barbieQ_paper Supplementary 1:
Preprocessing HSPC xenograft data
Liyang Fei
Initiated: 2025-04-03
Rendered: 2026-01-06
Last updated: 2026-01-06
Checks: 5 2
Knit directory: public_barcode_count/
This reproducible R Markdown analysis was created with workflowr (version 1.7.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown is untracked by Git. To know which version of the R
Markdown file created these results, you’ll want to first commit it to
the Git repo. If you’re still working on the analysis, you can ignore
this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
The global environment had objects present when the code in the R
Markdown file was run. These objects can affect the analysis in your R
Markdown file in unknown ways. For reproduciblity it’s best to always
run the code in an empty environment. Use wflow_publish or
wflow_build to ensure that the code is always run in an
empty environment.
The following objects were defined in the global environment when these results were created:
| Name | Class | Size |
|---|---|---|
| module | function | 5.6 Kb |
The command set.seed(20250112) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 34f5894. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .RData
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: public_barcode_count.Rproj
Untracked files:
Untracked: analysis/barbieQ_paper_FigureS1_AML.Rmd
Untracked: analysis/barbieQ_paper_FigureS1_Mixture.Rmd
Untracked: analysis/barbieQ_paper_FigureS1_xenoHSPC.Rmd
Untracked: analysis/figure/
Untracked: output/AML_barbieQ.rda
Untracked: output/fs1_aml.png
Untracked: output/fs1_mixture.png
Untracked: output/fs1_xeno.png
Untracked: output/fs1a_knowncluster.png
Untracked: output/xenoHSPC_barbieQ.rda
Unstaged changes:
Modified: analysis/barbieQ_paper_Figure2.Rmd
Deleted: analysis/barbieQ_paper_S1.Rmd
Modified: analysis/index.Rmd
Modified: data/BelderbosME/README.md
Modified: output/f2.png
Modified: output/monkeyHSPC_barbieQ.rda
Modified: output/monkeyHSPC_raw_barbieQ.rda
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
There are no past versions. Publish this analysis with
wflow_publish() to start tracking its development.
Links to preprocessing other datasets in the barbieQ paper:
1 Load Dependencies
library(readxl)
library(magrittr)
library(dplyr)
library(tidyr) # for pivot_longer
library(tibble) # for rownames_to _column
library(knitr) # for kable()
library(ggplot2)
library(patchwork)
library(scales)
library(ggVennDiagram)
library(ComplexHeatmap)
library(limma)
library(edgeR)
library(SummarizedExperiment)
library(SEtools)
library(S4Vectors)
library(devtools)
source("analysis/plotBarcodeHistogram.R") ## accommodated from bartools::plotBarcodehistogram
source("analysis/ggplot_theme.R") ## setting ggplot theme2 Install
barbieQ
Installing the latest devel version of barbieQ from
GitHub.
if (!requireNamespace("barbieQ", quietly = TRUE)) {
remotes::install_github("Oshlack/barbieQ")
}Warning: replacing previous import 'data.table::first' by 'dplyr::first' when
loading 'barbieQ'Warning: replacing previous import 'data.table::last' by 'dplyr::last' when
loading 'barbieQ'Warning: replacing previous import 'data.table::between' by 'dplyr::between'
when loading 'barbieQ'Warning: replacing previous import 'dplyr::as_data_frame' by
'igraph::as_data_frame' when loading 'barbieQ'Warning: replacing previous import 'dplyr::groups' by 'igraph::groups' when
loading 'barbieQ'Warning: replacing previous import 'dplyr::union' by 'igraph::union' when
loading 'barbieQ'Registered S3 method overwritten by 'formula.tools':
method from
as.character.formula openxlsxlibrary(barbieQ)Check the version of barbieQ.
packageVersion("barbieQ")[1] '1.1.3'3 Set seeds
set.seed(2025)4 Read data
Read every sheet in the excel.
8 mice: “C1” “C2” “C3” “C10” “C15” “C22” “C21” “C23”
overlapping barcodes: hundreds
mouse-specific barcodes: 1-2k.
According to the original publication, ‘library’ in columns may refer to lentiviral level barcode library - before integrated into cell genome.
sheets <- readxl::excel_sheets("data/BelderbosME/mmc2.xlsx")
data_list <- purrr::map(sheets, ~readxl::read_excel("data/BelderbosME/mmc2.xlsx", sheet = .x))New names:
New names:
New names:
• `` -> `...39`
• `` -> `...40`# bind sheets
# library(dplyr)
# combined_data <- bind_rows(data_list, .id = "sheet_name")
names(data_list) <- sheets
C1_bar <- data_list$C1$ID
C2_bar <- data_list$C2$`['ID'`
C3_bar <- data_list$C3$`['ID'`
C10_bar <- data_list$C10$ID
C15_bar <- data_list$C15$...1
C22_bar <- data_list$C22$ID
C21_bar <- data_list$C21$ID
C23_bar <- data_list$C23$ID
bar_list <- list(
C1 = C1_bar,
C2 = C2_bar,
C3 = C3_bar,
C10 = C10_bar,
C15 = C15_bar,
C22 = C22_bar,
C21 = C21_bar,
C23 = C23_bar
)
ggVennDiagram::ggVennDiagram(x = bar_list)Warning in ggVennDiagram::ggVennDiagram(x = bar_list): Only support 2-7
dimension Venn diagram. Will give a plain upset plot instead.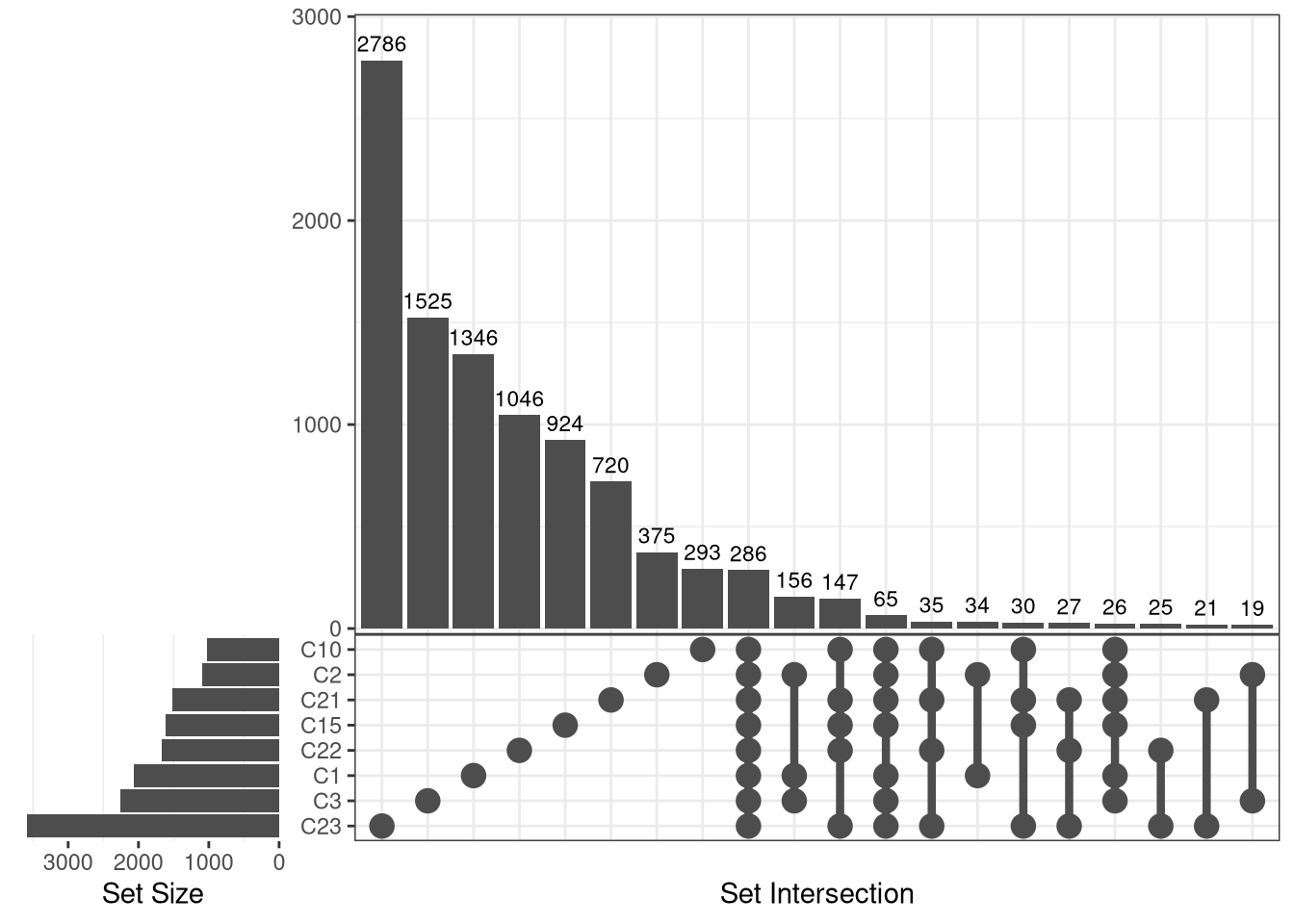
5 Clean up data
5.1 C1
class(data_list$C1)[1] "tbl_df" "tbl" "data.frame"data_list$C1data_list$C1 <- as.data.frame(data_list$C1)
class(data_list$C1)[1] "data.frame"# Subset the dataframe to exclude rows where the first column (barcodeID) is NA
any(is.na(data_list$C1[, 1]))[1] TRUEdata_list$C1 <- data_list$C1[!is.na(data_list$C1[, 1]), ]
dim(data_list$C1)[1] 2062 43# set up the first column of each mat as rownames (barcode ID)
rownames(data_list$C1) <- data_list$C1[,1]
data_list$C1 <- data_list$C1[,-1]
# delete empty or processed columns
del <- c("...39", "...40", "Present", "Total", "Log_total")
data_list$C1 <- data_list$C1[, !colnames(data_list$C1) %in% del]
# delete processed "sum" columns
data_list$C1 <- data_list$C1[, !grepl("_Sum", colnames(data_list$C1))]
data_list$C1 <- data_list$C1[, !grepl("_sum", colnames(data_list$C1))]
# delete processed "shannon" rows
rownames(data_list$C1) [!grepl("^[ATGC]+$", rownames(data_list$C1))][1] "TTGTGAAAGACGGCCAGTGACATCCCCCGGGCTGCACG\""
[2] "TAGTGCTAGACGGCCAGTGAGATCCCCCGGGCTGCACG\""
[3] "TTGTGTAAGACGGCCAGTGAGATCCCCCGGGCTGCACG\""
[4] "CCGTGGATGACGGCCAGTGCGATCCCCCAGGCTGCACG\""
[5] "Present"
[6] "Total"
[7] "Shannon index"
[8] "Shannon equitability"
[9] "Shannon count" rownames(data_list$C1) [!grepl("^[ATGC\\\"]+$", rownames(data_list$C1))][1] "Present" "Total" "Shannon index"
[4] "Shannon equitability" "Shannon count" data_list$C1 <- data_list$C1[grepl("^[ATGC\\\"]+$", rownames(data_list$C1)),]
dim(data_list$C1)[1] 2057 33# Convert NA values to 0 in the matrix
data_list$C1[is.na(data_list$C1)] <- 0
!any(is.na(data_list$C1))[1] TRUE# Convert character data to numeric
C1_count_numeric <- apply(data_list$C1, 2, as.numeric)
rownames(C1_count_numeric) <- rownames(data_list$C1)
data_list$C1 <- C1_count_numeric
# extract condition from colnames. I don;t know what GK / L means.
xCondition <- gsub("(.*)\\_.*", "\\1", colnames(data_list$C1))
xCondition <- gsub("(.*)\\_.*", "\\1", xCondition)
xCondition [1] "library" "GK" "GK" "GK" "GK" "GK" "GK"
[8] "GK" "L" "L" "L" "L" "L" "L"
[15] "GK" "GK" "GK" "GK" "GK" "GK" "L"
[22] "L" "L" "L" "L" "L" "L" "GK"
[29] "GK" "GK" "L" "L" "L" # extract time from colnames
Time <- gsub(".*\\_(wk[0-9]{1,}).*", "\\1", colnames(data_list$C1))
Time[grepl("\\_", Time)] <- "sac"
Time [1] "library" "wk6" "wk12" "wk19" "wk30" "wk37" "wk43"
[8] "sac" "wk6" "wk12" "wk19" "wk25" "wk30" "wk43"
[15] "wk25" "wk25" "wk25" "wk37" "wk37" "wk37" "wk25"
[22] "wk25" "wk25" "wk37" "wk37" "wk37" "sac" "sac"
[29] "sac" "sac" "sac" "sac" "sac" # extract tissue from colnames
Tissue <- gsub(".*\\_(BM)\\_.*", "\\1", colnames(data_list$C1))
Tissue <- gsub(".*\\_(Blood)\\_.*", "\\1", Tissue)
Tissue[grepl("\\_", Tissue)] <- "Blood"
Tissue [1] "library" "Blood" "Blood" "Blood" "Blood" "Blood" "Blood"
[8] "Blood" "Blood" "Blood" "Blood" "Blood" "Blood" "Blood"
[15] "Blood" "Blood" "Blood" "Blood" "Blood" "Blood" "Blood"
[22] "Blood" "Blood" "Blood" "Blood" "Blood" "Blood" "BM"
[29] "BM" "BM" "BM" "BM" "BM" # extract celltype from colnames
Celltype <- gsub(".*\\_.*\\_([A-Z])", "\\1", colnames(data_list$C1))
Celltype[grepl("\\_", Celltype)] <- "Unsorted"
Celltype[grepl("T", Celltype)] <- "Tcell"
Celltype[grepl("B", Celltype)] <- "Bcell"
Celltype[grepl("G", Celltype)] <- "Grn"
colnames(data_list$C1) [1] "library" "GK_wk6" "GK_wk12" "GK_wk19" "GK_wk30"
[6] "GK_wk37" "GK_wk43" "GK_sac" "L_wk6" "L_wk12"
[11] "L_wk19" "L_wk25" "L_wk30" "L_wk43" "GK_wk25_B"
[16] "GK_wk25_T" "GK_wk25_G" "GK_wk37_B" "GK_wk37_T" "GK_wk37_G"
[21] "L_wk25_B" "L_wk25_T" "L_wk25_G" "L_wk37_B" "L_wk37_T"
[26] "L_wk37_G" "L_Blood_sac" "GK_BM_B" "GK_BM_G" "GK_BM_T"
[31] "L_BM_B" "L_BM_G" "L_BM_T" C1_meta <- data.frame(
Sample = colnames(data_list$C1),
Donor = rep("C1", ncol(data_list$C1)),
xCondition = xCondition,
Time = Time,
Tissue = Tissue,
Celltype = Celltype
)
# View(data_list$C1)5.2 C2
# data_list$C2 %>% View
data_list$C2 <- as.data.frame(data_list$C2)
# Subset the dataframe to exclude rows where the first column (barcodeID) is NA
any(is.na(data_list$C2[, 1]))[1] FALSEdata_list$C2 <- data_list$C2[!is.na(data_list$C2[, 1]), ]
dim(data_list$C2)[1] 1090 43# set up the first column of each mat as rownames (barcode ID)
rownames(data_list$C2) <- data_list$C2[,1]
data_list$C2 <- data_list$C2[,-1]
colnames(data_list$C2) [1] "\"['library'\"" "GK_wk7" "GK_wk12"
[4] "GK_wk18" "GK_wk25" "GK_wk30"
[7] "GK_Liver" "GK_Spleen" "GK_BMFront"
[10] "GK_BMLeft" "GK_BMRight" "GK_BMSpine"
[13] "L_wk7" "L_wk12" "L_wk18"
[16] "L_wk24" "L_wk30" "L_wk36Sum"
[19] "L_BloodSacSum" "L_BMBcells" "L_BMGrn"
[22] "L_BMTcells" "L_BMOther" "L_BM_Sum"
[25] "R_wk7" "R_wk12" "R_wk18"
[28] "R_wk24" "R_wk30" "R_wk36_Bcells"
[31] "L_BloodSac" "R_BMBcells" "R_BMGrn"
[34] "R_BMOther" "R_BM_Sum" "L_wk36Bcells"
[37] "L_wk36_Grn" "L_wk36_Tcells" "L_BloodSac_Bcells"
[40] "L_BloodSac_Grn" "L_BloodSac_Tcells" "L_BloodSac_Other" # delete empty or processed columns
del <- c("...39", "...40", "Present", "Total", "Log_total")
data_list$C2 <- data_list$C2[, !colnames(data_list$C2) %in% del]
# delete processed "sum" columns
data_list$C2 <- data_list$C2[, !grepl("Sum", colnames(data_list$C2))]
data_list$C2 <- data_list$C2[, !grepl("sum", colnames(data_list$C2))]
# delete processed "shannon" rows
rownames(data_list$C2) [!grepl("^[ATGC]+$", rownames(data_list$C2))][1] "TTGTGAAAGACGGCCAGTGACATCCCCCGGGCTGCACG\""
[2] "TAGTGCTAGACGGCCAGTGAGATCCCCCGGGCTGCACG\""
[3] "TTGTGTAAGACGGCCAGTGAGATCCCCCGGGCTGCACG\""
[4] "CCGTGGATGACGGCCAGTGCGATCCCCCAGGCTGCACG\""rownames(data_list$C2) [!grepl("^[ATGC\\\"]+$", rownames(data_list$C2))]character(0)data_list$C2 <- data_list$C2[grepl("^[ATGC\\\"]+$", rownames(data_list$C2)),]
dim(data_list$C2)[1] 1090 38# Convert NA values to 0 in the matrix
data_list$C2[is.na(data_list$C2)] <- 0
!any(is.na(data_list$C2))[1] TRUE# Convert character data to numeric
C2_count_numeric <- apply(data_list$C2, 2, as.numeric)
rownames(C2_count_numeric) <- rownames(data_list$C2)
data_list$C2 <- C2_count_numeric
# View sample names
colnames(data_list$C2) [1] "\"['library'\"" "GK_wk7" "GK_wk12"
[4] "GK_wk18" "GK_wk25" "GK_wk30"
[7] "GK_Liver" "GK_Spleen" "GK_BMFront"
[10] "GK_BMLeft" "GK_BMRight" "GK_BMSpine"
[13] "L_wk7" "L_wk12" "L_wk18"
[16] "L_wk24" "L_wk30" "L_BMBcells"
[19] "L_BMGrn" "L_BMTcells" "L_BMOther"
[22] "R_wk7" "R_wk12" "R_wk18"
[25] "R_wk24" "R_wk30" "R_wk36_Bcells"
[28] "L_BloodSac" "R_BMBcells" "R_BMGrn"
[31] "R_BMOther" "L_wk36Bcells" "L_wk36_Grn"
[34] "L_wk36_Tcells" "L_BloodSac_Bcells" "L_BloodSac_Grn"
[37] "L_BloodSac_Tcells" "L_BloodSac_Other" # fix "library" clean any character that is not a letter, digit, or underscore with an empty string, effectively removing unwanted symbols.
colnames(data_list$C2) <- gsub("[^a-zA-Z0-9_]", "", colnames(data_list$C2))
# extract condition from colnames. I don;t know what GK / L means.
xCondition <- gsub("(.*)\\_.*", "\\1", colnames(data_list$C2))
xCondition <- gsub("(.*)\\_.*", "\\1", xCondition)
xCondition [1] "library" "GK" "GK" "GK" "GK" "GK" "GK"
[8] "GK" "GK" "GK" "GK" "GK" "L" "L"
[15] "L" "L" "L" "L" "L" "L" "L"
[22] "R" "R" "R" "R" "R" "R" "L"
[29] "R" "R" "R" "L" "L" "L" "L"
[36] "L" "L" "L" # extract time from colnames
Time <- gsub(".*\\_(wk[0-9]{1,}).*", "\\1", colnames(data_list$C2))
Time[grepl("\\_", Time)] <- "sac"
Time [1] "library" "wk7" "wk12" "wk18" "wk25" "wk30" "sac"
[8] "sac" "sac" "sac" "sac" "sac" "wk7" "wk12"
[15] "wk18" "wk24" "wk30" "sac" "sac" "sac" "sac"
[22] "wk7" "wk12" "wk18" "wk24" "wk30" "wk36" "sac"
[29] "sac" "sac" "sac" "wk36" "wk36" "wk36" "sac"
[36] "sac" "sac" "sac" # extract tissue from colnames
Tissue <- gsub(".*\\_(BM).*", "\\1", colnames(data_list$C2))
Tissue <- gsub(".*\\_(Liver).*", "\\1", Tissue)
Tissue <- gsub(".*\\_(Spleen).*", "\\1", Tissue)
Tissue <- gsub(".*\\_(Blood).*", "\\1", Tissue)
Tissue[grepl("\\_", Tissue)] <- "Blood"
Tissue [1] "library" "Blood" "Blood" "Blood" "Blood" "Blood" "Liver"
[8] "Spleen" "BM" "BM" "BM" "BM" "Blood" "Blood"
[15] "Blood" "Blood" "Blood" "BM" "BM" "BM" "BM"
[22] "Blood" "Blood" "Blood" "Blood" "Blood" "Blood" "Blood"
[29] "BM" "BM" "BM" "Blood" "Blood" "Blood" "Blood"
[36] "Blood" "Blood" "Blood" # extract position, like BMLeft, BMSpine
Position <- vector("character", length(Tissue))
Position[grepl("Left", colnames(data_list$C2))] <- "Left"
Position[grepl("Front", colnames(data_list$C2))] <- "Front"
Position[grepl("Right", colnames(data_list$C2))] <- "Right"
Position[grepl("Spine", colnames(data_list$C2))] <- "Spine"
# extract celltype from colnames
Celltype <- gsub(".*\\_.*\\_([A-Z])", "\\1", colnames(data_list$C2))
Celltype[grepl("Tcell", Celltype)] <- "Tcell"
Celltype[grepl("Bcell", Celltype)] <- "Bcell"
Celltype[grepl("Grn", Celltype)] <- "Grn"
Celltype[grepl("Other", Celltype)] <- "Other"
Celltype[grepl("\\_", Celltype)] <- "Unsorted"
Celltype [1] "library" "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Unsorted"
[7] "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Unsorted"
[13] "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Bcell"
[19] "Grn" "Tcell" "Other" "Unsorted" "Unsorted" "Unsorted"
[25] "Unsorted" "Unsorted" "Bcell" "Unsorted" "Bcell" "Grn"
[31] "Other" "Bcell" "Grn" "Tcell" "Bcell" "Grn"
[37] "Tcell" "Other" colnames(data_list$C2) [1] "library" "GK_wk7" "GK_wk12"
[4] "GK_wk18" "GK_wk25" "GK_wk30"
[7] "GK_Liver" "GK_Spleen" "GK_BMFront"
[10] "GK_BMLeft" "GK_BMRight" "GK_BMSpine"
[13] "L_wk7" "L_wk12" "L_wk18"
[16] "L_wk24" "L_wk30" "L_BMBcells"
[19] "L_BMGrn" "L_BMTcells" "L_BMOther"
[22] "R_wk7" "R_wk12" "R_wk18"
[25] "R_wk24" "R_wk30" "R_wk36_Bcells"
[28] "L_BloodSac" "R_BMBcells" "R_BMGrn"
[31] "R_BMOther" "L_wk36Bcells" "L_wk36_Grn"
[34] "L_wk36_Tcells" "L_BloodSac_Bcells" "L_BloodSac_Grn"
[37] "L_BloodSac_Tcells" "L_BloodSac_Other" C2_meta <- data.frame(
Sample = colnames(data_list$C2),
Donor = rep("C2", ncol(data_list$C2)),
xCondition = xCondition,
Time = Time,
Tissue = Tissue,
Position = Position,
Celltype = Celltype
)
# View(data_list$C2)5.3 C3
There are two GK_BM_T. One might be typo, originally GK_BM_G.
# data_list$C3 %>% View
data_list$C3 <- as.data.frame(data_list$C3)
# Subset the dataframe to exclude rows where the first column (barcodeID) is NA
any(is.na(data_list$C3[, 1]))[1] FALSEdata_list$C3 <- data_list$C3[!is.na(data_list$C3[, 1]), ]
dim(data_list$C3)[1] 2260 25# set up the first column of each mat as rownames (barcode ID)
rownames(data_list$C3) <- data_list$C3[,1]
data_list$C3 <- data_list$C3[,-1]
colnames(data_list$C3) [1] "\"['library'\""
[2] "\" 'TK3 BM Cells before transplant'\""
[3] "GK_wk6"
[4] "GK_wk12"
[5] "GK_wk18"
[6] "GK_wk24"
[7] "GK_wk30"
[8] "GK_wk36_B"
[9] "GK_wk42"
[10] "GK_wk47"
[11] "L_wk6"
[12] "L_wk12"
[13] "L_wk18"
[14] "L_wk24"
[15] "L_wk30"
[16] "L_wk42"
[17] "GK_BM_B"
[18] "GK_BM_T...19"
[19] "GK_BM_T...20"
[20] "GK_BM_Sum"
[21] "L_BM_B"
[22] "L_BM_G"
[23] "L_BM_T"
[24] "L_BM_Sum" # delete empty or processed columns
del <- c("...39", "...40", "Present", "Total", "Log_total")
data_list$C3 <- data_list$C3[, !colnames(data_list$C3) %in% del]
# delete processed "sum" columns
data_list$C3 <- data_list$C3[, !grepl("Sum", colnames(data_list$C3))]
data_list$C3 <- data_list$C3[, !grepl("sum", colnames(data_list$C3))]
# delete processed "shannon" rows
rownames(data_list$C3) [!grepl("^[ATGC]+$", rownames(data_list$C3))][1] "TTGTGAAAGACGGCCAGTGACATCCCCCGGGCTGCACG\""
[2] "TAGTGCTAGACGGCCAGTGAGATCCCCCGGGCTGCACG\""
[3] "TTGTGTAAGACGGCCAGTGAGATCCCCCGGGCTGCACG\""
[4] "CCGTGGATGACGGCCAGTGCGATCCCCCAGGCTGCACG\""rownames(data_list$C3) [!grepl("^[ATGC\\\"]+$", rownames(data_list$C3))]character(0)data_list$C3 <- data_list$C3[grepl("^[ATGC\\\"]+$", rownames(data_list$C3)),]
dim(data_list$C3)[1] 2260 22# Convert NA values to 0 in the matrix
data_list$C3[is.na(data_list$C3)] <- 0
!any(is.na(data_list$C3))[1] TRUE# Convert character data to numeric
C3_count_numeric <- apply(data_list$C3, 2, as.numeric)
rownames(C3_count_numeric) <- rownames(data_list$C3)
data_list$C3 <- C3_count_numeric
# View sample names
colnames(data_list$C3) [1] "\"['library'\""
[2] "\" 'TK3 BM Cells before transplant'\""
[3] "GK_wk6"
[4] "GK_wk12"
[5] "GK_wk18"
[6] "GK_wk24"
[7] "GK_wk30"
[8] "GK_wk36_B"
[9] "GK_wk42"
[10] "GK_wk47"
[11] "L_wk6"
[12] "L_wk12"
[13] "L_wk18"
[14] "L_wk24"
[15] "L_wk30"
[16] "L_wk42"
[17] "GK_BM_B"
[18] "GK_BM_T...19"
[19] "GK_BM_T...20"
[20] "L_BM_B"
[21] "L_BM_G"
[22] "L_BM_T" # fix "library" clean any character that is not a letter, digit, or underscore with an empty string, effectively removing unwanted symbols.
colnames(data_list$C3) <- gsub("[^a-zA-Z0-9_]", "", colnames(data_list$C3))
# extract condition from colnames. I don;t know what GK / L means.
xCondition <- gsub("(.*)\\_.*", "\\1", colnames(data_list$C3))
xCondition <- gsub("(.*)\\_.*", "\\1", xCondition)
xCondition[grepl("TK3BMCellsbeforetransplant", xCondition)] <- "time0"
xCondition [1] "library" "time0" "GK" "GK" "GK" "GK" "GK"
[8] "GK" "GK" "GK" "L" "L" "L" "L"
[15] "L" "L" "GK" "GK" "GK" "L" "L"
[22] "L" # extract time from colnames
Time <- gsub(".*\\_(wk[0-9]{1,}).*", "\\1", colnames(data_list$C3))
Time[grepl("\\_", Time)] <- "sac"
Time[grepl("TK3BMCellsbeforetransplant", Time)] <- "time0"
Time [1] "library" "time0" "wk6" "wk12" "wk18" "wk24" "wk30"
[8] "wk36" "wk42" "wk47" "wk6" "wk12" "wk18" "wk24"
[15] "wk30" "wk42" "sac" "sac" "sac" "sac" "sac"
[22] "sac" # extract tissue from colnames
Tissue <- gsub(".*\\_(BM).*", "\\1", colnames(data_list$C3))
Tissue <- gsub(".*\\_(Liver).*", "\\1", Tissue)
Tissue <- gsub(".*\\_(Spleen).*", "\\1", Tissue)
Tissue <- gsub(".*\\_(Blood).*", "\\1", Tissue)
Tissue[grepl("\\_", Tissue)] <- "Blood"
Tissue[grepl("TK3BMCellsbeforetransplant", Tissue)] <- "time0"
Tissue [1] "library" "time0" "Blood" "Blood" "Blood" "Blood" "Blood"
[8] "Blood" "Blood" "Blood" "Blood" "Blood" "Blood" "Blood"
[15] "Blood" "Blood" "BM" "BM" "BM" "BM" "BM"
[22] "BM" # extract celltype from colnames
Celltype <- gsub(".*\\_.*\\_([A-Z])", "\\1", colnames(data_list$C3))
Celltype[grepl("TK3BMCellsbeforetransplant", Celltype)] <- "time0"
Celltype[grepl("T", Celltype)] <- "Tcell"
Celltype[grepl("B", Celltype)] <- "Bcell"
Celltype[grepl("G", Celltype)] <- "Grn"
Celltype[grepl("\\_", Celltype)] <- "Unsorted"
Celltype [1] "library" "time0" "Grn" "Grn" "Grn" "Grn"
[7] "Grn" "Bcell" "Grn" "Grn" "Unsorted" "Unsorted"
[13] "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Bcell" "Tcell"
[19] "Tcell" "Bcell" "Grn" "Tcell" colnames(data_list$C3) [1] "library" "TK3BMCellsbeforetransplant"
[3] "GK_wk6" "GK_wk12"
[5] "GK_wk18" "GK_wk24"
[7] "GK_wk30" "GK_wk36_B"
[9] "GK_wk42" "GK_wk47"
[11] "L_wk6" "L_wk12"
[13] "L_wk18" "L_wk24"
[15] "L_wk30" "L_wk42"
[17] "GK_BM_B" "GK_BM_T19"
[19] "GK_BM_T20" "L_BM_B"
[21] "L_BM_G" "L_BM_T" C3_meta <- data.frame(
Sample = colnames(data_list$C3),
Donor = rep("C3", ncol(data_list$C3)),
xCondition = xCondition,
Time = Time,
Tissue = Tissue,
Celltype = Celltype
)
# View(data_list$C3)5.4 C10
# data_list$C10 %>% View
data_list$C10 <- as.data.frame(data_list$C10)
# Subset the dataframe to exclude rows where the first column (barcodeID) is NA
any(is.na(data_list$C10[, 1]))[1] FALSEdata_list$C10 <- data_list$C10[!is.na(data_list$C10[, 1]), ]
dim(data_list$C10)[1] 1026 30# set up the first column of each mat as rownames (barcode ID)
rownames(data_list$C10) <- data_list$C10[,1]
data_list$C10 <- data_list$C10[,-1]
colnames(data_list$C10) [1] "['Blood_wk_4_GK'" "'Blood_wk_10_GK'"
[3] "'Blood_wk_16_GK'" "'Blood_wk_22_GK'"
[5] "'Blood B_wk_24_GK'" "'Blood T_wk_24_GK'"
[7] "'Blood G_wk_24_GK'" "'Blood Other_wk_24_GK'"
[9] "'Front BM B_wk_24_GK'" "'Front BM T_wk_24_GK'"
[11] "'Front BM G_wk_24_GK'" "'Front BM Other_wk_24_GK'"
[13] "'Left BM B_wk_24_GK'" "'Left BM T_wk_24_GK'"
[15] "'Left BM G_wk_24_GK'" "'Left BM Other_wk_24_GK'"
[17] "'Left BM Unsorted_wk_24_GK'" "'Right BM B_wk_24_GK'"
[19] "'Right BM T_wk_24_GK'" "'Right BM G_wk_24_GK'"
[21] "'Right BM Other_wk_24_GK'" "'Right BM Unsorted_wk_24_GK'"
[23] "'Spine BM B_wk_24_GK'" "'Spine BM T_wk_24_GK'"
[25] "'Spine BM G_wk_24_GK'" "'Spine BM Other_wk_24_GK'"
[27] "'Spine BM Unsorted_wk_24_GK'" "'Liver Liver_wk_24_GK'"
[29] "'Spleen Spleen_wk_24_GK']" # delete empty or processed columns
del <- c("...39", "...40", "Present", "Total", "Log_total")
data_list$C10 <- data_list$C10[, !colnames(data_list$C10) %in% del]
# delete processed "sum" columns
data_list$C10 <- data_list$C10[, !grepl("Sum", colnames(data_list$C10))]
data_list$C10 <- data_list$C10[, !grepl("sum", colnames(data_list$C10))]
# delete processed "shannon" rows
rownames(data_list$C10) [!grepl("^[ATGC]+$", rownames(data_list$C10))]character(0)rownames(data_list$C10) [!grepl("^[ATGC\\\"]+$", rownames(data_list$C10))]character(0)data_list$C10 <- data_list$C10[grepl("^[ATGC\\\"]+$", rownames(data_list$C10)),]
dim(data_list$C10)[1] 1026 29# Convert NA values to 0 in the matrix
data_list$C10[is.na(data_list$C10)] <- 0
!any(is.na(data_list$C10))[1] TRUE# Convert character data to numeric
C10_count_numeric <- apply(data_list$C10, 2, as.numeric)
rownames(C10_count_numeric) <- rownames(data_list$C10)
data_list$C10 <- C10_count_numeric
# View sample names
colnames(data_list$C10) [1] "['Blood_wk_4_GK'" "'Blood_wk_10_GK'"
[3] "'Blood_wk_16_GK'" "'Blood_wk_22_GK'"
[5] "'Blood B_wk_24_GK'" "'Blood T_wk_24_GK'"
[7] "'Blood G_wk_24_GK'" "'Blood Other_wk_24_GK'"
[9] "'Front BM B_wk_24_GK'" "'Front BM T_wk_24_GK'"
[11] "'Front BM G_wk_24_GK'" "'Front BM Other_wk_24_GK'"
[13] "'Left BM B_wk_24_GK'" "'Left BM T_wk_24_GK'"
[15] "'Left BM G_wk_24_GK'" "'Left BM Other_wk_24_GK'"
[17] "'Left BM Unsorted_wk_24_GK'" "'Right BM B_wk_24_GK'"
[19] "'Right BM T_wk_24_GK'" "'Right BM G_wk_24_GK'"
[21] "'Right BM Other_wk_24_GK'" "'Right BM Unsorted_wk_24_GK'"
[23] "'Spine BM B_wk_24_GK'" "'Spine BM T_wk_24_GK'"
[25] "'Spine BM G_wk_24_GK'" "'Spine BM Other_wk_24_GK'"
[27] "'Spine BM Unsorted_wk_24_GK'" "'Liver Liver_wk_24_GK'"
[29] "'Spleen Spleen_wk_24_GK']" # clean any character that is not a letter, digit, or underscore with an empty string, effectively removing unwanted symbols.
colnames(data_list$C10) <- gsub("[^a-zA-Z0-9_]", "", colnames(data_list$C10))
# extract condition from colnames. I don;t know what GK / L means.
xCondition <- gsub(".*\\_.*\\_.*\\_(.*)", "\\1", colnames(data_list$C10))
xCondition [1] "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK"
[16] "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK" "GK"# extract time from colnames
Time <- gsub(".*\\_(wk\\_[0-9]{1,}).*", "\\1", colnames(data_list$C10))
Time <- gsub("\\_", "", Time)
Time [1] "wk4" "wk10" "wk16" "wk22" "wk24" "wk24" "wk24" "wk24" "wk24" "wk24"
[11] "wk24" "wk24" "wk24" "wk24" "wk24" "wk24" "wk24" "wk24" "wk24" "wk24"
[21] "wk24" "wk24" "wk24" "wk24" "wk24" "wk24" "wk24" "wk24" "wk24"# extract tissue from colnames
Tissue <- gsub("(.*)\\_.*\\_.*\\_.*", "\\1", colnames(data_list$C10))
Tissue[grepl("Blood", Tissue)] <- "Blood"
Tissue[grepl("BM", Tissue)] <- "BM"
Tissue[grepl("Liver", Tissue)] <- "Liver"
Tissue[grepl("Spleen", Tissue)] <- "Spleen"
Tissue [1] "Blood" "Blood" "Blood" "Blood" "Blood" "Blood" "Blood" "Blood"
[9] "BM" "BM" "BM" "BM" "BM" "BM" "BM" "BM"
[17] "BM" "BM" "BM" "BM" "BM" "BM" "BM" "BM"
[25] "BM" "BM" "BM" "Liver" "Spleen"# extract position
mix <- gsub("(.*)\\_.*\\_.*\\_.*", "\\1", colnames(data_list$C10))
Position <- vector("character", length(mix))
Position[grepl("BM", mix)] <- gsub("(.*)BM.*", "\\1", mix)[grepl("BM", mix)]
Position [1] "" "" "" "" "" "" "" "" "Front"
[10] "Front" "Front" "Front" "Left" "Left" "Left" "Left" "Left" "Right"
[19] "Right" "Right" "Right" "Right" "Spine" "Spine" "Spine" "Spine" "Spine"
[28] "" "" # extract celltype from colnames
mix <- gsub("(.*)\\_.*\\_.*\\_.*", "\\1", colnames(data_list$C10))
Celltype <- vector("character", length(mix))
Celltype[grepl("Blood", mix)] <- gsub(".*Blood(.*)", "\\1", mix)[grepl("Blood", mix)]
Celltype[grepl("BM", mix)] <- gsub(".*BM(.*)", "\\1", mix)[grepl("BM", mix)]
Celltype[Celltype == ""] <- "Unsorted"
Celltype[Celltype == "B"] <- "Bcell"
Celltype[Celltype == "T"] <- "Tcell"
Celltype[Celltype == "G"] <- "Grn"
Celltype [1] "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Bcell" "Tcell"
[7] "Grn" "Other" "Bcell" "Tcell" "Grn" "Other"
[13] "Bcell" "Tcell" "Grn" "Other" "Unsorted" "Bcell"
[19] "Tcell" "Grn" "Other" "Unsorted" "Bcell" "Tcell"
[25] "Grn" "Other" "Unsorted" "Unsorted" "Unsorted"colnames(data_list$C10) [1] "Blood_wk_4_GK" "Blood_wk_10_GK"
[3] "Blood_wk_16_GK" "Blood_wk_22_GK"
[5] "BloodB_wk_24_GK" "BloodT_wk_24_GK"
[7] "BloodG_wk_24_GK" "BloodOther_wk_24_GK"
[9] "FrontBMB_wk_24_GK" "FrontBMT_wk_24_GK"
[11] "FrontBMG_wk_24_GK" "FrontBMOther_wk_24_GK"
[13] "LeftBMB_wk_24_GK" "LeftBMT_wk_24_GK"
[15] "LeftBMG_wk_24_GK" "LeftBMOther_wk_24_GK"
[17] "LeftBMUnsorted_wk_24_GK" "RightBMB_wk_24_GK"
[19] "RightBMT_wk_24_GK" "RightBMG_wk_24_GK"
[21] "RightBMOther_wk_24_GK" "RightBMUnsorted_wk_24_GK"
[23] "SpineBMB_wk_24_GK" "SpineBMT_wk_24_GK"
[25] "SpineBMG_wk_24_GK" "SpineBMOther_wk_24_GK"
[27] "SpineBMUnsorted_wk_24_GK" "LiverLiver_wk_24_GK"
[29] "SpleenSpleen_wk_24_GK" C10_meta <- data.frame(
Sample = colnames(data_list$C10),
Donor = rep("C10", ncol(data_list$C10)),
xCondition = xCondition,
Time = Time,
Tissue = Tissue,
Position = Position,
Celltype = Celltype
)
# View(data_list$C10)5.5 C15
# data_list$C15 %>% View
data_list$C15 <- as.data.frame(data_list$C15)
# Subset the dataframe to exclude rows where the first column (barcodeID) is NA
any(is.na(data_list$C15[, 1]))[1] FALSEdata_list$C15 <- data_list$C15[!is.na(data_list$C15[, 1]), ]
dim(data_list$C15)[1] 1613 21# set up the first column of each mat as rownames (barcode ID)
rownames(data_list$C15) <- data_list$C15[,1]
data_list$C15 <- data_list$C15[,-1]
colnames(data_list$C15) [1] "wk4" "wk12" "wk18_T" "wk18_O" "BMFront_B"
[6] "BMFront_T" "BMFront_G" "BMFront_O" "BMLeft_B" "BMLeft_T"
[11] "BMLeft_G" "BMLeft_O" "BM_Left_Sum" "BMRight_B" "BMRight_G"
[16] "BMRight_O" "BMSpine_B" "BMSpine_T" "BMSpine_G" "BMSpine_O" # delete empty or processed columns
del <- c("...39", "...40", "Present", "Total", "Log_total")
data_list$C15 <- data_list$C15[, !colnames(data_list$C15) %in% del]
# delete processed "sum" columns
data_list$C15 <- data_list$C15[, !grepl("Sum", colnames(data_list$C15))]
data_list$C15 <- data_list$C15[, !grepl("sum", colnames(data_list$C15))]
# delete processed "shannon" rows
rownames(data_list$C15) [!grepl("^[ATGC]+$", rownames(data_list$C15))]character(0)rownames(data_list$C15) [!grepl("^[ATGC\\\"]+$", rownames(data_list$C15))]character(0)data_list$C15 <- data_list$C15[grepl("^[ATGC\\\"]+$", rownames(data_list$C15)),]
dim(data_list$C15)[1] 1613 19# Convert NA values to 0 in the matrix
data_list$C15[is.na(data_list$C15)] <- 0
!any(is.na(data_list$C15))[1] TRUE# Convert character data to numeric
C15_count_numeric <- apply(data_list$C15, 2, as.numeric)
rownames(C15_count_numeric) <- rownames(data_list$C15)
data_list$C15 <- C15_count_numeric
# View sample names
colnames(data_list$C15) [1] "wk4" "wk12" "wk18_T" "wk18_O" "BMFront_B" "BMFront_T"
[7] "BMFront_G" "BMFront_O" "BMLeft_B" "BMLeft_T" "BMLeft_G" "BMLeft_O"
[13] "BMRight_B" "BMRight_G" "BMRight_O" "BMSpine_B" "BMSpine_T" "BMSpine_G"
[19] "BMSpine_O"# clean any character that is not a letter, digit, or underscore with an empty string, effectively removing unwanted symbols.
colnames(data_list$C15) <- gsub("[^a-zA-Z0-9_]", "", colnames(data_list$C15))
# extract time from colnames
Time <- gsub(".*(wk[0-9]{1,}).*", "\\1", colnames(data_list$C15))
Time[grepl("\\_", Time)] <- "sac"
Time [1] "wk4" "wk12" "wk18" "wk18" "sac" "sac" "sac" "sac" "sac" "sac"
[11] "sac" "sac" "sac" "sac" "sac" "sac" "sac" "sac" "sac" # extract tissue from colnames
Tissue <- vector("character", ncol(data_list$C15))
Tissue[grepl("BM", colnames(data_list$C15))] <- "BM"
Tissue[Tissue == ""] <- "Blood"
Tissue [1] "Blood" "Blood" "Blood" "Blood" "BM" "BM" "BM" "BM" "BM"
[10] "BM" "BM" "BM" "BM" "BM" "BM" "BM" "BM" "BM"
[19] "BM" # extract position
mix <- gsub("(.*)\\_.*", "\\1", colnames(data_list$C15))
Position <- vector("character", ncol(data_list$C15))
Position[grepl("BM", mix)] <- gsub("BM(.*)", "\\1", mix)[grepl("BM", mix)]
Position [1] "" "" "" "" "Front" "Front" "Front" "Front" "Left"
[10] "Left" "Left" "Left" "Right" "Right" "Right" "Spine" "Spine" "Spine"
[19] "Spine"# extract celltype from colnames
Celltype <- vector("character", ncol(data_list$C15))
Celltype[grepl("\\_",colnames(data_list$C15))] <- gsub(".*\\_(.*)", "\\1", colnames(data_list$C15))[grepl("\\_",colnames(data_list$C15))]
Celltype[Celltype == ""] <- "Unsorted"
Celltype[Celltype == "B"] <- "Bcell"
Celltype[Celltype == "T"] <- "Tcell"
Celltype[Celltype == "G"] <- "Grn"
Celltype[Celltype == "O"] <- "Other"
Celltype [1] "Unsorted" "Unsorted" "Tcell" "Other" "Bcell" "Tcell"
[7] "Grn" "Other" "Bcell" "Tcell" "Grn" "Other"
[13] "Bcell" "Grn" "Other" "Bcell" "Tcell" "Grn"
[19] "Other" colnames(data_list$C15) [1] "wk4" "wk12" "wk18_T" "wk18_O" "BMFront_B" "BMFront_T"
[7] "BMFront_G" "BMFront_O" "BMLeft_B" "BMLeft_T" "BMLeft_G" "BMLeft_O"
[13] "BMRight_B" "BMRight_G" "BMRight_O" "BMSpine_B" "BMSpine_T" "BMSpine_G"
[19] "BMSpine_O"C15_meta <- data.frame(
Sample = colnames(data_list$C15),
Donor = rep("C15", ncol(data_list$C15)),
Time = Time,
Tissue = Tissue,
Position = Position,
Celltype = Celltype
)
# View(data_list$C15)5.6 C21
# data_list$C21 %>% View
data_list$C21 <- as.data.frame(data_list$C21)
# Subset the dataframe to exclude rows where the first column (barcodeID) is NA
any(is.na(data_list$C21[, 1]))[1] FALSEdata_list$C21 <- data_list$C21[!is.na(data_list$C21[, 1]), ]
dim(data_list$C21)[1] 1515 36# set up the first column of each mat as rownames (barcode ID)
rownames(data_list$C21) <- data_list$C21[,1]
data_list$C21 <- data_list$C21[,-1]
colnames(data_list$C21) [1] "wk9" "wk14" "wk20"
[4] "wk22_unsorted" "wk22_B" "wk22_T"
[7] "wk22_G" "wk22_sum" "BMFront_unsorted"
[10] "BMFront_B" "BMFront_T" "BMFront_G"
[13] "BMFront_sum" "BMLeft_unsorted" "BMLeft_B"
[16] "BMLeft_T" "BMLeft_G" "BMLeft_sum"
[19] "BMRight_unsorted" "BMRight_B" "BMRight_T"
[22] "BMRight_G" "BMRight_sum" "BMSpine_unsorted"
[25] "BMSpine_B" "BMSpine_T" "BMSpine_G"
[28] "BMSpine_sum" "BMPelvis_unsorted" "BMPelvis_B"
[31] "BMPelvis_T" "BMPelvis_G" "BMPelvis_sum"
[34] "Spleen" "Liver" # delete empty or processed columns
del <- c("...39", "...40", "Present", "Total", "Log_total")
data_list$C21 <- data_list$C21[, !colnames(data_list$C21) %in% del]
# delete processed "sum" columns
data_list$C21 <- data_list$C21[, !grepl("Sum", colnames(data_list$C21))]
data_list$C21 <- data_list$C21[, !grepl("sum", colnames(data_list$C21))]
# delete processed "shannon" rows
rownames(data_list$C21) [!grepl("^[ATGC]+$", rownames(data_list$C21))]character(0)rownames(data_list$C21) [!grepl("^[ATGC\\\"]+$", rownames(data_list$C21))]character(0)data_list$C21 <- data_list$C21[grepl("^[ATGC\\\"]+$", rownames(data_list$C21)),]
dim(data_list$C21)[1] 1515 29# Convert NA values to 0 in the matrix
data_list$C21[is.na(data_list$C21)] <- 0
!any(is.na(data_list$C21))[1] TRUE# Convert character data to numeric
C21_count_numeric <- apply(data_list$C21, 2, as.numeric)
rownames(C21_count_numeric) <- rownames(data_list$C21)
data_list$C21 <- C21_count_numeric
# View sample names
colnames(data_list$C21) [1] "wk9" "wk14" "wk20"
[4] "wk22_unsorted" "wk22_B" "wk22_T"
[7] "wk22_G" "BMFront_unsorted" "BMFront_B"
[10] "BMFront_T" "BMFront_G" "BMLeft_unsorted"
[13] "BMLeft_B" "BMLeft_T" "BMLeft_G"
[16] "BMRight_unsorted" "BMRight_B" "BMRight_T"
[19] "BMRight_G" "BMSpine_unsorted" "BMSpine_B"
[22] "BMSpine_T" "BMSpine_G" "BMPelvis_unsorted"
[25] "BMPelvis_B" "BMPelvis_T" "BMPelvis_G"
[28] "Spleen" "Liver" # clean any character that is not a letter, digit, or underscore with an empty string, effectively removing unwanted symbols.
colnames(data_list$C21) <- gsub("[^a-zA-Z0-9_]", "", colnames(data_list$C21))
# extract time from colnames
Time <- gsub("(wk[0-9]{1,}).*", "\\1", colnames(data_list$C21))
Time[grepl("BM|Spleen|Liver", Time)] <- "sac"
Time [1] "wk9" "wk14" "wk20" "wk22" "wk22" "wk22" "wk22" "sac" "sac" "sac"
[11] "sac" "sac" "sac" "sac" "sac" "sac" "sac" "sac" "sac" "sac"
[21] "sac" "sac" "sac" "sac" "sac" "sac" "sac" "sac" "sac" # extract tissue from colnames
Tissue <- gsub("(BM|Spleen|Liver).*", "\\1", colnames(data_list$C21))
Tissue[grepl("wk", Tissue)] <- "Blood"
Tissue [1] "Blood" "Blood" "Blood" "Blood" "Blood" "Blood" "Blood" "BM"
[9] "BM" "BM" "BM" "BM" "BM" "BM" "BM" "BM"
[17] "BM" "BM" "BM" "BM" "BM" "BM" "BM" "BM"
[25] "BM" "BM" "BM" "Spleen" "Liver" # extract position
Position <- vector("character", ncol(data_list$C21))
Position[grepl("BM", colnames(data_list$C21))] <- gsub("BM(.*)\\_.*", "\\1", colnames(data_list$C21))[grepl("BM", colnames(data_list$C21))]
Position [1] "" "" "" "" "" "" "" "Front"
[9] "Front" "Front" "Front" "Left" "Left" "Left" "Left" "Right"
[17] "Right" "Right" "Right" "Spine" "Spine" "Spine" "Spine" "Pelvis"
[25] "Pelvis" "Pelvis" "Pelvis" "" "" # extract celltype from colnames
Celltype <- vector("character", ncol(data_list$C21))
Celltype[grepl("\\_", colnames(data_list$C21))] <- gsub(".*\\_(.*)", "\\1", colnames(data_list$C21))[grepl("\\_", colnames(data_list$C21))]
Celltype[Celltype == ""] <- "Unsorted"
Celltype[Celltype == "unsorted"] <- "Unsorted"
Celltype[Celltype == "B"] <- "Bcell"
Celltype[Celltype == "T"] <- "Tcell"
Celltype[Celltype == "G"] <- "Grn"
Celltype[Celltype == "O"] <- "Other"
Celltype [1] "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Bcell" "Tcell"
[7] "Grn" "Unsorted" "Bcell" "Tcell" "Grn" "Unsorted"
[13] "Bcell" "Tcell" "Grn" "Unsorted" "Bcell" "Tcell"
[19] "Grn" "Unsorted" "Bcell" "Tcell" "Grn" "Unsorted"
[25] "Bcell" "Tcell" "Grn" "Unsorted" "Unsorted"colnames(data_list$C21) [1] "wk9" "wk14" "wk20"
[4] "wk22_unsorted" "wk22_B" "wk22_T"
[7] "wk22_G" "BMFront_unsorted" "BMFront_B"
[10] "BMFront_T" "BMFront_G" "BMLeft_unsorted"
[13] "BMLeft_B" "BMLeft_T" "BMLeft_G"
[16] "BMRight_unsorted" "BMRight_B" "BMRight_T"
[19] "BMRight_G" "BMSpine_unsorted" "BMSpine_B"
[22] "BMSpine_T" "BMSpine_G" "BMPelvis_unsorted"
[25] "BMPelvis_B" "BMPelvis_T" "BMPelvis_G"
[28] "Spleen" "Liver" C21_meta <- data.frame(
Sample = colnames(data_list$C21),
Donor = rep("C21", ncol(data_list$C21)),
Time = Time,
Tissue = Tissue,
Position = Position,
Celltype = Celltype
)
# View(data_list$C21)5.7 C22
# data_list$C22 %>% View
data_list$C22 <- as.data.frame(data_list$C22)
# Subset the dataframe to exclude rows where the first column (barcodeID) is NA
any(is.na(data_list$C22[, 1]))[1] FALSEdata_list$C22 <- data_list$C22[!is.na(data_list$C22[, 1]), ]
dim(data_list$C22)[1] 1673 19# set up the first column of each mat as rownames (barcode ID)
rownames(data_list$C22) <- data_list$C22[,1]
data_list$C22 <- data_list$C22[,-1]
colnames(data_list$C22) [1] "wk10" "wk14" "wk20"
[4] "wk27" "wk33" "sac"
[7] "BM_Front" "BM_Left" "BM_Right_unsorted"
[10] "BM_Right_T" "BM_Right_G" "BM_Right_O"
[13] "BM_Spine_B" "BM_Spine_T" "BM_Spine_G"
[16] "BM_Spine_O" "BM_Spine_sum" "Spleen" # delete empty or processed columns
del <- c("...39", "...40", "Present", "Total", "Log_total")
data_list$C22 <- data_list$C22[, !colnames(data_list$C22) %in% del]
# delete processed "sum" columns
data_list$C22 <- data_list$C22[, !grepl("Sum", colnames(data_list$C22))]
data_list$C22 <- data_list$C22[, !grepl("sum", colnames(data_list$C22))]
# delete processed "shannon" rows
rownames(data_list$C22) [!grepl("^[ATGC]+$", rownames(data_list$C22))] [1] "AGTAAGGACTGAGTATACGGTATACTACNNNNNNGCCGACG"
[2] "AAGGACAACTCAGTCCCCGAAATACNNNANCATGATTGACG"
[3] "AAGGACAACTCAGTCCCCGAAATACGNCNCCATGATTGACG"
[4] "AAGGACAACTCAGTCCCCGAAATACNNNNNNATGATTGACG"
[5] "AAGGACAACTCAGTCCCCGAAATACNNCACCATGATTGACG"
[6] "AAGGATTACAATGTTGCCGGCCTATNNCATGTTGTATGACG"
[7] "AAGGACAACTCAGTCCCCGAAATACGNNACCATGATTGACG"
[8] "AAGGGGAACGGTGTACACGAACTATNNNNNNATGGTAGACG"
[9] "AAGGGCGACGCTGTATTCGACCTATNNNANTATGGAAGACG"
[10] "AAGGGCGACGCTGTATTCGACCTATNNCACTATGGAAGACG"
[11] "AAGGATTACAATGTTGCCGGCCTATGNNNNNNNNNNTGACG"
[12] "AAGGAGTGCGTGGTTTTCGCAATAANNNATGTTGTNNNACG"
[13] "AAGGGGAACGGTGTACACGAACTATNNNANTATGGTAGACG"
[14] "AAGGGCGACGCTGTATTCGACCTATANCNCTATGGAAGACG"
[15] "AAGGGGAACGGTGTACACGAACTATNNCACTATGGTAGACG"
[16] "AAGGGGAACGGTGTACACGAACTATNCCACTATGGNAGACG"
[17] "AAGGGCGACGCTGTATTCGACCTATANNNNNNNNNNAGACG"
[18] "AAGGATTACAATGTTGCCGGCCTATNNNANGTTGTATGACG"
[19] "AAGGATTACAATGTTGCCGGCCTATNGCATGTTGTNTGACG"
[20] "AAGGATTACAATGTTGCCGGCCTATGNNNNNTTGTATGACG"
[21] "AAGGATTACAATGTTGCCGGCCTATGNCNTGTTGTATGACG"
[22] "AAGGAGTGCGTGGTTTTCGCAATAANNCATGTTGTCCGACG"
[23] "AAGGATGACCACGTGATCGACTTAGCNNAAAATGGACGACG"
[24] "AAGGTGAACGTAGTAAACGGCGTACNNCACAATGATCGACG"
[25] "AAGGCACACATTGTTTGCGTAGTATTNNACTNNGACAGACG"
[26] "AAGGGGAACGGTGTACACGAACTATNNNNNNANGGTAGACG"
[27] "AAGGAGTGCGTGGTTTTCGCAATAACNCNTGTTGTCCGACG"
[28] "AAGGGCGACGCTGTATTCGACCTATNTCNCNATGGAAGACG"
[29] "AAGGGCGACGCTGTATTCGACCTATNNNNNTATGGAAGACG"
[30] "AAGGGCGACGCTGTATTCGACCTATNNNNNNNNNNNNGACG"
[31] "AAGGATTACAATGTTGCCGGCCTATGNNNTNNNNNNTGACG"
[32] "AAGGACGACGTTGTGTGCGATATAGNNCAGTATGTTCGACG"
[33] "AAGGCATACCGTGTATCCGTCATACANNNNNCTGTAGGACG"
[34] "AAGGAGTGCGTGGTTTTCGCAATAANACNNNTTGTCCGACG"rownames(data_list$C22) [!grepl("^[ATGC\\\"]+$", rownames(data_list$C22))] [1] "AGTAAGGACTGAGTATACGGTATACTACNNNNNNGCCGACG"
[2] "AAGGACAACTCAGTCCCCGAAATACNNNANCATGATTGACG"
[3] "AAGGACAACTCAGTCCCCGAAATACGNCNCCATGATTGACG"
[4] "AAGGACAACTCAGTCCCCGAAATACNNNNNNATGATTGACG"
[5] "AAGGACAACTCAGTCCCCGAAATACNNCACCATGATTGACG"
[6] "AAGGATTACAATGTTGCCGGCCTATNNCATGTTGTATGACG"
[7] "AAGGACAACTCAGTCCCCGAAATACGNNACCATGATTGACG"
[8] "AAGGGGAACGGTGTACACGAACTATNNNNNNATGGTAGACG"
[9] "AAGGGCGACGCTGTATTCGACCTATNNNANTATGGAAGACG"
[10] "AAGGGCGACGCTGTATTCGACCTATNNCACTATGGAAGACG"
[11] "AAGGATTACAATGTTGCCGGCCTATGNNNNNNNNNNTGACG"
[12] "AAGGAGTGCGTGGTTTTCGCAATAANNNATGTTGTNNNACG"
[13] "AAGGGGAACGGTGTACACGAACTATNNNANTATGGTAGACG"
[14] "AAGGGCGACGCTGTATTCGACCTATANCNCTATGGAAGACG"
[15] "AAGGGGAACGGTGTACACGAACTATNNCACTATGGTAGACG"
[16] "AAGGGGAACGGTGTACACGAACTATNCCACTATGGNAGACG"
[17] "AAGGGCGACGCTGTATTCGACCTATANNNNNNNNNNAGACG"
[18] "AAGGATTACAATGTTGCCGGCCTATNNNANGTTGTATGACG"
[19] "AAGGATTACAATGTTGCCGGCCTATNGCATGTTGTNTGACG"
[20] "AAGGATTACAATGTTGCCGGCCTATGNNNNNTTGTATGACG"
[21] "AAGGATTACAATGTTGCCGGCCTATGNCNTGTTGTATGACG"
[22] "AAGGAGTGCGTGGTTTTCGCAATAANNCATGTTGTCCGACG"
[23] "AAGGATGACCACGTGATCGACTTAGCNNAAAATGGACGACG"
[24] "AAGGTGAACGTAGTAAACGGCGTACNNCACAATGATCGACG"
[25] "AAGGCACACATTGTTTGCGTAGTATTNNACTNNGACAGACG"
[26] "AAGGGGAACGGTGTACACGAACTATNNNNNNANGGTAGACG"
[27] "AAGGAGTGCGTGGTTTTCGCAATAACNCNTGTTGTCCGACG"
[28] "AAGGGCGACGCTGTATTCGACCTATNTCNCNATGGAAGACG"
[29] "AAGGGCGACGCTGTATTCGACCTATNNNNNTATGGAAGACG"
[30] "AAGGGCGACGCTGTATTCGACCTATNNNNNNNNNNNNGACG"
[31] "AAGGATTACAATGTTGCCGGCCTATGNNNTNNNNNNTGACG"
[32] "AAGGACGACGTTGTGTGCGATATAGNNCAGTATGTTCGACG"
[33] "AAGGCATACCGTGTATCCGTCATACANNNNNCTGTAGGACG"
[34] "AAGGAGTGCGTGGTTTTCGCAATAANACNNNTTGTCCGACG"data_list$C22 <- data_list$C22[grepl("^[ATGC\\\"]+$", rownames(data_list$C22)),]
dim(data_list$C22)[1] 1639 17# Convert NA values to 0 in the matrix
data_list$C22[is.na(data_list$C22)] <- 0
!any(is.na(data_list$C22))[1] TRUE# Convert character data to numeric
C22_count_numeric <- apply(data_list$C22, 2, as.numeric)
rownames(C22_count_numeric) <- rownames(data_list$C22)
data_list$C22 <- C22_count_numeric
# View sample names
colnames(data_list$C22) [1] "wk10" "wk14" "wk20"
[4] "wk27" "wk33" "sac"
[7] "BM_Front" "BM_Left" "BM_Right_unsorted"
[10] "BM_Right_T" "BM_Right_G" "BM_Right_O"
[13] "BM_Spine_B" "BM_Spine_T" "BM_Spine_G"
[16] "BM_Spine_O" "Spleen" # clean any character that is not a letter, digit, or underscore with an empty string, effectively removing unwanted symbols.
colnames(data_list$C22) <- gsub("[^a-zA-Z0-9_]", "", colnames(data_list$C22))
# extract time from colnames
Time <- gsub(".*\\_(wk\\_[0-9]{1,}).*", "\\1", colnames(data_list$C22))
Time[grepl("\\_", Time)] <- "sac"
Time[Time == "Spleen"] <- "sac"
Time [1] "wk10" "wk14" "wk20" "wk27" "wk33" "sac" "sac" "sac" "sac" "sac"
[11] "sac" "sac" "sac" "sac" "sac" "sac" "sac" # extract tissue from colnames
Tissue <- gsub("(.*)\\_.*\\_.*", "\\1", colnames(data_list$C22))
Tissue <- gsub("(.*)\\_.*", "\\1", Tissue)
Tissue[grepl("wk", Tissue)] <- "Blood"
Tissue [1] "Blood" "Blood" "Blood" "Blood" "Blood" "sac" "BM" "BM"
[9] "BM" "BM" "BM" "BM" "BM" "BM" "BM" "BM"
[17] "Spleen"# extract position
Position <- vector("character", ncol(data_list$C22))
Position[grepl("BM", colnames(data_list$C22))] <- gsub("BM\\_(.*)\\_.*", "\\1", colnames(data_list$C22))[grepl("BM", colnames(data_list$C22))]
Position[grepl("BM", Position)] <- gsub("BM\\_(.*)", "\\1", Position)[grepl("BM", Position)]
Position [1] "" "" "" "" "" "" "Front" "Left" "Right"
[10] "Right" "Right" "Right" "Spine" "Spine" "Spine" "Spine" "" # extract celltype from colnames
Celltype <- vector("character", ncol(data_list$C22))
Celltype[grepl("BM\\_.*\\_.*", colnames(data_list$C22))] <- gsub("BM\\_.*\\_(.*)", "\\1", colnames(data_list$C22))[grepl("BM\\_.*\\_.*", colnames(data_list$C22))]
Celltype[Celltype == ""] <- "Unsorted"
Celltype[Celltype == "unsorted"] <- "Unsorted"
Celltype[Celltype == "B"] <- "Bcell"
Celltype[Celltype == "T"] <- "Tcell"
Celltype[Celltype == "G"] <- "Grn"
Celltype[Celltype == "O"] <- "Other"
Celltype [1] "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Unsorted"
[7] "Unsorted" "Unsorted" "Unsorted" "Tcell" "Grn" "Other"
[13] "Bcell" "Tcell" "Grn" "Other" "Unsorted"colnames(data_list$C22) [1] "wk10" "wk14" "wk20"
[4] "wk27" "wk33" "sac"
[7] "BM_Front" "BM_Left" "BM_Right_unsorted"
[10] "BM_Right_T" "BM_Right_G" "BM_Right_O"
[13] "BM_Spine_B" "BM_Spine_T" "BM_Spine_G"
[16] "BM_Spine_O" "Spleen" C22_meta <- data.frame(
Sample = colnames(data_list$C22),
Donor = rep("C22", ncol(data_list$C22)),
Time = Time,
Tissue = Tissue,
Position = Position,
Celltype = Celltype
)
# View(data_list$C22)5.8 C23
# data_list$C23 %>% View
data_list$C23 <- as.data.frame(data_list$C23)
# Subset the dataframe to exclude rows where the first column (barcodeID) is NA
any(is.na(data_list$C23[, 1]))[1] FALSEdata_list$C23 <- data_list$C23[!is.na(data_list$C23[, 1]), ]
dim(data_list$C23)[1] 3590 14# set up the first column of each mat as rownames (barcode ID)
rownames(data_list$C23) <- data_list$C23[,1]
data_list$C23 <- data_list$C23[,-1]
colnames(data_list$C23) [1] "wk11" "Sac" "BM_Front_unsorted"
[4] "BM_Left_unsorted" "BM_Right_unsorted" "BM_Right_T"
[7] "BM_Right_G" "BM_Spine_B" "BM_Spine_T"
[10] "BM_Spine_G" "BM_Spine_sum" "Spleen"
[13] "BM_CD34" # delete empty or processed columns
del <- c("...39", "...40", "Present", "Total", "Log_total")
data_list$C23 <- data_list$C23[, !colnames(data_list$C23) %in% del]
# delete processed "sum" columns
data_list$C23 <- data_list$C23[, !grepl("Sum", colnames(data_list$C23))]
data_list$C23 <- data_list$C23[, !grepl("sum", colnames(data_list$C23))]
# delete processed "shannon" rows
rownames(data_list$C23) [!grepl("^[ATGC]+$", rownames(data_list$C23))] [1] "AAGGTATACACAGTGGACGTTTTATNNNNNNCTGTTGGACG"
[2] "AAGGTATACACAGTGGACGTTTTATNNNNNNNNNNNNGACG"
[3] "AAGGTATACACAGTGGACGTTTTATNNNATTCTGTTNGACG"
[4] "AAGGTATACACAGTGGACGTTTTATGNNNNNCTGTTGGANG"
[5] "AAGGAACACTTTGTATGCGTATTAGNNCAATTTGCCGGACG"
[6] "AAGGGAGACGGTGTTCTCGGATTATNNNNCGNNGNNNNANG"
[7] "AAGGATGACAAGGTCTCCGCCATATNNCATAATGCAAGACG"
[8] "AAGGTCCACTAAGTGACCGTACTACANNATAATGTTCGACG"
[9] "AAGGTGCACGTTGTGATCGCCGTAGNNCAGAGTGAGAGACG"
[10] "AAGGGGGACAGTGTTCCCGAATTATANCNTTATGTTAGACG"
[11] "AAGGCAAACTGCGTCTACGTGGTACNNCAACATGCAGGACG"
[12] "AAGGAGTGCGTGGTTTTCGCAATAACNNNNNTTGTCCGACG"rownames(data_list$C23) [!grepl("^[ATGC\\\"]+$", rownames(data_list$C23))] [1] "AAGGTATACACAGTGGACGTTTTATNNNNNNCTGTTGGACG"
[2] "AAGGTATACACAGTGGACGTTTTATNNNNNNNNNNNNGACG"
[3] "AAGGTATACACAGTGGACGTTTTATNNNATTCTGTTNGACG"
[4] "AAGGTATACACAGTGGACGTTTTATGNNNNNCTGTTGGANG"
[5] "AAGGAACACTTTGTATGCGTATTAGNNCAATTTGCCGGACG"
[6] "AAGGGAGACGGTGTTCTCGGATTATNNNNCGNNGNNNNANG"
[7] "AAGGATGACAAGGTCTCCGCCATATNNCATAATGCAAGACG"
[8] "AAGGTCCACTAAGTGACCGTACTACANNATAATGTTCGACG"
[9] "AAGGTGCACGTTGTGATCGCCGTAGNNCAGAGTGAGAGACG"
[10] "AAGGGGGACAGTGTTCCCGAATTATANCNTTATGTTAGACG"
[11] "AAGGCAAACTGCGTCTACGTGGTACNNCAACATGCAGGACG"
[12] "AAGGAGTGCGTGGTTTTCGCAATAACNNNNNTTGTCCGACG"data_list$C23 <- data_list$C23[grepl("^[ATGC\\\"]+$", rownames(data_list$C23)),]
dim(data_list$C23)[1] 3578 12# Convert NA values to 0 in the matrix
data_list$C23[is.na(data_list$C23)] <- 0
!any(is.na(data_list$C23))[1] TRUE# Convert character data to numeric
C23_count_numeric <- apply(data_list$C23, 2, as.numeric)
rownames(C23_count_numeric) <- rownames(data_list$C23)
data_list$C23 <- C23_count_numeric
# View sample names
colnames(data_list$C23) [1] "wk11" "Sac" "BM_Front_unsorted"
[4] "BM_Left_unsorted" "BM_Right_unsorted" "BM_Right_T"
[7] "BM_Right_G" "BM_Spine_B" "BM_Spine_T"
[10] "BM_Spine_G" "Spleen" "BM_CD34" # clean any character that is not a letter, digit, or underscore with an empty string, effectively removing unwanted symbols.
colnames(data_list$C23) <- gsub("[^a-zA-Z0-9_]", "", colnames(data_list$C23))
# extract time from colnames
Time <- gsub("(wk[0-9]{1,}).*", "\\1", colnames(data_list$C23))
Time[grepl("BM|Spleen|Liver|Sac", Time)] <- "sac"
Time [1] "wk11" "sac" "sac" "sac" "sac" "sac" "sac" "sac" "sac" "sac"
[11] "sac" "sac" # extract tissue from colnames
Tissue <- gsub("(BM|Spleen|Liver).*", "\\1", colnames(data_list$C23))
Tissue[grepl("wk", Tissue)] <- "Blood"
Tissue[grepl("Sac", Tissue)] <- "sac"
Tissue [1] "Blood" "sac" "BM" "BM" "BM" "BM" "BM" "BM"
[9] "BM" "BM" "Spleen" "BM" # extract position
Position <- vector("character", ncol(data_list$C23))
Position[grepl("BM\\_.*\\_.*", colnames(data_list$C23))] <- gsub("BM\\_(.*)\\_.*", "\\1", colnames(data_list$C23))[grepl("BM\\_.*\\_.*", colnames(data_list$C23))]
Position [1] "" "" "Front" "Left" "Right" "Right" "Right" "Spine" "Spine"
[10] "Spine" "" "" # extract celltype from colnames
Celltype <- vector("character", ncol(data_list$C23))
Celltype[grepl("\\_", colnames(data_list$C23))] <- gsub(".*\\_(.*)", "\\1", colnames(data_list$C23))[grepl("\\_", colnames(data_list$C23))]
Celltype[Celltype == ""] <- "Unsorted"
Celltype[Celltype == "unsorted"] <- "Unsorted"
Celltype[Celltype == "B"] <- "Bcell"
Celltype[Celltype == "T"] <- "Tcell"
Celltype[Celltype == "G"] <- "Grn"
Celltype[Celltype == "O"] <- "Other"
Celltype [1] "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Unsorted" "Tcell"
[7] "Grn" "Bcell" "Tcell" "Grn" "Unsorted" "CD34" colnames(data_list$C23) [1] "wk11" "Sac" "BM_Front_unsorted"
[4] "BM_Left_unsorted" "BM_Right_unsorted" "BM_Right_T"
[7] "BM_Right_G" "BM_Spine_B" "BM_Spine_T"
[10] "BM_Spine_G" "Spleen" "BM_CD34" C23_meta <- data.frame(
Sample = colnames(data_list$C23),
Donor = rep("C23", ncol(data_list$C23)),
Time = Time,
Tissue = Tissue,
Position = Position,
Celltype = Celltype
)
# View(data_list$C23)5.9 Combine mice
Metadata <- dplyr::bind_rows(C1_meta, C2_meta, C3_meta, C10_meta, C15_meta, C22_meta, C21_meta, C23_meta)
Metadata[is.na(Metadata)] <- ""
data_list <- lapply(data_list, function(mat) as.data.frame(t(mat)))
CountData <- dplyr::bind_rows(data_list)
CountData <- t(CountData)6 Save to a barbieQ object
Note that clones (barcodes) between different donors don’t match.
Clones from each donor were labelled separately, using the same barcode (lentiviral) library.
Same barcode across donors doesn’t refer to the same clone!
For convenience, herein we combined samples from all donors together.
## convert all NA into zero and continue
CountData %>% is.na() %>% any()[1] TRUECountData[is.na(CountData)] <- 0
CountData %>% is.na() %>% any()[1] FALSExenoHSPC <- createBarbieQ(object = CountData, sampleMetadata = Metadata)sample names set up in line with `object`continuing with missing `factorColors`.7 Tag top barcodes
dim(xenoHSPC)[1] 10149 199## find minimum group size
targets <- as.data.frame(xenoHSPC$sampleMetadata) %>%
mutate(Group = paste0(Donor, Celltype))
targets$Group %>% table().
C10Bcell C10Grn C10Other C10Tcell C10Unsorted C15Bcell
5 5 5 5 9 4
C15Grn C15Other C15Tcell C15Unsorted C1Bcell C1Grn
4 5 4 2 6 6
C1library C1Tcell C1Unsorted C21Bcell C21Grn C21Tcell
1 6 14 6 6 6
C21Unsorted C22Bcell C22Grn C22Other C22Tcell C22Unsorted
11 1 2 2 2 10
C23Bcell C23CD34 C23Grn C23Tcell C23Unsorted C2Bcell
1 1 2 2 6 5
C2Grn C2library C2Other C2Tcell C2Unsorted C3Bcell
4 1 3 3 22 3
C3Grn C3library C3Tcell C3time0 C3Unsorted
8 1 3 1 6 ## select sample to consider, exclude TP0 when tagging
xenoHSPC <- barbieQ::tagTopBarcodes(barbieQ = xenoHSPC,
# activeSamples = xenoHSPC$sampleMetadata$Sample != "library",
nSampleThreshold = 3
)
rowData(xenoHSPC)$isTopBarcode$isTop %>% table().
FALSE TRUE
9278 871 According to the original paper, for each donor, barcode library contains around 800 unique barcodes.
Check out filtered “top” barcodes.
n_uni_BC <- xenoHSPC@assays@data$isTopAssay %>% colSums()
n_uni_BC %>% sort(decreasing = T)TK3BMCellsbeforetransplant library...1
527 499
library...34 library...72
499 499
Spleen...198 BMSpine_T...180
492 470
BMFront_unsorted BMRight_O
399 386
BMRight_unsorted BM_Spine_B...195
372 346
BMSpine_G...140 BMLeft_T...172
337 334
wk12 Sac
332 313
BMRight_T BM_Right_G...194
307 304
wk9 BMSpine_unsorted
302 299
BM_CD34 BMLeft_B...131
297 292
BM_Front_unsorted BM_Right_unsorted...192
280 280
wk20...161 BM_Spine_G...197
275 270
BMSpine_B...179 BMSpine_G...181
268 263
BMFront_B...127 BMFront_B...167
258 255
BM_Spine_T...196 BM_Left_unsorted
253 252
BMRight_G...177 Spleen...186
231 213
L_wk12...10 wk22_unsorted
193 191
RightBMG_wk_24_GK wk22_T
190 185
RightBMOther_wk_24_GK FrontBMB_wk_24_GK
184 179
SpineBMG_wk_24_GK wk18_O
171 169
wk14...160 BMLeft_B...171
164 163
BMPelvis_unsorted wk22_B
151 143
BMFront_T...168 GK_wk6...2
138 127
SpineBMOther_wk_24_GK GK_wk6...74
125 119
wk11 BM_Right_T...193
117 115
BMLeft_G...173 L_wk6...9
114 113
BM_Spine_G...156 L_wk7
109 98
L_BMGrn FrontBMOther_wk_24_GK
97 92
BMSpine_O BM_Left
89 85
L_wk30...13 BMLeft_G...133
83 83
L_BloodSac L_wk43
82 78
GK_Spleen BM_Spine_O
77 73
BMPelvis_B BM_Right_O
72 71
LeftBMG_wk_24_GK BMLeft_T...132
68 67
BM_Spine_B...154 BM_Spine_T...155
65 64
BMRight_B...175 LeftBMUnsorted_wk_24_GK
64 60
L_wk6...82 FrontBMT_wk_24_GK
57 55
L_BMOther BMFront_G...129
53 53
L_wk19 SpineBMB_wk_24_GK
52 51
L_BM_B...31 Spleen...158
48 48
GK_BMLeft GK_wk12...75
47 47
GK_BM_T19 L_wk37_T
47 46
GK_BMSpine FrontBMG_wk_24_GK
46 46
GK_BMFront L_wk25
45 44
L_wk25_B L_wk37_B
44 44
SpleenSpleen_wk_24_GK L_BMBcells
43 42
R_BMGrn L_BMTcells
42 41
RightBMUnsorted_wk_24_GK L_BM_T...33
41 40
L_BloodSac_Bcells BM_Front
40 39
wk20...144 GK_wk37
38 37
R_wk12 GK_wk37_B
37 36
GK_wk7 GK_Liver
36 36
R_wk7 GK_BM_T
36 35
L_wk12...83 BM_Right_unsorted...150
35 34
GK_sac BloodT_wk_24_GK
33 33
LeftBMT_wk_24_GK GK_wk43
33 32
L_Blood_sac L_wk18...48
31 30
L_wk30...50 L_BM_G...32
30 29
wk14...143 GK_BMRight
29 28
Blood_wk_16_GK sac
28 28
GK_BM_G GK_wk12...36
27 27
GK_wk18...76 GK_wk30...78
27 26
GK_wk36_B GK_BM_B...28
26 25
Blood_wk_10_GK R_wk36_Bcells
25 24
BloodB_wk_24_GK LeftBMOther_wk_24_GK
24 24
LiverLiver_wk_24_GK wk10
24 24
GK_wk37_G GK_wk24
23 23
BloodG_wk_24_GK GK_BM_B...88
23 22
L_BloodSac_Other wk27
21 21
wk33 L_wk24...49
21 20
R_BMOther wk18_T
20 20
L_wk36Bcells RightBMT_wk_24_GK
19 19
L_wk37_G L_wk42
18 18
L_BM_G...92 GK_wk30...5
18 17
R_BMBcells L_wk18...84
17 15
GK_BM_T20 BM_Right_G...152
15 15
GK_wk25 L_wk36_Tcells
14 14
SpineBMT_wk_24_GK GK_wk12...3
14 13
L_wk12...47 L_wk36_Grn
13 13
L_BloodSac_Tcells BM_Right_T...151
13 13
GK_wk42 Blood_wk_4_GK
12 12
GK_wk18...37 GK_wk47
11 11
L_BM_T...93 LeftBMB_wk_24_GK
11 11
R_wk30 L_BM_B...91
10 10
GK_wk30...39 Blood_wk_22_GK
9 7
R_wk18 GK_wk19
6 4
L_wk25_G R_wk24
3 3
L_wk30...86 BMSpine_T...139
3 3
GK_wk25_B GK_wk25_T
2 2
GK_wk37_T L_wk25_T
2 2
BloodOther_wk_24_GK SpineBMUnsorted_wk_24_GK
2 2
wk4 GK_wk25_G
2 1
L_BloodSac_Grn L_wk24...85
1 1
RightBMB_wk_24_GK BMFront_T...128
1 1
BMFront_O BMLeft_O
1 1
BMRight_B...135 BMRight_G...136
1 1
BMSpine_B...138 wk22_G
1 1
BMFront_G...169 BMLeft_unsorted
1 1
BMPelvis_T BMPelvis_G
1 1
Liver
1 barbieQ::plotBarcodePareto(barbieQ = xenoHSPC)Warning: Removed 10 rows containing missing values or values outside the scale range
(`geom_bar()`).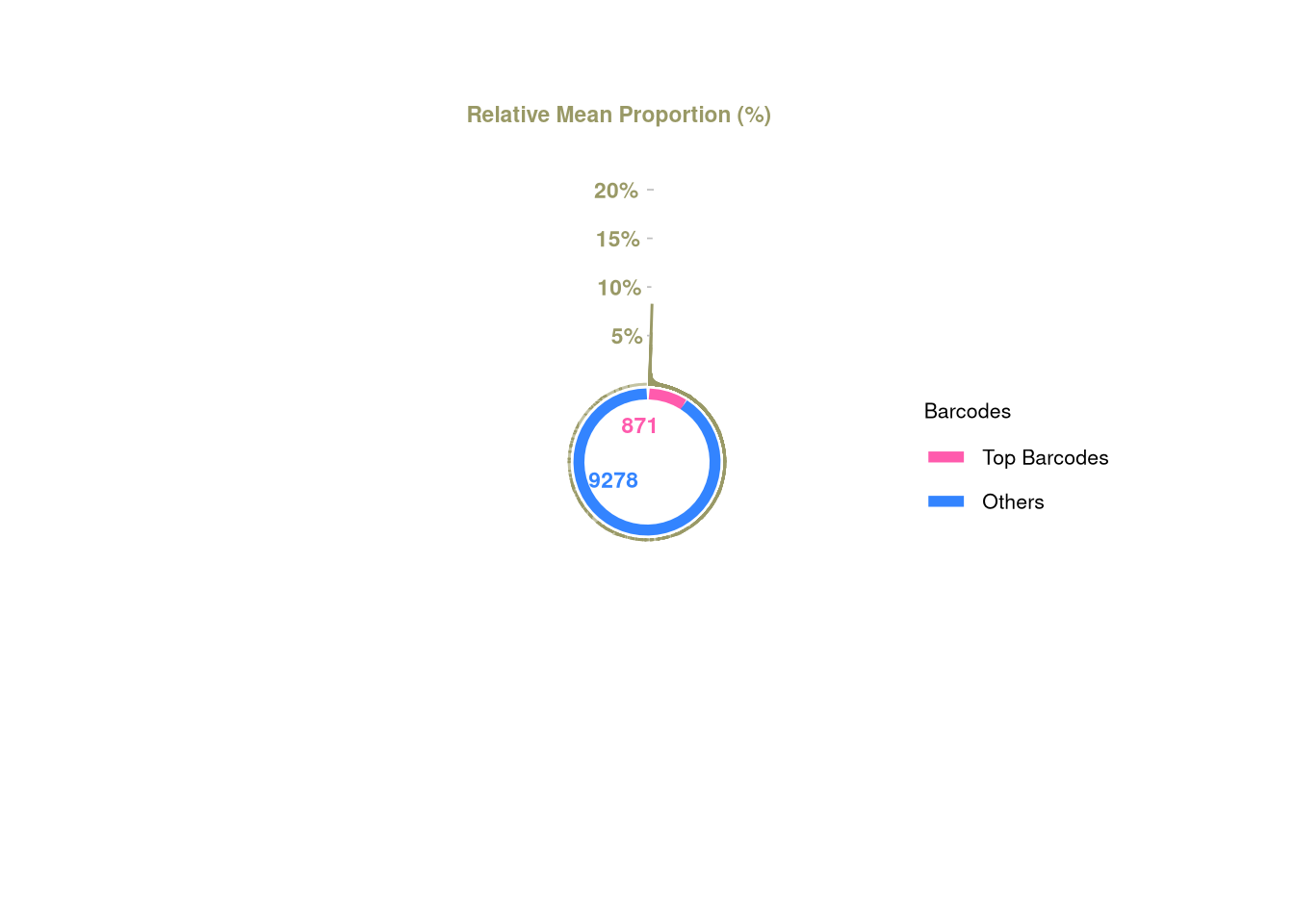
barbieQ::plotBarcodeSankey(barbieQ = xenoHSPC)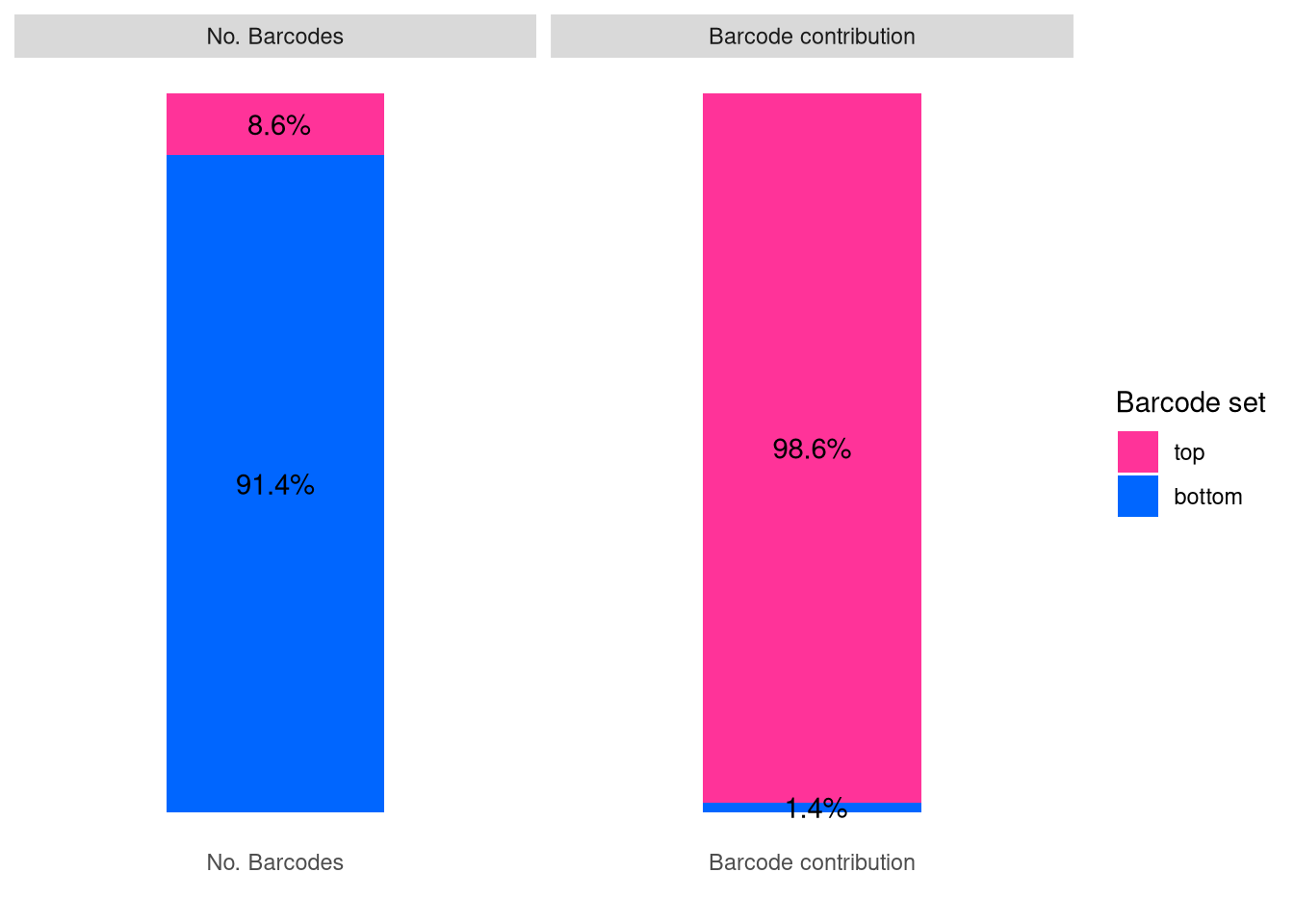
8 Cluster barcodes
8.1 FS1C: pairwise correlation
plotBarcodePairCorrelation(
barbieQ = xenoHSPC[rowData(xenoHSPC)$isTopBarcode$isTop,],
propThresh = 0.005, transformation = "none") +
theme(legend.position = "top") +
labs(title = "HSPC xenograft data") -> fs1cprocessing Barcode pairwise pearson correlation on propotion (none transformation).fs1cWarning: The dot-dot notation (`..y..`) was deprecated in ggplot2 3.4.0.
ℹ Please use `after_stat(y)` instead.
ℹ The deprecated feature was likely used in the barbieQ package.
Please report the issue at <https://github.com/Oshlack/barbieQ>.
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_bar()`).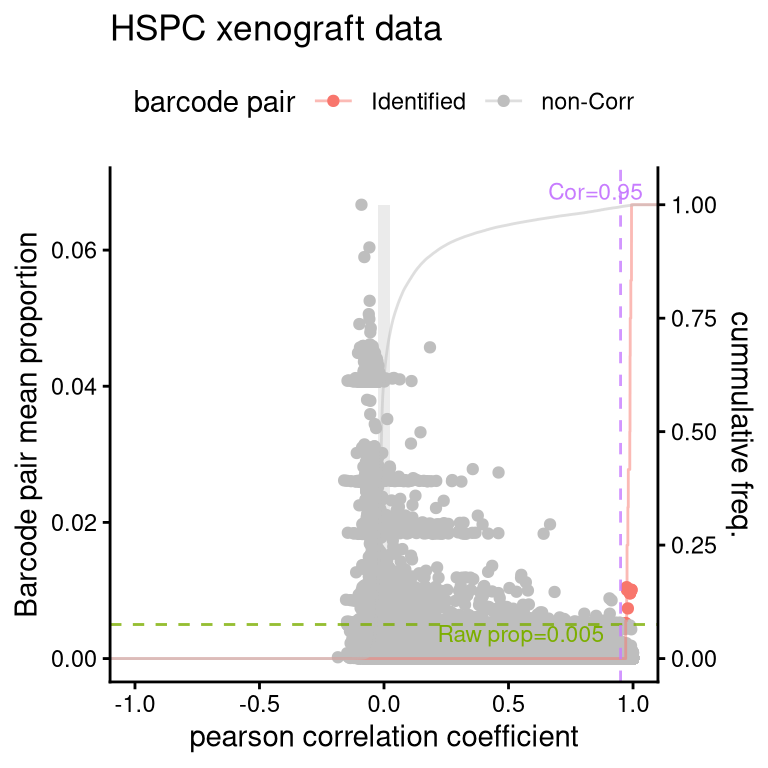
8.2 FS1F: clusters barcodes
## cluster barcoes
xenoHSPC_top0 <- barbieQ::clusterCorrelatingBarcodes(
barbieQ = xenoHSPC[rowData(xenoHSPC)$isTopBarcode$isTop,],
transformation = "none", propThresh = 0.005)processing Barcode pairwise pearson correlation on propotion (none transformation).identified 3 clusters, including 9 Barcodes.p_cluster_list <- barbieQ::inspectCorrelatingBarcodes(xenoHSPC_top0)
p_cluster_list$p_cluster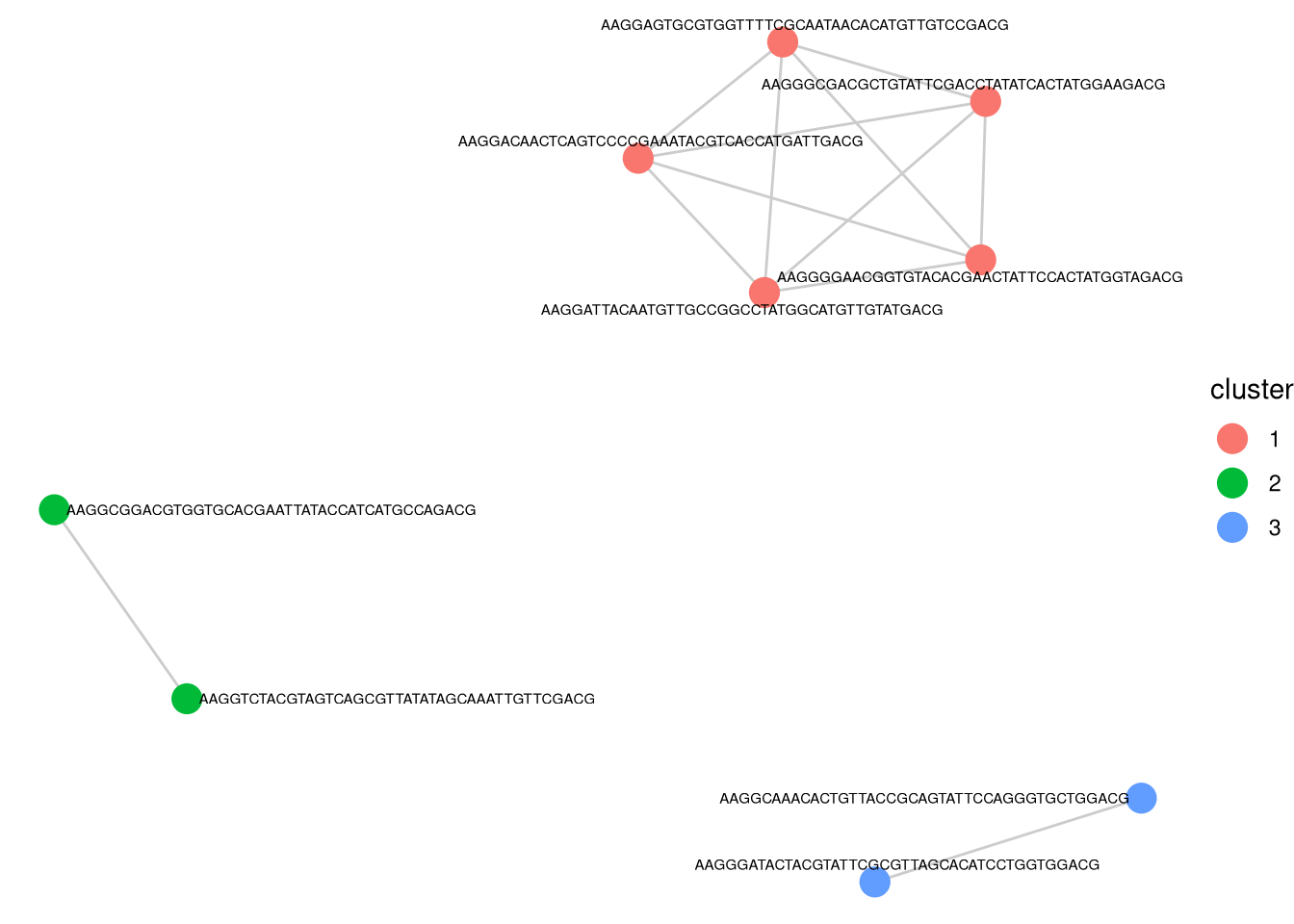
$p_cluster_size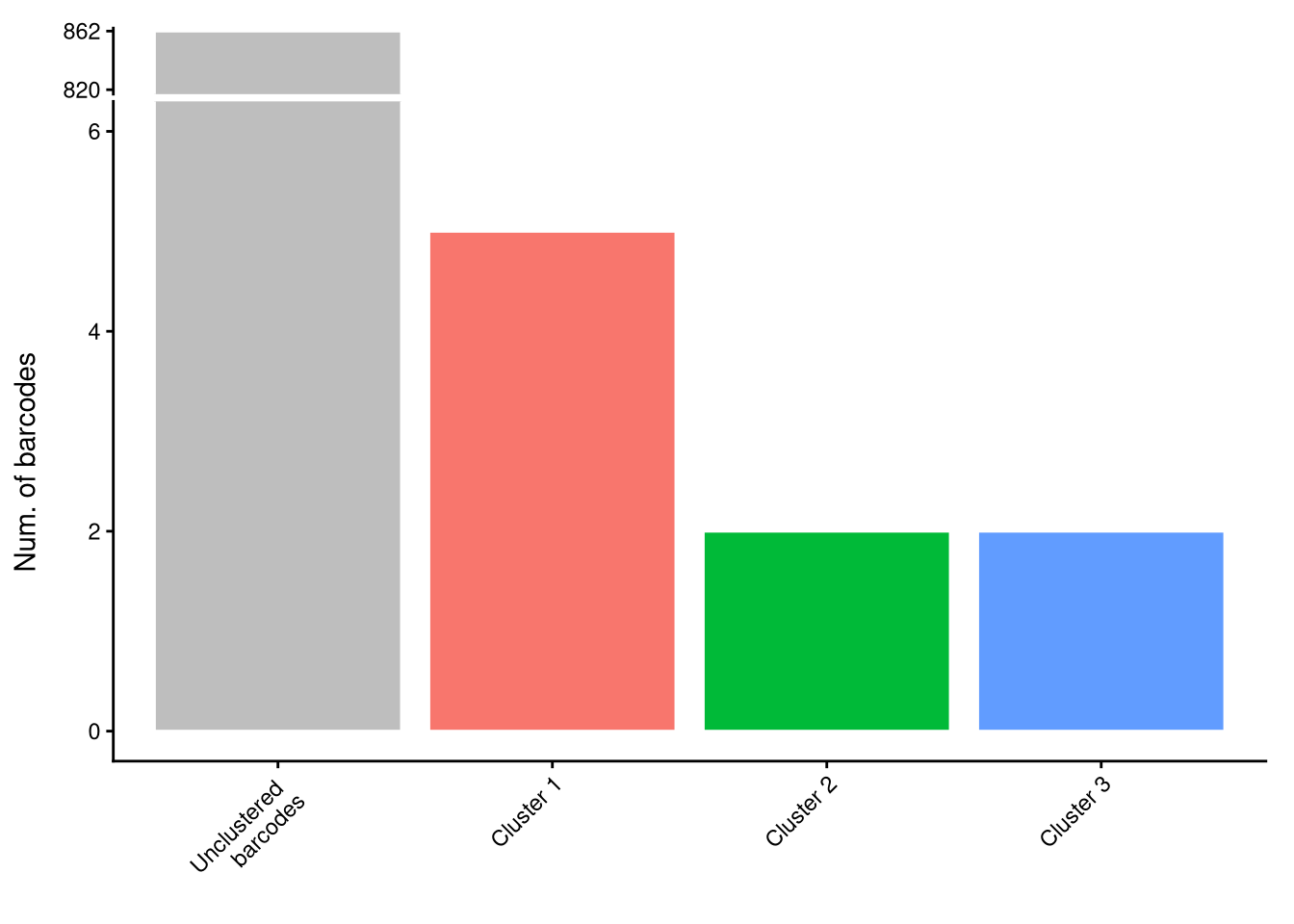
$p_cluster_prop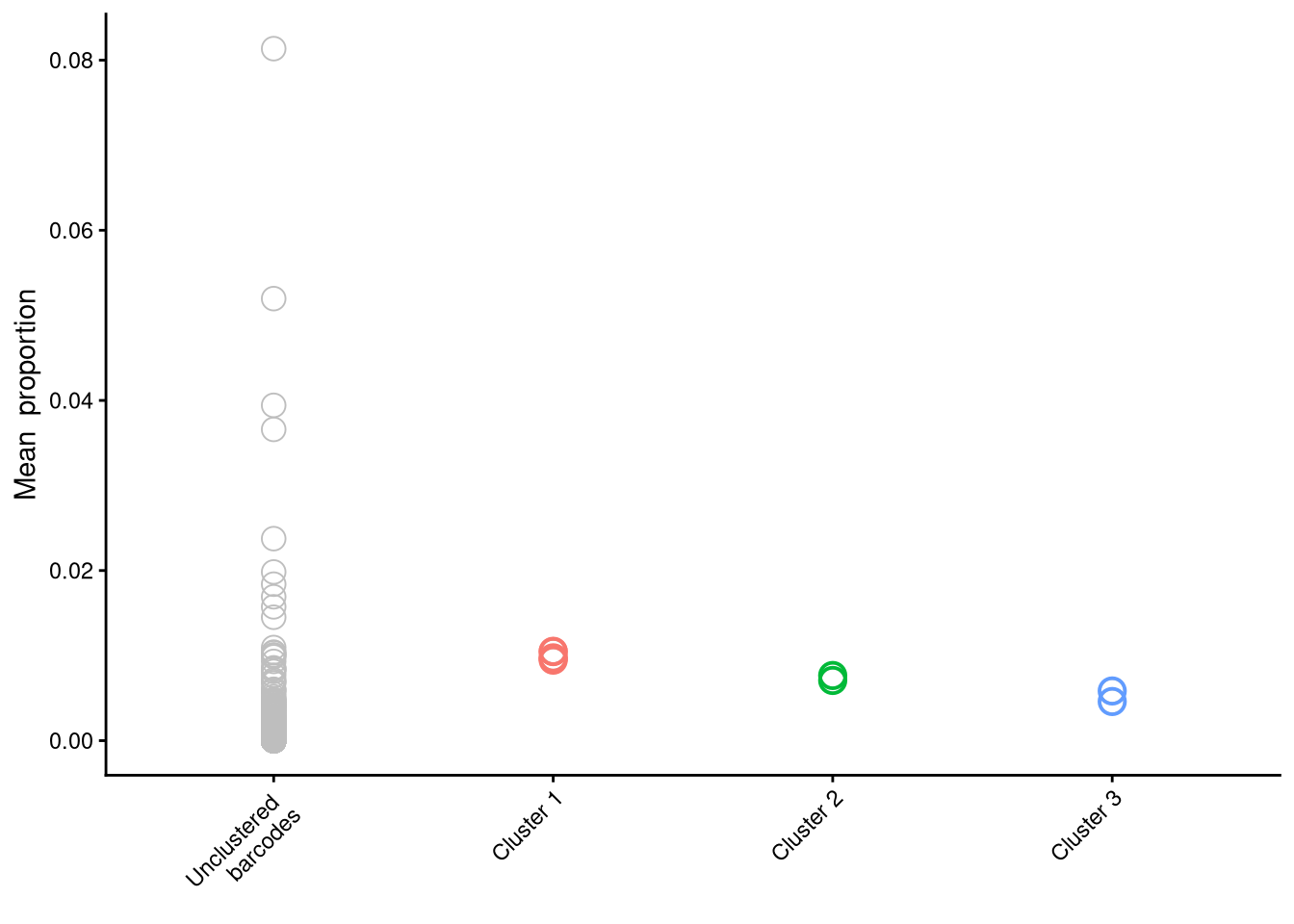
layer_index <- which(sapply(p_cluster_list$p_cluster$layers, function(x) inherits(x$geom, "GeomTextRepel")))
## update the size for that layer
p_cluster_list$p_cluster$layers[[layer_index]]$aes_params$size <- 1.4 # new size
fs1f <- p_cluster_list$p_cluster + theme(legend.position = "none")
fs1f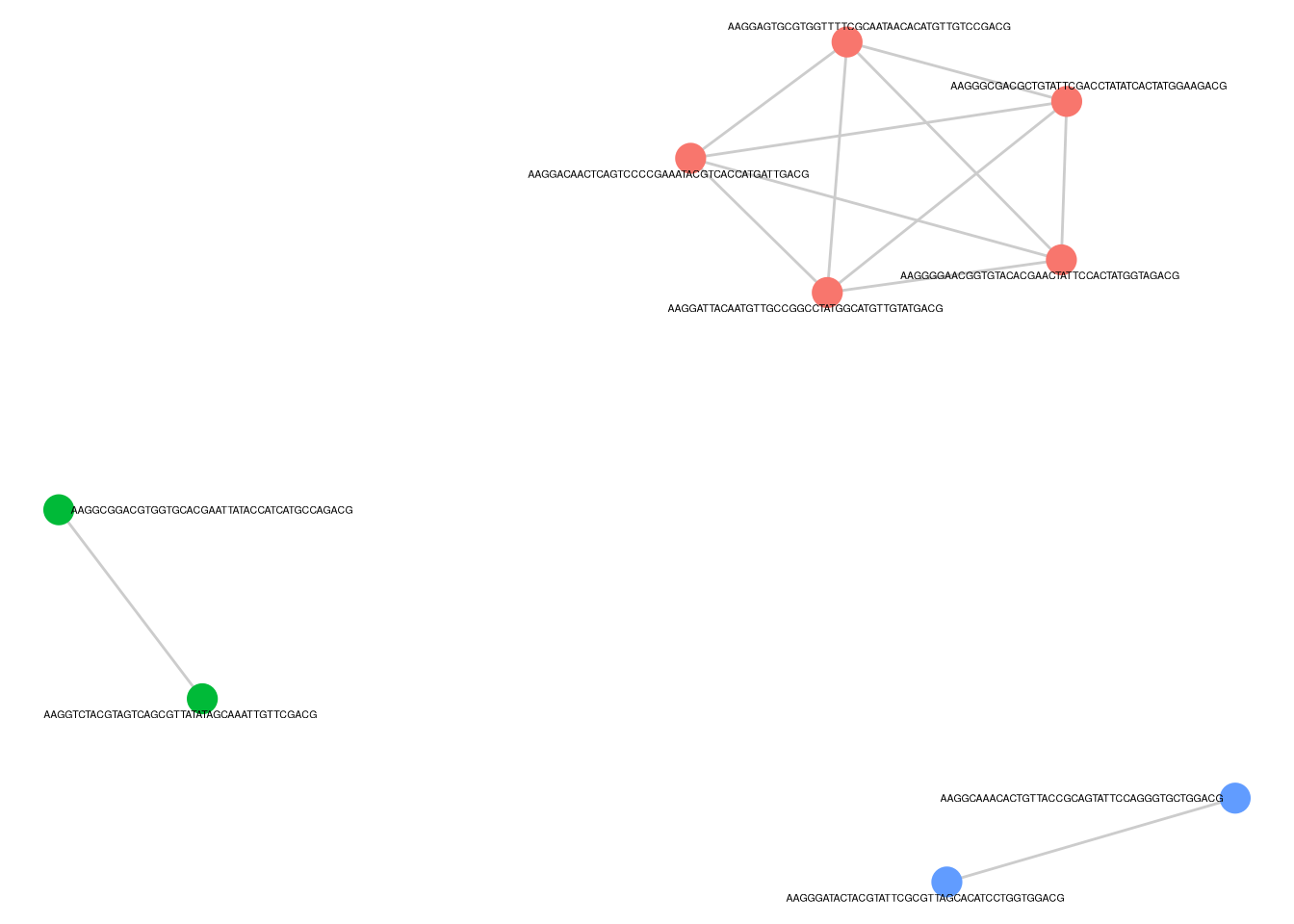
8.3 merge clusters
xenoHSPC_merged <- barbieQ::mergeCorrelatingBarcodes(barbieQ_clustered = xenoHSPC_top0, barbieQ_toMerge = xenoHSPC)[1] "printing removed barcodes: AAGGGGAACGGTGTACACGAACTATTCCACTATGGTAGACG"
[2] "printing removed barcodes: AAGGATTACAATGTTGCCGGCCTATGGCATGTTGTATGACG"
[3] "printing removed barcodes: AAGGGCGACGCTGTATTCGACCTATATCACTATGGAAGACG"
[4] "printing removed barcodes: AAGGTCTACGTAGTCAGCGTTATATAGCAAATTGTTCGACG"
[5] "printing removed barcodes: AAGGGATACTACGTATTCGCGTTAGCACATCCTGGTGGACG"
[6] "printing removed barcodes: AAGGACAACTCAGTCCCCGAAATACGTCACCATGATTGACG"sample names set up in line with `object`continuing with missing `factorColors`.!! re-computing barcode proportion, CPM, rank... from the selected barcodes.dim(xenoHSPC_merged)[1] 10143 199xenoHSPC_merged@metadata$predicted_barcode_clustersIGRAPH e9531bf UN-- 9 12 --
+ attr: name (v/c)
+ edges from e9531bf (vertex names):
[1] AAGGGGAACGGTGTACACGAACTATTCCACTATGGTAGACG--AAGGATTACAATGTTGCCGGCCTATGGCATGTTGTATGACG
[2] AAGGGGAACGGTGTACACGAACTATTCCACTATGGTAGACG--AAGGAGTGCGTGGTTTTCGCAATAACACATGTTGTCCGACG
[3] AAGGATTACAATGTTGCCGGCCTATGGCATGTTGTATGACG--AAGGAGTGCGTGGTTTTCGCAATAACACATGTTGTCCGACG
[4] AAGGGGAACGGTGTACACGAACTATTCCACTATGGTAGACG--AAGGGCGACGCTGTATTCGACCTATATCACTATGGAAGACG
[5] AAGGATTACAATGTTGCCGGCCTATGGCATGTTGTATGACG--AAGGGCGACGCTGTATTCGACCTATATCACTATGGAAGACG
[6] AAGGAGTGCGTGGTTTTCGCAATAACACATGTTGTCCGACG--AAGGGCGACGCTGTATTCGACCTATATCACTATGGAAGACG
[7] AAGGCGGACGTGGTGCACGAATTATACCATCATGCCAGACG--AAGGTCTACGTAGTCAGCGTTATATAGCAAATTGTTCGACG
+ ... omitted several edgesxenoHSPC_merged@metadata$removed_barcodes[1] "AAGGGGAACGGTGTACACGAACTATTCCACTATGGTAGACG"
[2] "AAGGATTACAATGTTGCCGGCCTATGGCATGTTGTATGACG"
[3] "AAGGGCGACGCTGTATTCGACCTATATCACTATGGAAGACG"
[4] "AAGGTCTACGTAGTCAGCGTTATATAGCAAATTGTTCGACG"
[5] "AAGGGATACTACGTATTCGCGTTAGCACATCCTGGTGGACG"
[6] "AAGGACAACTCAGTCCCCGAAATACGTCACCATGATTGACG"9 Tag top barcode after collapsing
9.1 FS1H,K: inspect top/bottom
xenoHSPC_merged <- tagTopBarcodes(
barbieQ = xenoHSPC_merged,
# activeSamples = xenoHSPC$sampleMetadata$Sample != "library",
nSampleThreshold = 3)
barbieQ::plotBarcodePareto(barbieQ = xenoHSPC_merged) +
ylim(-8, 12) +
annotate("text", x = c(pi * 0.05), y = c(12),
label = c("Relative proportion %"), color = "#999966",
size = 3, angle = 0, fontface = "bold", hjust = 1) -> fs1hScale for y is already present.
Adding another scale for y, which will replace the existing scale.fs1hWarning: Removed 10 rows containing missing values or values outside the scale range
(`geom_bar()`).Warning: Removed 1 row containing missing values or values outside the scale range
(`geom_segment()`).
Removed 1 row containing missing values or values outside the scale range
(`geom_segment()`).Warning: Removed 3 rows containing missing values or values outside the scale range
(`geom_text()`).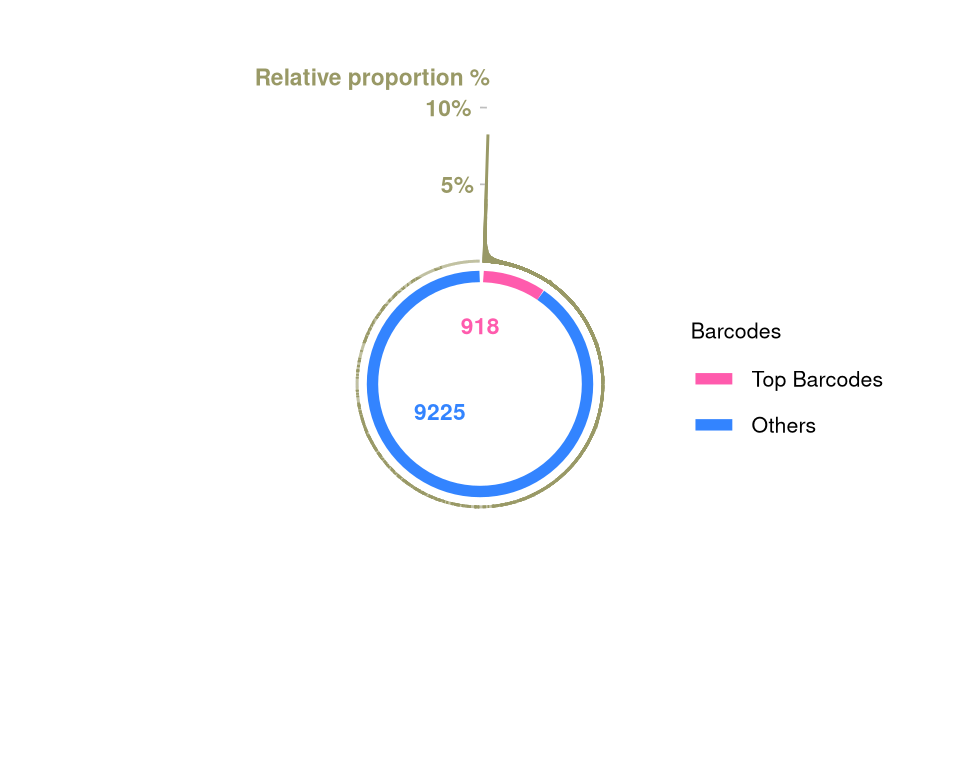
barbieQ::plotBarcodeSankey(barbieQ = xenoHSPC_merged) +
theme(legend.position = "top") -> fs1k
fs1k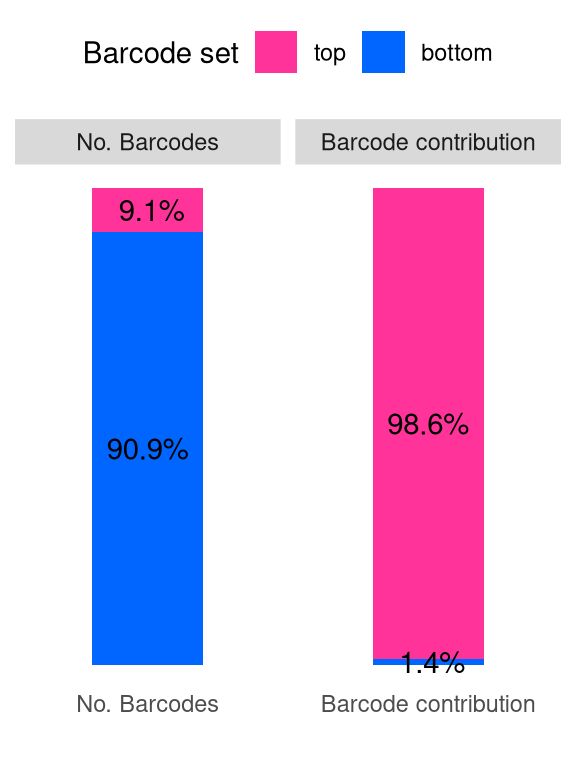
9.2 Filter “top” barcodes
xenoHSPC_top <- xenoHSPC_merged[rowData(xenoHSPC_merged)$isTopBarcode$isTop,]9.3 Heatmap
## pre-filtering
Heatmap(name = "asin-sqrt prop.",
matrix = asin(sqrt(assays(xenoHSPC_merged)$proportion)),
row_title = paste0(nrow(xenoHSPC_merged), " Barcodes"),
show_row_names = FALSE, show_column_names = FALSE)The automatically generated colors map from the 1^st and 99^th of the
values in the matrix. There are outliers in the matrix whose patterns
might be hidden by this color mapping. You can manually set the color
to `col` argument.
Use `suppressMessages()` to turn off this message.`use_raster` is automatically set to TRUE for a matrix with more than
2000 rows. You can control `use_raster` argument by explicitly setting
TRUE/FALSE to it.
Set `ht_opt$message = FALSE` to turn off this message.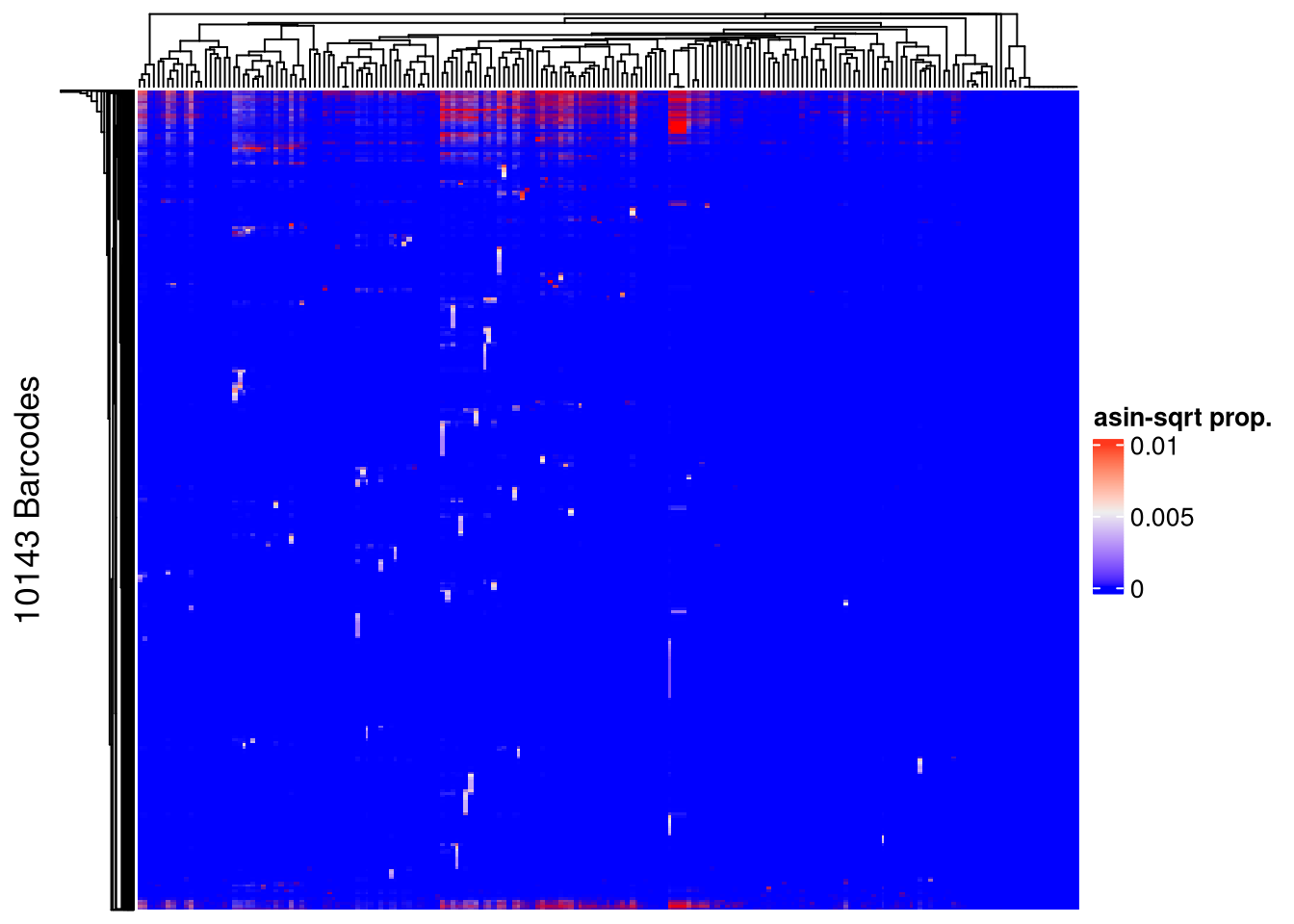
## post-filtering
Heatmap(name = "asin-sqrt prop.",
matrix = asin(sqrt(assays(xenoHSPC_top)$proportion)),
row_title = paste0(nrow(xenoHSPC_top), " Barcodes"),
show_row_names = FALSE, show_column_names = FALSE)The automatically generated colors map from the 1^st and 99^th of the
values in the matrix. There are outliers in the matrix whose patterns
might be hidden by this color mapping. You can manually set the color
to `col` argument.
Use `suppressMessages()` to turn off this message.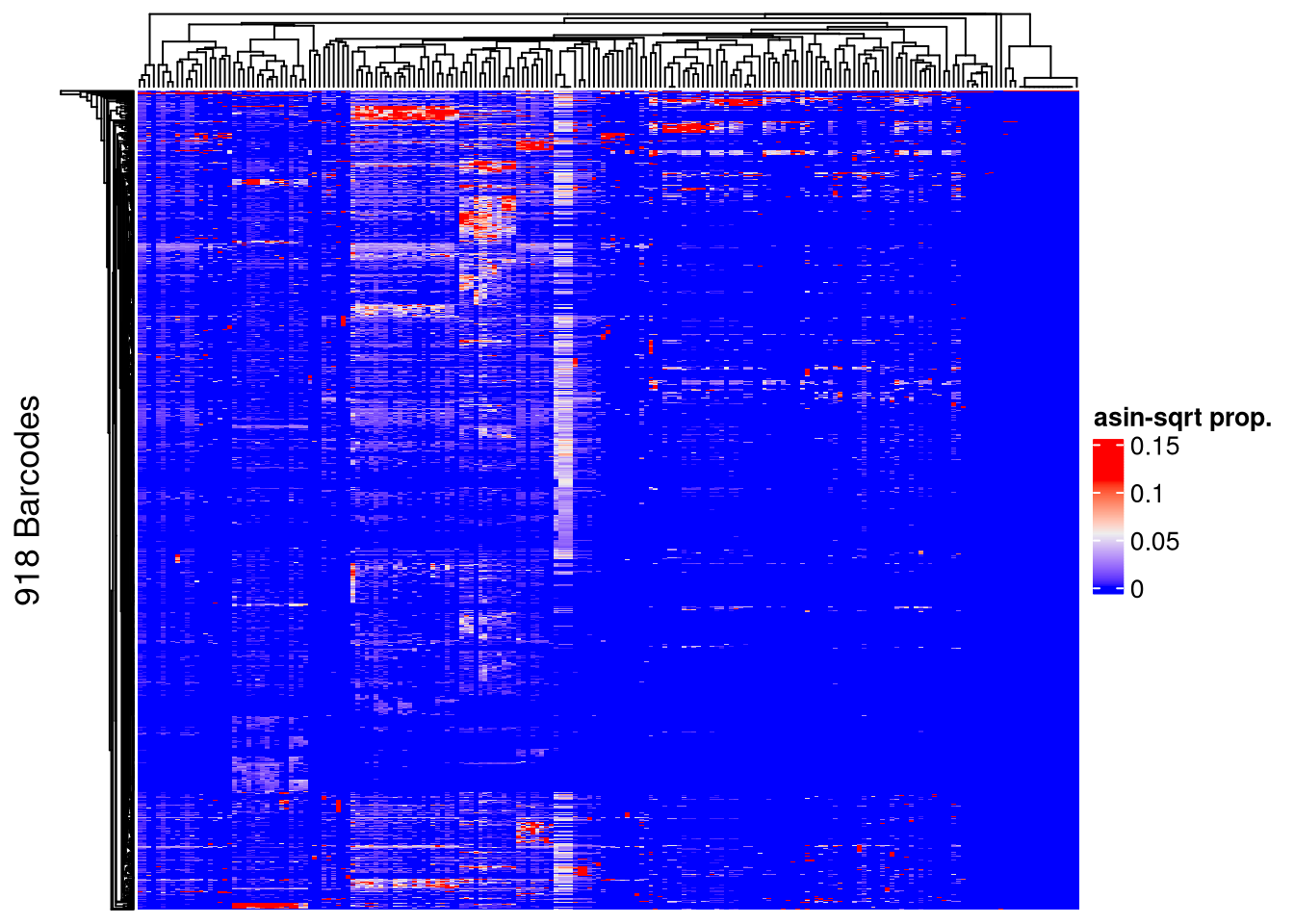
10 Save rds
save(xenoHSPC, xenoHSPC_merged, xenoHSPC_top, file = "output/xenoHSPC_barbieQ.rda")11 Top10
tagTop10 <- function(col) {
ranks <- rank(-col)
# summary(ranks)
isTop <- ranks <= 10
# summary(isTop)
return(isTop)
}
tester <- tagTop10(assays(xenoHSPC_merged)$CPM[,1])
summary(tester) Mode FALSE TRUE
logical 10133 10 colTags <- lapply(as.data.frame(assays(xenoHSPC_merged)$CPM),
function(col) tagTop10(col))
colTagMat <- do.call(cbind, colTags)
colTagVec <- rowSums(colTagMat[,xenoHSPC_merged$sampleMetadata$Sample != "library"]) >= 1
summary(colTagVec) Mode FALSE TRUE
logical 9734 409 xenoHSPC_merged_top10 <- xenoHSPC_merged
rowData(xenoHSPC_merged_top10)$isTopBarcode <- DataFrame(isTop = colTagVec)
summary(rowData(xenoHSPC_merged_top10)$isTopBarcode$isTop) Mode FALSE TRUE
logical 9734 409 barbieQ::plotBarcodePareto(barbieQ = xenoHSPC_merged_top10)Warning: Removed 10 rows containing missing values or values outside the scale range
(`geom_bar()`).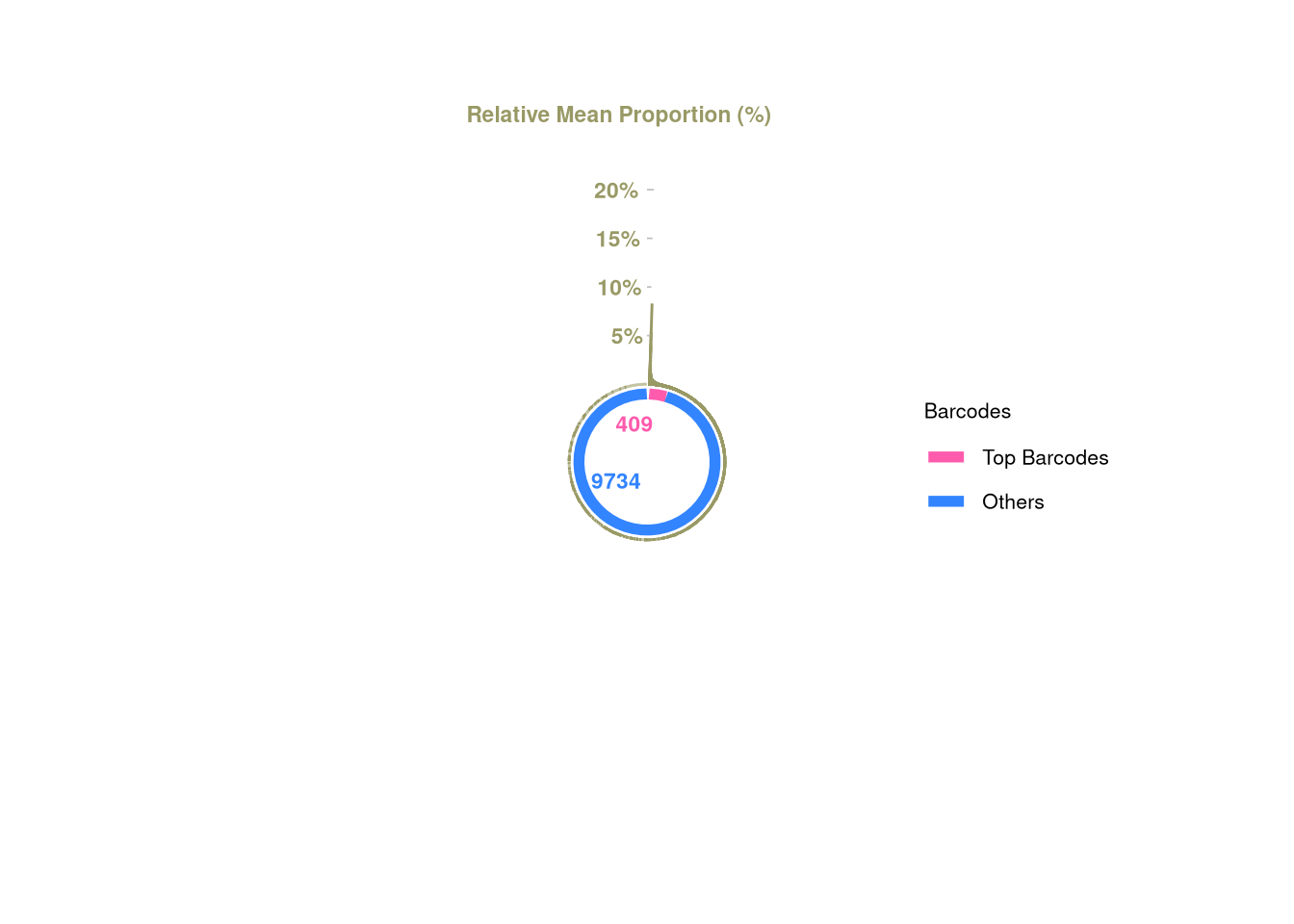
barbieQ::plotBarcodeSankey(barbieQ = xenoHSPC_merged_top10)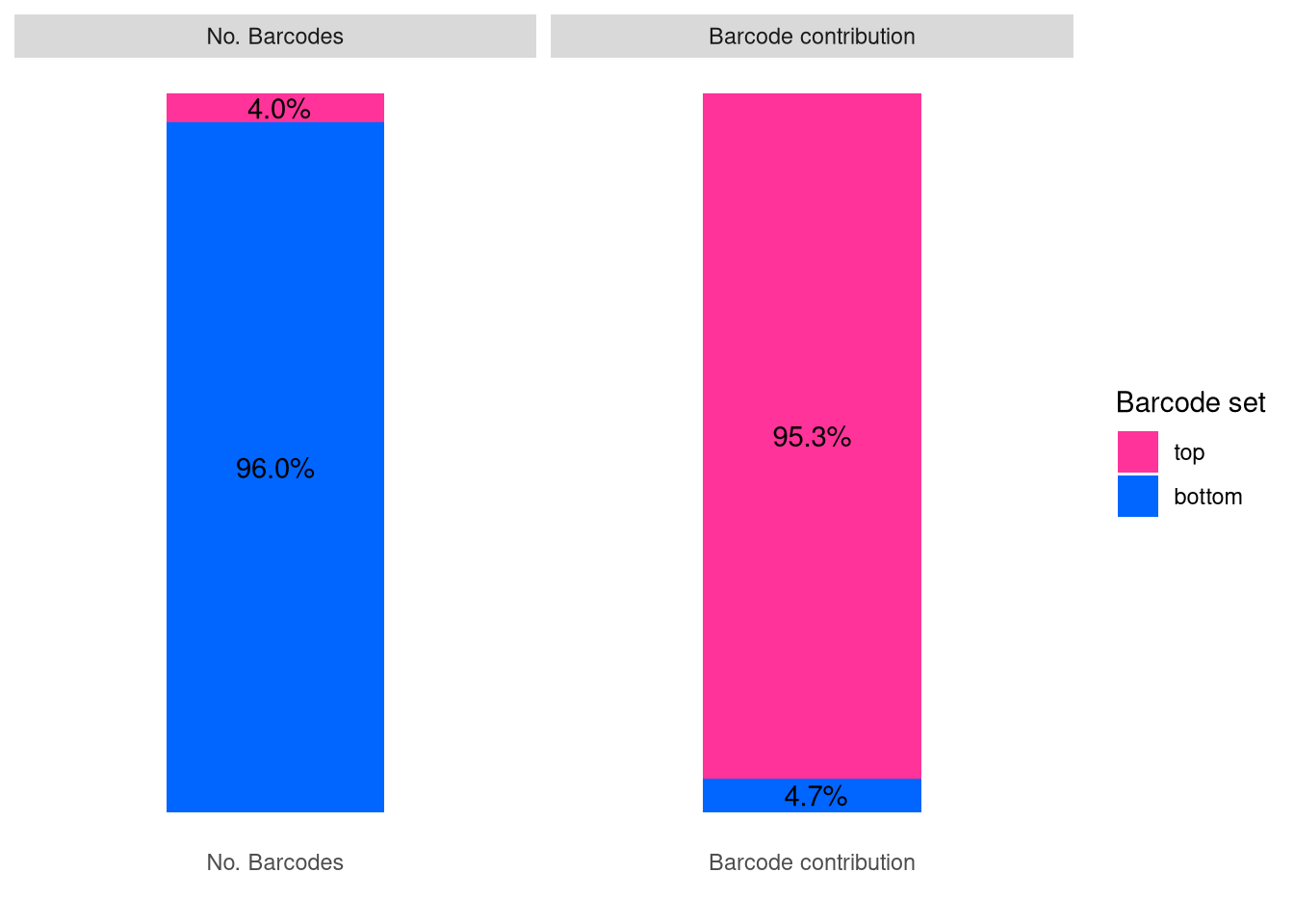
12 Save FigureS1-AML
layout = "
C
F
H
K
"
fs1_xeno <- (
wrap_elements(fs1c + theme(plot.margin = unit(c(0,0,0,0), "line"))) +
wrap_elements(fs1f + theme(plot.margin = unit(rep(0,4), "cm"))) +
wrap_elements(fs1h + theme(plot.margin = unit(rep(0,4), "cm"))) +
wrap_elements(fs1k + theme(plot.margin = unit(rep(0,4), "cm")))
) +
plot_layout(design = layout) +
plot_annotation(tag_levels = list(c("C","F","H", "K"))) &
theme(
plot.tag = element_text(size = 20, face = "bold", family = "arial"),
axis.title = element_text(size = 17),
axis.text = element_text(size = 12),
legend.title = element_text(size = 13),
legend.text = element_text(size = 11))
fs1_xenoWarning: Removed 2 rows containing missing values or values outside the scale range
(`geom_bar()`).Warning: Removed 10 rows containing missing values or values outside the scale range
(`geom_bar()`).Warning: Removed 1 row containing missing values or values outside the scale range
(`geom_segment()`).
Removed 1 row containing missing values or values outside the scale range
(`geom_segment()`).Warning: Removed 3 rows containing missing values or values outside the scale range
(`geom_text()`).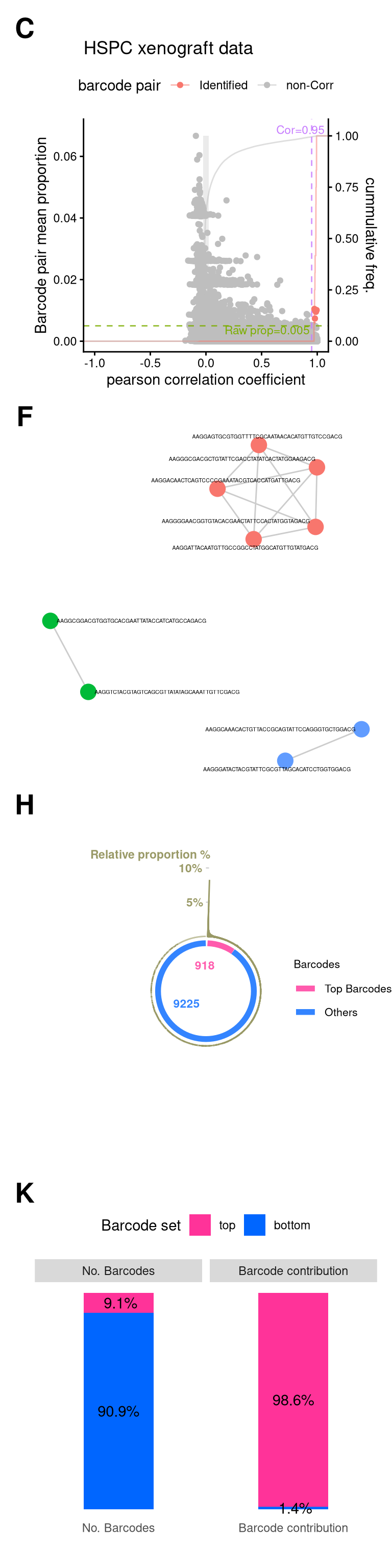
ggsave(
filename = "output/fs1_xeno.png",
plot = fs1_xeno,
width = 4,
height = 16,
units = "in", # for Rmd r chunk fig size, unit default to inch
dpi = 350
)Warning: Removed 2 rows containing missing values or values outside the scale range
(`geom_bar()`).Warning: Removed 10 rows containing missing values or values outside the scale range
(`geom_bar()`).Warning: Removed 1 row containing missing values or values outside the scale range
(`geom_segment()`).
Removed 1 row containing missing values or values outside the scale range
(`geom_segment()`).Warning: Removed 3 rows containing missing values or values outside the scale range
(`geom_text()`).Saving this figure in fs1_xenoHSPC
sessionInfo()R version 4.5.0 (2025-04-11)
Platform: x86_64-pc-linux-gnu
Running under: Red Hat Enterprise Linux 9.6 (Plow)
Matrix products: default
BLAS/LAPACK: FlexiBLAS OPENBLAS-OPENMP; LAPACK version 3.9.0
locale:
[1] LC_CTYPE=en_AU.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_AU.UTF-8 LC_COLLATE=en_AU.UTF-8
[5] LC_MONETARY=en_AU.UTF-8 LC_MESSAGES=en_AU.UTF-8
[7] LC_PAPER=en_AU.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_AU.UTF-8 LC_IDENTIFICATION=C
time zone: Australia/Melbourne
tzcode source: system (glibc)
attached base packages:
[1] stats4 grid stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] barbieQ_1.1.3 devtools_2.4.6
[3] usethis_3.2.1 SEtools_1.22.0
[5] sechm_1.16.0 SummarizedExperiment_1.38.1
[7] Biobase_2.68.0 GenomicRanges_1.60.0
[9] GenomeInfoDb_1.44.3 IRanges_2.42.0
[11] S4Vectors_0.48.0 BiocGenerics_0.54.0
[13] generics_0.1.4 MatrixGenerics_1.20.0
[15] matrixStats_1.5.0 edgeR_4.6.3
[17] limma_3.64.3 ComplexHeatmap_2.24.1
[19] ggVennDiagram_1.5.4 scales_1.4.0
[21] patchwork_1.3.2 ggplot2_4.0.0
[23] knitr_1.50 tibble_3.3.0
[25] tidyr_1.3.1 dplyr_1.1.4
[27] magrittr_2.0.4 readxl_1.4.5
[29] workflowr_1.7.2
loaded via a namespace (and not attached):
[1] splines_4.5.0 later_1.4.4 ggplotify_0.1.3
[4] cellranger_1.1.0 polyclip_1.10-7 rpart_4.1.24
[7] XML_3.99-0.20 lifecycle_1.0.4 Rdpack_2.6.4
[10] formula.tools_1.7.1 doParallel_1.0.17 rprojroot_2.1.1
[13] processx_3.8.6 lattice_0.22-6 MASS_7.3-65
[16] backports_1.5.0 openxlsx_4.2.8.1 sass_0.4.10
[19] rmarkdown_2.30 jquerylib_0.1.4 yaml_2.3.10
[22] remotes_2.5.0 httpuv_1.6.16 zip_2.3.3
[25] sessioninfo_1.2.3 pkgbuild_1.4.8 minqa_1.2.8
[28] DBI_1.2.3 RColorBrewer_1.1-3 abind_1.4-8
[31] pkgload_1.4.1 Rtsne_0.17 purrr_1.1.0
[34] ggraph_2.2.2 nnet_7.3-20 yulab.utils_0.2.1
[37] tweenr_2.0.3 rappdirs_0.3.3 git2r_0.36.2
[40] sva_3.56.0 circlize_0.4.16 seriation_1.5.8
[43] GenomeInfoDbData_1.2.14 ggrepel_0.9.6 genefilter_1.90.0
[46] pheatmap_1.0.13 annotate_1.86.1 codetools_0.2-20
[49] DelayedArray_0.34.1 ggforce_0.5.0 tidyselect_1.2.1
[52] shape_1.4.6.1 aplot_0.2.9 UCSC.utils_1.4.0
[55] farver_2.1.2 lme4_1.1-37 viridis_0.6.5
[58] TSP_1.2.6 jsonlite_2.0.0 GetoptLong_1.0.5
[61] mitml_0.4-5 ellipsis_0.3.2 tidygraph_1.3.1
[64] ggbreak_0.1.6 randomcoloR_1.1.0.1 survival_3.8-3
[67] iterators_1.0.14 systemfonts_1.3.1 foreach_1.5.2
[70] tools_4.5.0 ragg_1.5.0 Rcpp_1.1.0
[73] glue_1.8.0 pan_1.9 gridExtra_2.3
[76] SparseArray_1.8.1 xfun_0.53 mgcv_1.9-1
[79] DESeq2_1.48.2 logistf_1.26.1 ca_0.71.1
[82] withr_3.0.2 fastmap_1.2.0 boot_1.3-31
[85] callr_3.7.6 digest_0.6.37 R6_2.6.1
[88] gridGraphics_0.5-1 textshaping_1.0.3 mice_3.18.0
[91] colorspace_2.1-2 RSQLite_2.4.5 data.table_1.17.8
[94] graphlayouts_1.2.2 httr_1.4.7 S4Arrays_1.8.1
[97] whisker_0.4.1 pkgconfig_2.0.3 gtable_0.3.6
[100] blob_1.2.4 registry_0.5-1 S7_0.2.0
[103] XVector_0.48.0 htmltools_0.5.8.1 clue_0.3-66
[106] png_0.1-8 reformulas_0.4.1 ggfun_0.2.0
[109] rstudioapi_0.17.1 rjson_0.2.23 nloptr_2.2.1
[112] nlme_3.1-168 curl_7.0.0 cachem_1.1.0
[115] GlobalOptions_0.1.2 stringr_1.5.2 operator.tools_1.6.3
[118] parallel_4.5.0 AnnotationDbi_1.70.0 pillar_1.11.1
[121] vctrs_0.6.5 promises_1.3.3 jomo_2.7-6
[124] xtable_1.8-4 cluster_2.1.8.1 evaluate_1.0.5
[127] magick_2.9.0 cli_3.6.5 locfit_1.5-9.12
[130] compiler_4.5.0 rlang_1.1.6 crayon_1.5.3
[133] labeling_0.4.3 ps_1.9.1 forcats_1.0.1
[136] getPass_0.2-4 fs_1.6.6 stringi_1.8.7
[139] viridisLite_0.4.2 BiocParallel_1.42.2 Biostrings_2.76.0
[142] glmnet_4.1-10 V8_8.0.1 Matrix_1.7-3
[145] bit64_4.6.0-1 KEGGREST_1.48.1 statmod_1.5.0
[148] rbibutils_2.3 broom_1.0.10 igraph_2.1.4
[151] memoise_2.0.1 bslib_0.9.0 bit_4.6.0