extract FC from testing
coff_FC <- monkeyHSPC_top10each_79@elementMetadata$testingBarcode$meanDiff
FC_top10each$testing_coeff <- coff_FC
long_FC_top10each <- FC_top10each %>%
mutate(Barcode = rownames(monkeyHSPC_top10each_79)) %>%
pivot_longer(
cols = -c(Barcode, identifiedBy, testing_coeff), # keep Celltype and Months fixed
names_to = "Month",
values_to = "FC"
)Draw box plot, barcodes ordered by mean LFC
## order BC in x axis by contrast coefficient
barcode_order <- long_FC_top10each %>%
group_by(Barcode) %>%
summarise(mean_LFC = mean((testing_coeff), na.rm = TRUE)) %>%
arrange(mean_LFC) %>%
pull(Barcode)
long_FC_top10each$identifiedBy <- factor(
long_FC_top10each$identifiedBy,
levels = c("barbieQ only", "heuristics only", "both", "neither")
)
## remove level "heuristics only"
long_FC_top10each$identifiedBy <- factor(
long_FC_top10each$identifiedBy,
levels = c("barbieQ only", "both", "neither")
)
p_box_coef <- ggplot(long_FC_top10each, aes(x = factor(Barcode, levels = barcode_order), y = log2(FC), group = (Barcode), color = identifiedBy)) +
geom_boxplot(varwidth = T) +
# geom_jitter()+
theme_classic() +
theme(axis.text.x = element_blank(), legend.position = "right", axis.title = element_text(size = 14)) +
geom_abline(slope = 0, intercept = 0, linetype = 2, colour = "snow4") +
labs(x = "",
y = "LFC",
title = "Barocdes selected by top10each filtering") +
scale_color_manual(
values = c("both" = "slateblue", "barbieQ only" = "hotpink", "heuristics only" = "steelblue1", "neither" = "slategray"),
drop = FALSE)
p_box_coef_2 <- ggplot(long_FC_top10each, aes(x = factor(Barcode, levels = barcode_order), y = testing_coeff, group = (Barcode), color = identifiedBy)) +
geom_boxplot(varwidth = T) +
theme_classic() +
theme(axis.text.x = element_blank(), legend.position = "none", axis.title = element_text(size = 14)) +
geom_abline(slope = 0, intercept = 0, linetype = 2, colour = "snow4") +
labs(x = "Barocdes ordered by barbieQ-estimated size effect",
y = "Size effect") +
scale_color_manual(values = c("both" = "slateblue", "barbieQ only" = "hotpink", "heuristics only" = "steelblue1", "neither" = "slategray"))
f4f <- p_box_coef / p_box_coef_2 + plot_layout(heights = c(3, 1))
f4f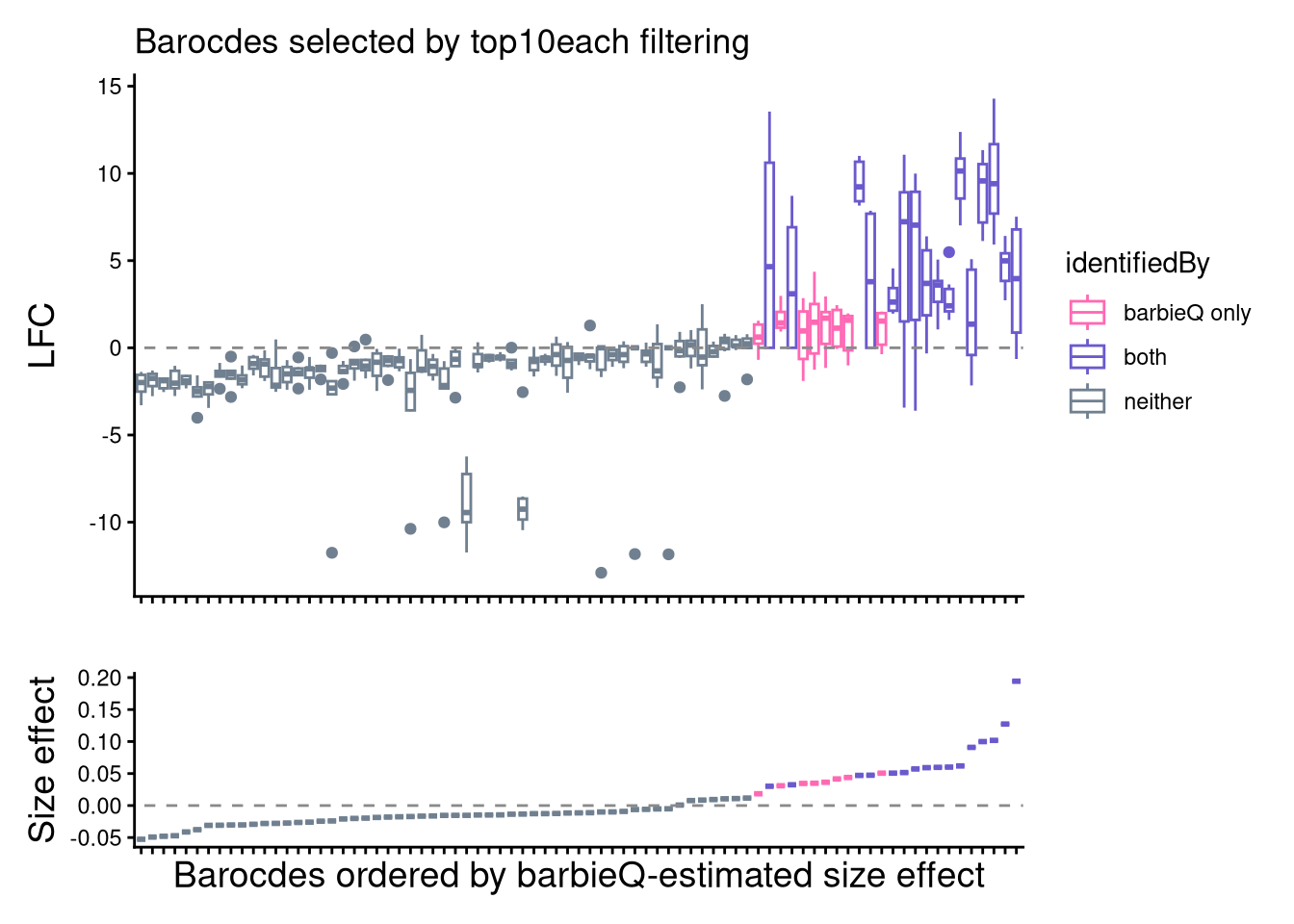
9 Compare fitering with barbieQ testing
9.1 top10each + barbieQ test
##need to take Time into account
designs <- monkeyHSPC_merged_top$sampleMetadata %>% as.data.frame() %>%
mutate(Months = as.factor(Months)) %>%
with(model.matrix(~0+ Celltype + Months))
monkeyHSPC_merged_top <- testBarcodeSignif(
barbieQ = monkeyHSPC_merged_top, designMatrix = designs,
contrastFormula = "(CelltypeNK_CD56n_CD16p) - (CelltypeB+CelltypeGr+CelltypeT+CelltypeNK_CD56p_CD16n)/4",
method = "diffProp", transformation = "asin-sqrt"
)setting Celltype as the primary factor in `sampleMetadata`.deleting 3 nested factor(s) because `designMatrix` must be full rank.no block specified, so there are no duplicate measurements.# check design
monkeyHSPC_merged_top@elementMetadata$testingBarcode@metadata$contrasts Contrasts
Levels (CelltypeNK_CD56n_CD16p) - (CelltypeB+CelltypeGr+CelltypeT+CelltypeNK_CD56p_CD16n)/4
CelltypeT -0.25
CelltypeB -0.25
CelltypeGr -0.25
CelltypeNK_CD56p_CD16n -0.25
CelltypeNK_CD56n_CD16p 1.00
Months12 0.00
Months17 0.00
Months27 0.00
Months36 0.00
Months42 0.00monkeyHSPC_merged_top@elementMetadata$testingBarcode@metadata$design %>% data.table(keep.rownames = T)9.2 F4H: stacked proportion
plot_validation sourced from “F4_validation.R”
f4h <- (plot_validation(monkeyHSPC_top10each_79) +
theme( plot.title = element_text(hjust = 0.5, vjust = 1)) + labs(title = "Based on top10each")) +
(plot_validation(monkeyHSPC_merged_top) +
theme( plot.title = element_text(hjust = 0.5, vjust = 1)) + labs(title = "Based on barbieQ"))
f4hWarning: Removed 5 rows containing missing values or values outside the scale range
(`geom_bar()`).Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_tile()`).Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_bar()`).Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_tile()`).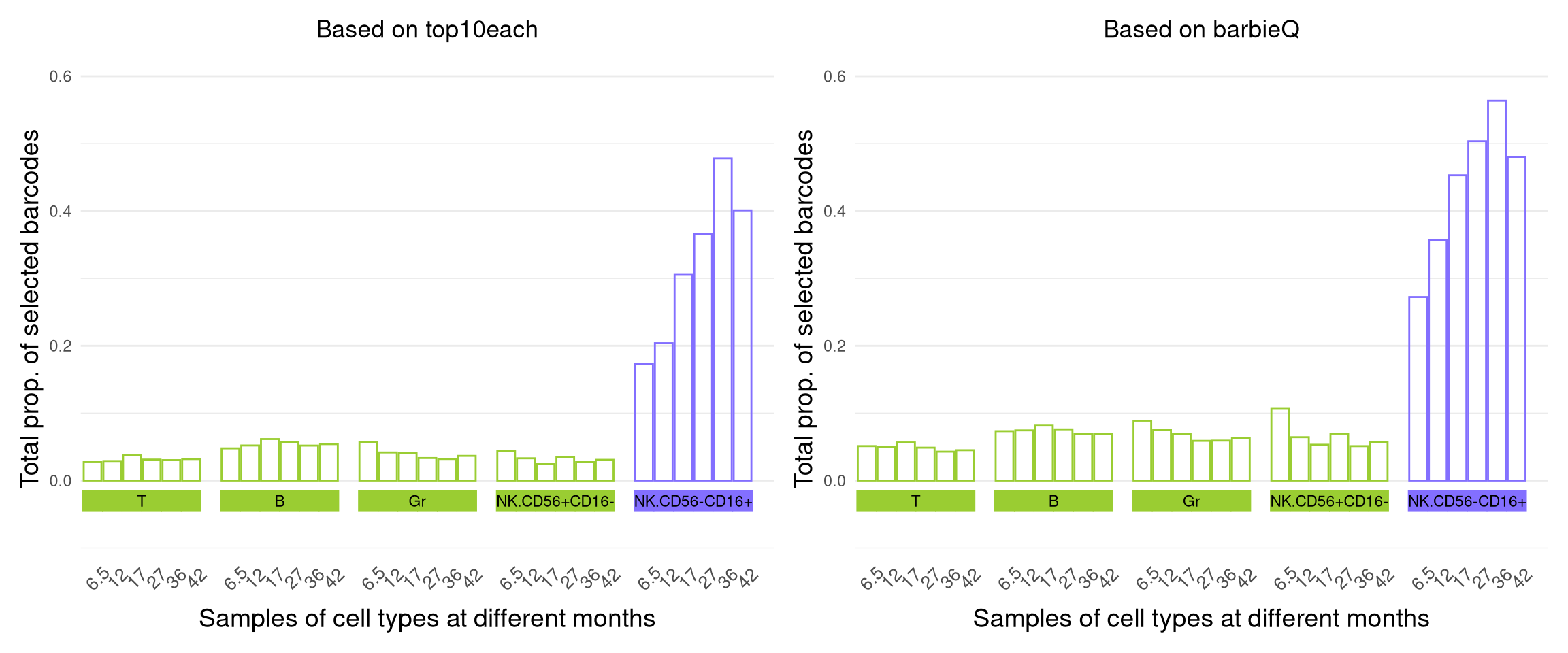
9.3 overlap
- compare sets of significant barcodes
## signif. BC
filter_venn_signif_list <- list(
## signif. BC
top10each = rownames(monkeyHSPC_top10each_79)[monkeyHSPC_top10each_79@elementMetadata$testingBarcode$tendencyTo == "CelltypeNK_CD56n_CD16p"],
barbieQ = rownames(monkeyHSPC_merged_top)[monkeyHSPC_merged_top@elementMetadata$testingBarcode$tendencyTo == "CelltypeNK_CD56n_CD16p"],
## all BC in test
top10each_all = rownames(monkeyHSPC_top10each_79),
barbieQ_filter_all = rownames(monkeyHSPC_merged_top))
venn_filter_signif <- eulerr::euler(filter_venn_signif_list[1:2]) %>%
plot(fills = c("skyblue", "pink"),
alpha = 0.5, labels = list(font = 4, cex = 1), quantities = list(font = 1, cex = 1))
venn_filter_signif %>% plot() %>% grid.grabExpr() %>% wrap_plots()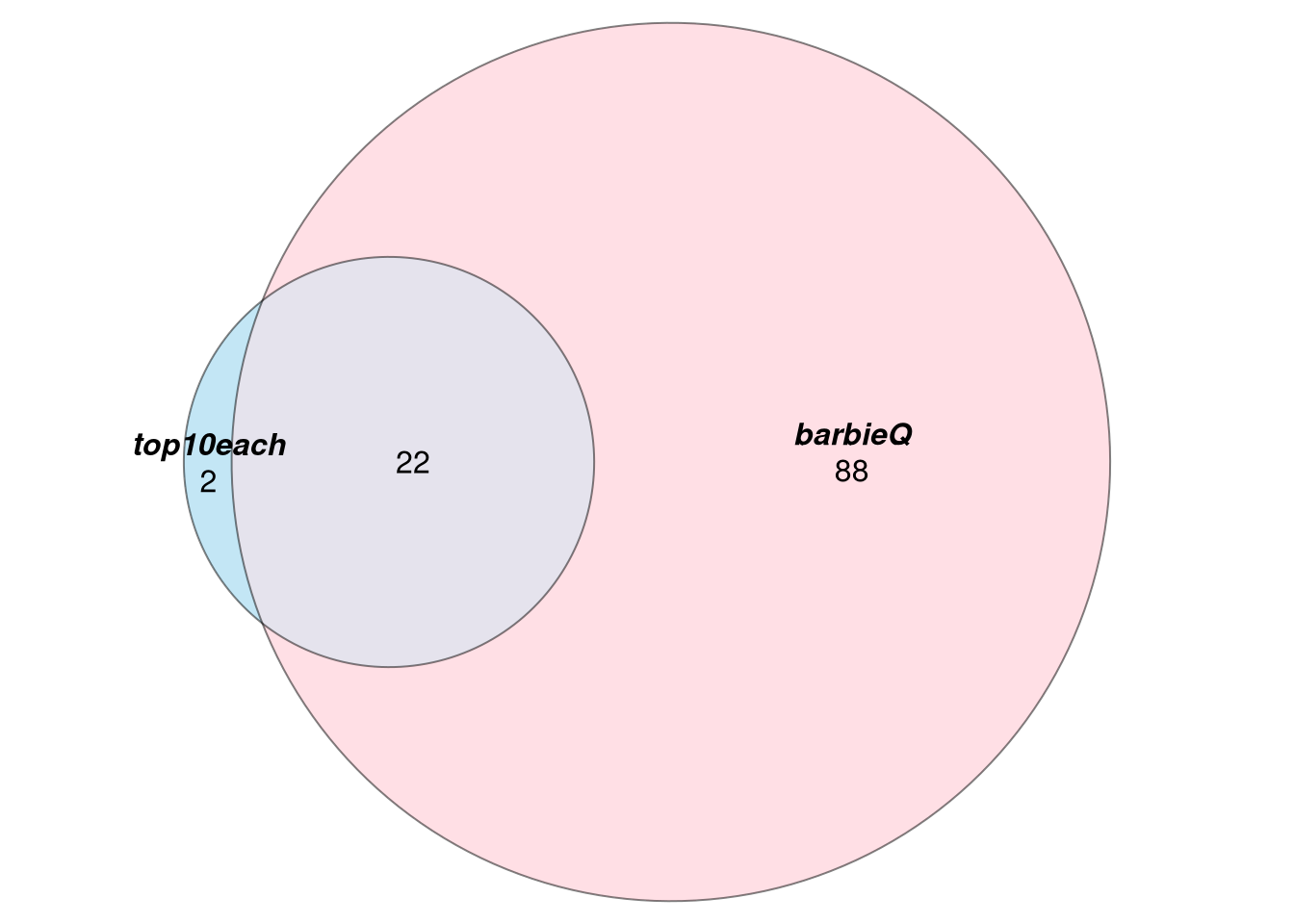
| Version | Author | Date |
|---|---|---|
| dfa737a | FeiLiyang | 2025-12-17 |
9.4 F4I: inspect two sets: boxplot
- box plot on two sets of significant barcodes based on two filtering appraoches
BC_filter_sig <- filter_venn_signif_list[1:2] %>% unlist %>% unique()
FC_filter_sig <- calc_bias_LFC(barbieQ = monkeyHSPC_merged[BC_filter_sig,], get_FC = T) %>%
as.data.frame()
Mean_filter_sig <- calc_bias_LFC(barbieQ = monkeyHSPC_merged[BC_filter_sig,], get_AMean = T) %>%
as.data.frame()
rownames(FC_filter_sig) <- BC_filter_sig
## annotating BC by barbieQ seletion
FC_filter_sig$top10each = 0
FC_filter_sig$bbq = 0
## annotating BC by heuristic selection, `filter_venn_signif_list` obtained from last chunk
FC_filter_sig[filter_venn_signif_list[[1]], "top10each"] = 1
FC_filter_sig[filter_venn_signif_list[[2]], "bbq"] = 1
## adding BC name
FC_filter_sig <- FC_filter_sig %>%
mutate(Barcode = BC_filter_sig) %>%
## summarizing two selecting methods overlaps
mutate(FilteringMethod = ifelse(top10each == 1,
ifelse(bbq == 1, "both", "top10each"),
ifelse(bbq == 1, "barbieQ", "neither"))) %>%
dplyr::select(-c(top10each, bbq))
## extract FC from testing
long_FC_filter_sig <- FC_filter_sig %>%
mutate(Barcode = BC_filter_sig) %>%
pivot_longer(
cols = -c(Barcode, FilteringMethod), # keep Celltype and Months fixed
names_to = "Month",
values_to = "FC"
)
# Add barcode column to Mean_filter_sig
Mean_filter_sig <- Mean_filter_sig %>%
mutate(Barcode = BC_filter_sig)
long_Mean <- Mean_filter_sig %>%
pivot_longer(
cols = -Barcode,
names_to = "Month",
values_to = "Mean"
)
long_FC_filter_sig <- long_FC_filter_sig %>%
left_join(long_Mean, by = c("Barcode", "Month"))barcode_order_filter_sig <- long_FC_filter_sig %>%
group_by(Barcode) %>%
summarise(AMean = mean(Mean, na.rm = TRUE)) %>%
arrange(AMean) %>%
pull(Barcode)
barcode_order_df_filter_sig <- long_FC_filter_sig %>%
group_by(Barcode, FilteringMethod) %>%
summarise(AMean = mean(Mean, na.rm = TRUE)) %>%
arrange(AMean)`summarise()` has grouped output by 'Barcode'. You can override using the
`.groups` argument.p_AMean_0 <- ggplot()+
geom_point(data = barcode_order_df_filter_sig,
aes(x = factor(Barcode, levels = barcode_order_filter_sig),
y = log10(AMean),
color = FilteringMethod),
size = 1.5, shape = 15,
inherit.aes = FALSE) +
theme_classic() +
theme(axis.text.x = element_blank(), legend.position = "none", axis.title = element_text(size = 14)) +
labs(x = "Barocdes ordered by mean proportion across cell types across time",
y = "Log10 prop.") +
scale_color_manual(values = c("both" = "slateblue", "barbieQ" = "hotpink", "top10each" = "steelblue1"))
p_box_0 <- ggplot(long_FC_filter_sig, aes(x = factor(Barcode, levels = barcode_order_filter_sig), y = log2(FC), group = (Barcode), color = FilteringMethod)) +
geom_boxplot(varwidth = T) +
theme_classic() +
theme(axis.text.x = element_blank(), legend.position = "right", axis.title = element_text(size = 14)) +
geom_abline(slope = 0, intercept = 0, linetype = 2, colour = "grey") +
labs(x = "",
y = "LFC",
title = "Barcodes significant in barbieQ test, selected by either barbieQ or top10each filtering") +
scale_color_manual(values = c("both" = "slateblue", "barbieQ" = "hotpink", "top10each" = "steelblue1"))
f4i <- p_box_0 / p_AMean_0 + plot_layout(heights = c(3, 1))
f4i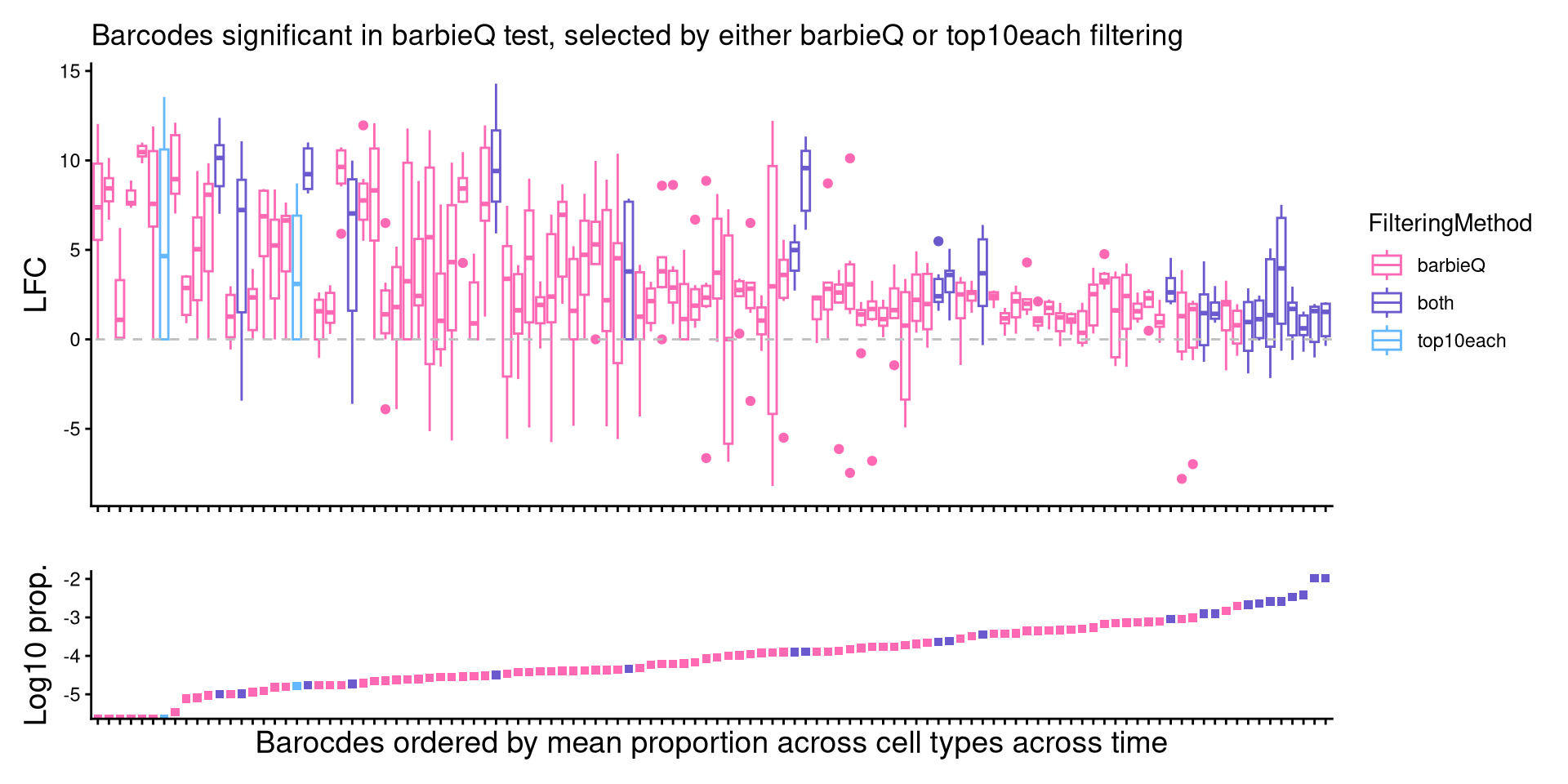
10 F4J: Example difference over time
designs <- monkeyHSPC_top10each_79$sampleMetadata %>% as.data.frame() %>%
with(model.matrix(~0+ Celltype+ Celltype:Months))
designs <- designs[,c(1:5, 9:13)]
colnames(designs) <- gsub(":", ".", colnames(designs))
monkeyHSPC_top10each_79_NK <- testBarcodeSignif(
barbieQ = monkeyHSPC_top10each_79,
sampleMetadata = designs, designMatrix = designs, sampleGroup = "CelltypeNK_CD56n_CD16p.Months",
method = "diffProp", transformation = "asin-sqrt"
)setting CelltypeNK_CD56n_CD16p.Months as the primary factor in `sampleMetadata`.setting up contrastFormula: CelltypeNK_CD56n_CD16p.Monthsno block specified, so there are no duplicate measurements.monkeyHSPC_top10each_79_NK@elementMetadata$testingBarcode$tendencyTo %>% table().
CelltypeNK_CD56n_CD16p.Monthsdowntrend CelltypeNK_CD56n_CD16p.Monthsuptrend
16 16
n.s.
47 monkeyHSPC_top10each_79_NK@elementMetadata$testingBarcode@metadata$design %>% data.table()monkeyHSPC_top10each_79_NK@elementMetadata$testingBarcode@metadata$contrasts Contrasts
Levels CelltypeNK_CD56n_CD16p.Months
CelltypeT 0
CelltypeB 0
CelltypeGr 0
CelltypeNK_CD56p_CD16n 0
CelltypeNK_CD56n_CD16p 0
CelltypeT.Months 0
CelltypeB.Months 0
CelltypeGr.Months 0
CelltypeNK_CD56p_CD16n.Months 0
CelltypeNK_CD56n_CD16p.Months 1monkeyHSPC_top10each_79_NK@elementMetadata$testingBarcode@metadata$contrastGroups levelLow
"CelltypeNK_CD56n_CD16p.Monthsdowntrend"
levelHigh
"CelltypeNK_CD56n_CD16p.Monthsuptrend" NKnp_fig <- monkeyHSPC_top10each_79_NK[,monkeyHSPC_top10each_79_NK$sampleMetadata$Celltype == "NK_CD56n_CD16p"]BC_kinetic <- NKnp_fig@elementMetadata$testingBarcode$tendencyTo
pch_fun <- ifelse(NKnp_fig@elementMetadata$testingBarcode$direction != 0, "*", NA)
kinetic_ha <- rowAnnotation(
Kinetic = anno_simple(
BC_kinetic,
col = NKnp_fig@metadata$factorColors$testingBarcode,
pch = pch_fun
),
show_annotation_name = TRUE,
annotation_name_side = "top",
annotation_name_rot = 45
)
HP_anno_kinetic <- Heatmap(
asin(sqrt(NKnp_fig@assays@data$proportion)), name = "asin-sqrt prop.",
show_row_names = FALSE, column_labels = NKnp_fig$sampleMetadata$Months,
column_title = "Months", column_names_side = "top", column_names_rot = 45,
row_title = "Barcodes", row_km = 4,
width = unit(4, "cm"), height = unit(12, "cm"),
column_order = order(NKnp_fig$sampleMetadata$Months),
# col = colorRamp2(c(0, -2, -4.602), c("red", "white", "blue")),
# left_annotation = row_ha,
right_annotation = kinetic_ha
)The automatically generated colors map from the 1^st and 99^th of the
values in the matrix. There are outliers in the matrix whose patterns
might be hidden by this color mapping. You can manually set the color
to `col` argument.
Use `suppressMessages()` to turn off this message.lgd_sig = Legend(pch = "*", type = "points", labels = "signif.")
lgd_kinetic = Legend(labels = c("up", "down", "n.s."), title = "Kinetic",
legend_gp = gpar(fill = c("#FF5959", "#33AAFF", "#FFC000"))
)
## heatmap_legend_side = "top", annotation_legend_side = "top"
f4j <- draw(HP_anno_kinetic, annotation_legend_list = list(lgd_kinetic, lgd_sig),
annotation_legend_side = "bottom", heatmap_legend_side = "bottom") %>%
grid.grabExpr() %>% list() %>% wrap_plots()
f4j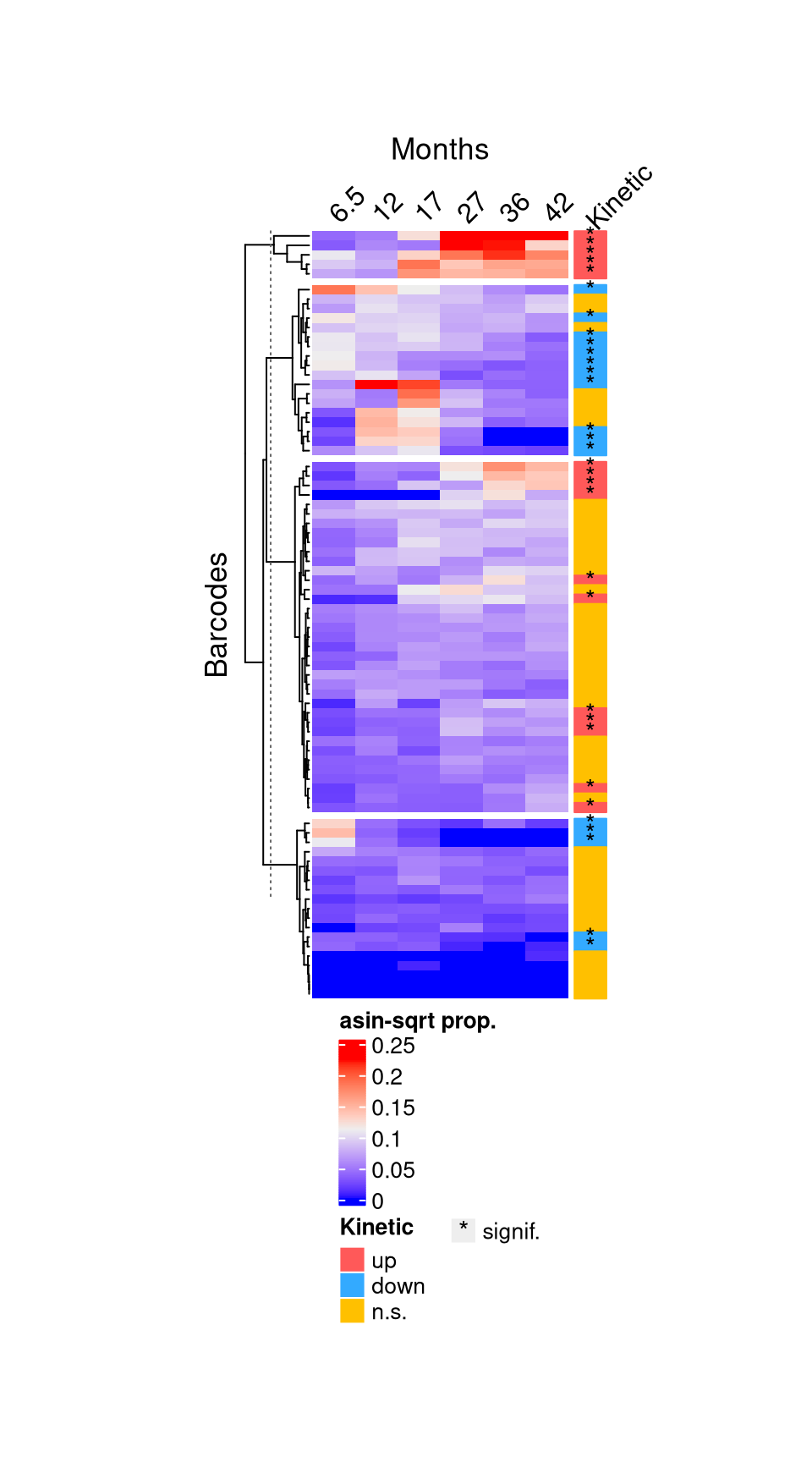
| Version | Author | Date |
|---|---|---|
| dfa737a | FeiLiyang | 2025-12-17 |
11 Figure 4
layout = "
AAAAAAAAAAAAA##
AAAAAAAAAAAAA##
BBBBBBCCCCEEEEE
BBBBBBCCCCEEEEE
BBBBBBCCCCEEEEE
DDDDDDDDDDEEEEE
DDDDDDDDDDEEEEE
DDDDDDDDDDEEEEE
FFFFFFFFFFGGGG#
FFFFFFFFFFGGGG#
FFFFFFFFFFGGGG#
HHHHHHHHHHJJJJ#
HHHHHHHHHHJJJJ#
HHHHHHHHHHJJJJ#
HHHHHHHHHHJJJJ#
IIIIIIIIIIJJJJ#
IIIIIIIIIIJJJJ#
IIIIIIIIIIJJJJ#
"
f4 <- (wrap_elements(full = f4a) +
wrap_elements(f4b + theme(plot.margin = unit(rep(0,4), "cm"))) +
wrap_elements(grid.grabExpr(print(f4c))) +
wrap_elements(f4d + theme(plot.margin = unit(rep(0,4), "cm"))) +
wrap_plots(list(f4e %>% draw() %>% grid.grabExpr())) +
wrap_elements(f4f + theme(plot.margin = unit(rep(0,4), "cm"))) +
wrap_elements(grid.grabExpr(print(f4g))) +
wrap_elements(f4h + theme(plot.margin = unit(c(0,0,0,0), "cm"))) +
wrap_elements(f4i + theme(plot.margin = unit(rep(0,4), "cm"))) +
# wrap_elements(f4j + theme(plot.margin = unit(rep(0,4), "cm")))
wrap_plots(list(HP_anno_kinetic %>%
draw(annotation_legend_list = list(lgd_kinetic, lgd_sig),
annotation_legend_side = "top",heatmap_legend_side = "top") %>%
grid.grabExpr()))
)+
plot_layout(design = layout) +
plot_annotation(tag_levels = list(c("A","B","C", "D", "E","F","G","H", "I", "J"))) &
theme(
plot.tag = element_text(size = 24, face = "bold", family = "arial"),
axis.title = element_text(size = 20),
axis.text = element_text(size = 14),
legend.title = element_text(size = 13),
legend.text = element_text(size = 11))
f4Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_bar()`).Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_tile()`).Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_bar()`).Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_tile()`).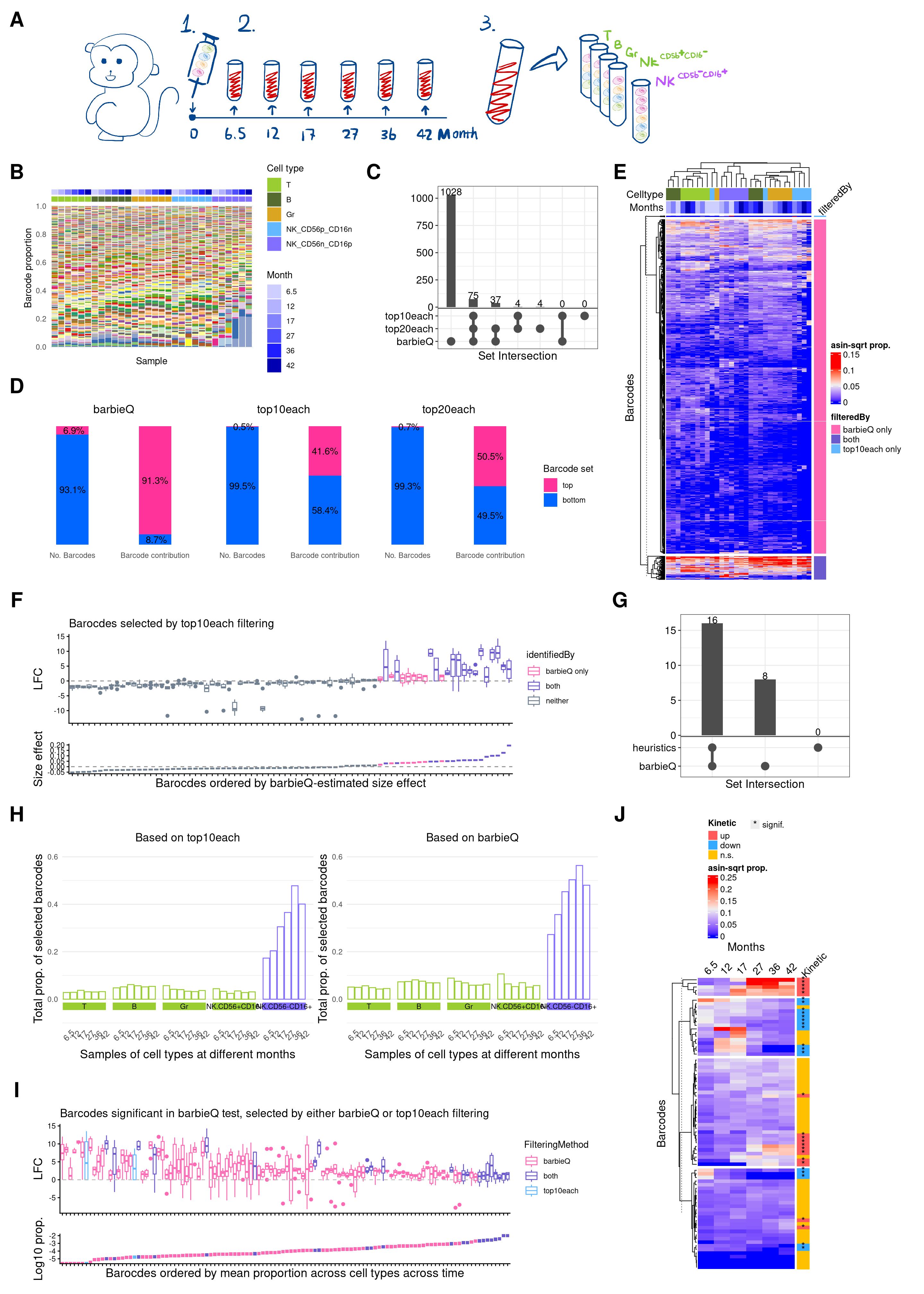
| Version | Author | Date |
|---|---|---|
| dfa737a | FeiLiyang | 2025-12-17 |
ggsave(
filename = "output/f4.png",
plot = f4,
width = 15,
height = 21,
units = "in", # for Rmd r chunk fig size, unit default to inch
dpi = 350
)Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_bar()`).Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_tile()`).Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_bar()`).Warning: Removed 5 rows containing missing values or values outside the scale range
(`geom_tile()`).Saving this figure in f4
sessionInfo()R version 4.5.0 (2025-04-11)
Platform: x86_64-pc-linux-gnu
Running under: Red Hat Enterprise Linux 9.6 (Plow)
Matrix products: default
BLAS/LAPACK: FlexiBLAS OPENBLAS-OPENMP; LAPACK version 3.9.0
locale:
[1] LC_CTYPE=en_AU.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_AU.UTF-8 LC_COLLATE=en_AU.UTF-8
[5] LC_MONETARY=en_AU.UTF-8 LC_MESSAGES=en_AU.UTF-8
[7] LC_PAPER=en_AU.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_AU.UTF-8 LC_IDENTIFICATION=C
time zone: Australia/Melbourne
tzcode source: system (glibc)
attached base packages:
[1] grid stats4 stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] barbieQ_1.1.3 edgeR_4.6.3
[3] limma_3.64.3 eulerr_7.0.4
[5] magick_2.9.0 ComplexHeatmap_2.24.1
[7] ggVennDiagram_1.5.4 scales_1.4.0
[9] patchwork_1.3.2 ggnewscale_0.5.2
[11] ggbreak_0.1.6 ggplot2_4.0.0
[13] SummarizedExperiment_1.38.1 Biobase_2.68.0
[15] GenomicRanges_1.60.0 GenomeInfoDb_1.44.3
[17] IRanges_2.42.0 S4Vectors_0.48.0
[19] BiocGenerics_0.54.0 generics_0.1.4
[21] MatrixGenerics_1.20.0 matrixStats_1.5.0
[23] data.table_1.17.8 knitr_1.50
[25] tibble_3.3.0 tidyr_1.3.1
[27] dplyr_1.1.4 magrittr_2.0.4
[29] readxl_1.4.5 workflowr_1.7.2
loaded via a namespace (and not attached):
[1] RColorBrewer_1.1-3 rstudioapi_0.17.1 jsonlite_2.0.0
[4] shape_1.4.6.1 jomo_2.7-6 nloptr_2.2.1
[7] farver_2.1.2 logistf_1.26.1 rmarkdown_2.30
[10] ragg_1.5.0 GlobalOptions_0.1.2 fs_1.6.6
[13] vctrs_0.6.5 minqa_1.2.8 memoise_2.0.1
[16] forcats_1.0.1 htmltools_0.5.8.1 S4Arrays_1.8.1
[19] broom_1.0.10 cellranger_1.1.0 SparseArray_1.8.1
[22] gridGraphics_0.5-1 mitml_0.4-5 sass_0.4.10
[25] bslib_0.9.0 cachem_1.1.0 whisker_0.4.1
[28] igraph_2.1.4 lifecycle_1.0.4 iterators_1.0.14
[31] pkgconfig_2.0.3 Matrix_1.7-3 R6_2.6.1
[34] fastmap_1.2.0 rbibutils_2.3 GenomeInfoDbData_1.2.14
[37] clue_0.3-66 digest_0.6.37 aplot_0.2.9
[40] colorspace_2.1-2 ps_1.9.1 rprojroot_2.1.1
[43] textshaping_1.0.3 labeling_0.4.3 httr_1.4.7
[46] polyclip_1.10-7 abind_1.4-8 mgcv_1.9-1
[49] compiler_4.5.0 withr_3.0.2 doParallel_1.0.17
[52] backports_1.5.0 S7_0.2.0 viridis_0.6.5
[55] ggforce_0.5.0 pan_1.9 MASS_7.3-65
[58] rappdirs_0.3.3 DelayedArray_0.34.1 rjson_0.2.23
[61] tools_4.5.0 httpuv_1.6.16 nnet_7.3-20
[64] glue_1.8.0 callr_3.7.6 nlme_3.1-168
[67] promises_1.3.3 polylabelr_0.3.0 getPass_0.2-4
[70] cluster_2.1.8.1 operator.tools_1.6.3 gtable_0.3.6
[73] formula.tools_1.7.1 tidygraph_1.3.1 XVector_0.48.0
[76] ggrepel_0.9.6 foreach_1.5.2 pillar_1.11.1
[79] stringr_1.5.2 yulab.utils_0.2.1 later_1.4.4
[82] circlize_0.4.16 splines_4.5.0 tweenr_2.0.3
[85] lattice_0.22-6 survival_3.8-3 tidyselect_1.2.1
[88] locfit_1.5-9.12 git2r_0.36.2 reformulas_0.4.1
[91] gridExtra_2.3 xfun_0.53 graphlayouts_1.2.2
[94] statmod_1.5.0 stringi_1.8.7 UCSC.utils_1.4.0
[97] boot_1.3-31 ggfun_0.2.0 yaml_2.3.10
[100] evaluate_1.0.5 codetools_0.2-20 ggraph_2.2.2
[103] ggplotify_0.1.3 cli_3.6.5 rpart_4.1.24
[106] systemfonts_1.3.1 Rdpack_2.6.4 processx_3.8.6
[109] jquerylib_0.1.4 Rcpp_1.1.0 png_0.1-8
[112] parallel_4.5.0 lme4_1.1-37 glmnet_4.1-10
[115] viridisLite_0.4.2 purrr_1.1.0 crayon_1.5.3
[118] GetoptLong_1.0.5 rlang_1.1.6 mice_3.18.0