barbieQ_paper Supplementary 2: Type I error
rate using AML data
Liyang Fei
Initiated: 2025-04-04 Rendered: 2026-01-07
Last updated: 2026-01-07
Checks: 5 2
Knit directory: public_barcode_count/
This reproducible R Markdown analysis was created with workflowr (version 1.7.2). The Checks tab describes the reproducibility checks that were applied when the results were created. The Past versions tab lists the development history.
The R Markdown is untracked by Git. To know which version of the R
Markdown file created these results, you’ll want to first commit it to
the Git repo. If you’re still working on the analysis, you can ignore
this warning. When you’re finished, you can run
wflow_publish to commit the R Markdown file and build the
HTML.
The global environment had objects present when the code in the R
Markdown file was run. These objects can affect the analysis in your R
Markdown file in unknown ways. For reproduciblity it’s best to always
run the code in an empty environment. Use wflow_publish or
wflow_build to ensure that the code is always run in an
empty environment.
The following objects were defined in the global environment when these results were created:
| Name | Class | Size |
|---|---|---|
| module | function | 5.6 Kb |
The command set.seed(20250112) was run prior to running
the code in the R Markdown file. Setting a seed ensures that any results
that rely on randomness, e.g. subsampling or permutations, are
reproducible.
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
Nice! There were no cached chunks for this analysis, so you can be confident that you successfully produced the results during this run.
Great job! Using relative paths to the files within your workflowr project makes it easier to run your code on other machines.
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility.
The results in this page were generated with repository version 34f5894. See the Past versions tab to see a history of the changes made to the R Markdown and HTML files.
Note that you need to be careful to ensure that all relevant files for
the analysis have been committed to Git prior to generating the results
(you can use wflow_publish or
wflow_git_commit). workflowr only checks the R Markdown
file, but you know if there are other scripts or data files that it
depends on. Below is the status of the Git repository when the results
were generated:
Ignored files:
Ignored: .RData
Ignored: .Rhistory
Ignored: .Rproj.user/
Ignored: public_barcode_count.Rproj
Untracked files:
Untracked: analysis/barbieQ_paper_FigureS1_AML.Rmd
Untracked: analysis/barbieQ_paper_FigureS1_Mixture.Rmd
Untracked: analysis/barbieQ_paper_FigureS1_xenoHSPC.Rmd
Untracked: analysis/barbieQ_paper_FigureS2_AML.Rmd
Untracked: analysis/barbieQ_paper_FigureS2_xenoHSPC.Rmd
Untracked: output/AML_barbieQ.rda
Untracked: output/AML_negative_simulation.rda
Untracked: output/fs1_aml.png
Untracked: output/fs1_mixture.png
Untracked: output/fs1_xeno.png
Untracked: output/fs1a_knowncluster.png
Untracked: output/fs2_aml.png
Untracked: output/fs2_xeno.png
Untracked: output/xenoC21_negative_simulation.rda
Untracked: output/xenoC22_negative_simulation.rda
Untracked: output/xenoC23_negative_simulation.rda
Untracked: output/xenoHSPC_barbieQ.rda
Unstaged changes:
Modified: analysis/F3_simulation.R
Modified: analysis/barbieQ_paper_Figure2.Rmd
Deleted: analysis/barbieQ_paper_S1.Rmd
Modified: analysis/index.Rmd
Modified: data/BelderbosME/README.md
Modified: output/f2.png
Modified: output/monkeyHSPC_barbieQ.rda
Modified: output/monkeyHSPC_raw_barbieQ.rda
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.
There are no past versions. Publish this analysis with
wflow_publish() to start tracking its development.
Links to Type I error rate assessment using other datasets in the barbieQ paper:
1 Load Dependencies
library(readxl)
library(magrittr)
library(dplyr)
library(tidyr) # for pivot_longer
library(tibble) # for rownames_to _column
library(knitr) # for kable()
library(ggplot2)
library(patchwork)
library(scales)
library(ggVennDiagram)
library(ComplexHeatmap)
library(limma)
library(edgeR)
library(SummarizedExperiment)
library(SEtools)
library(S4Vectors)
library(devtools)
# devtools::install_github("DaneVass/bartools", dependencies = TRUE, force = TRUE)
# library(bartools)
source("analysis/plotBarcodeHistogram.R") ## accommodated from bartools::plotBarcodehistogram
source("analysis/F3_simulation.R") ## for negative simulation
source("analysis/ggplot_theme.R") ## setting ggplot theme2 Install
barbieQ
Installing the latest devel version of barbieQ from
GitHub.
if (!requireNamespace("barbieQ", quietly = TRUE)) {
remotes::install_github("Oshlack/barbieQ")
}Warning: replacing previous import 'data.table::first' by 'dplyr::first' when
loading 'barbieQ'Warning: replacing previous import 'data.table::last' by 'dplyr::last' when
loading 'barbieQ'Warning: replacing previous import 'data.table::between' by 'dplyr::between'
when loading 'barbieQ'Warning: replacing previous import 'dplyr::as_data_frame' by
'igraph::as_data_frame' when loading 'barbieQ'Warning: replacing previous import 'dplyr::groups' by 'igraph::groups' when
loading 'barbieQ'Warning: replacing previous import 'dplyr::union' by 'igraph::union' when
loading 'barbieQ'Registered S3 method overwritten by 'formula.tools':
method from
as.character.formula openxlsxlibrary(barbieQ)Check the version of barbieQ.
packageVersion("barbieQ")[1] '1.1.3'3 Set seeds
set.seed(2025)5 Samples level QC
5.1 PCA
pca_results <- (assays(AML_top)$proportion) %>% sqrt() %>% asin() %>%
t() %>%
prcomp()
var_pct <- pca_results$sdev^2 / sum(pca_results$sdev^2) * 100
var_pct [1] 4.076279e+01 2.507797e+01 1.531253e+01 5.537856e+00 3.346360e+00
[6] 2.044106e+00 1.625127e+00 1.106829e+00 7.862013e-01 7.028069e-01
[11] 6.121548e-01 5.392358e-01 3.813225e-01 2.848237e-01 2.636818e-01
[16] 2.415583e-01 2.224249e-01 1.783012e-01 1.293819e-01 1.141377e-01
[21] 9.559165e-02 8.685013e-02 7.950005e-02 6.285928e-02 5.813735e-02
[26] 4.994494e-02 4.071874e-02 3.821716e-02 3.269847e-02 2.767458e-02
[31] 2.619478e-02 2.492990e-02 2.221603e-02 2.060769e-02 1.861592e-02
[36] 1.411699e-02 1.063725e-02 8.653785e-03 7.951038e-03 4.283570e-03
[41] 8.039042e-30pca_df <- data.frame(
PC1 = pca_results$x[, 1],
PC2 = pca_results$x[, 2],
AML_top$sampleMetadata
)ggplot(pca_df, aes(x = PC1, y = PC2, color = Treatment)) +
geom_point(size = 3) +
labs(title = "PCA on asin-sqrt prop.",
x = paste0("PC1 (", round(var_pct[1], 2), " %)"),
y = paste0("PC2 (", round(var_pct[2], 2), " %)")) +
theme_classic() +
theme(aspect.ratio = 1)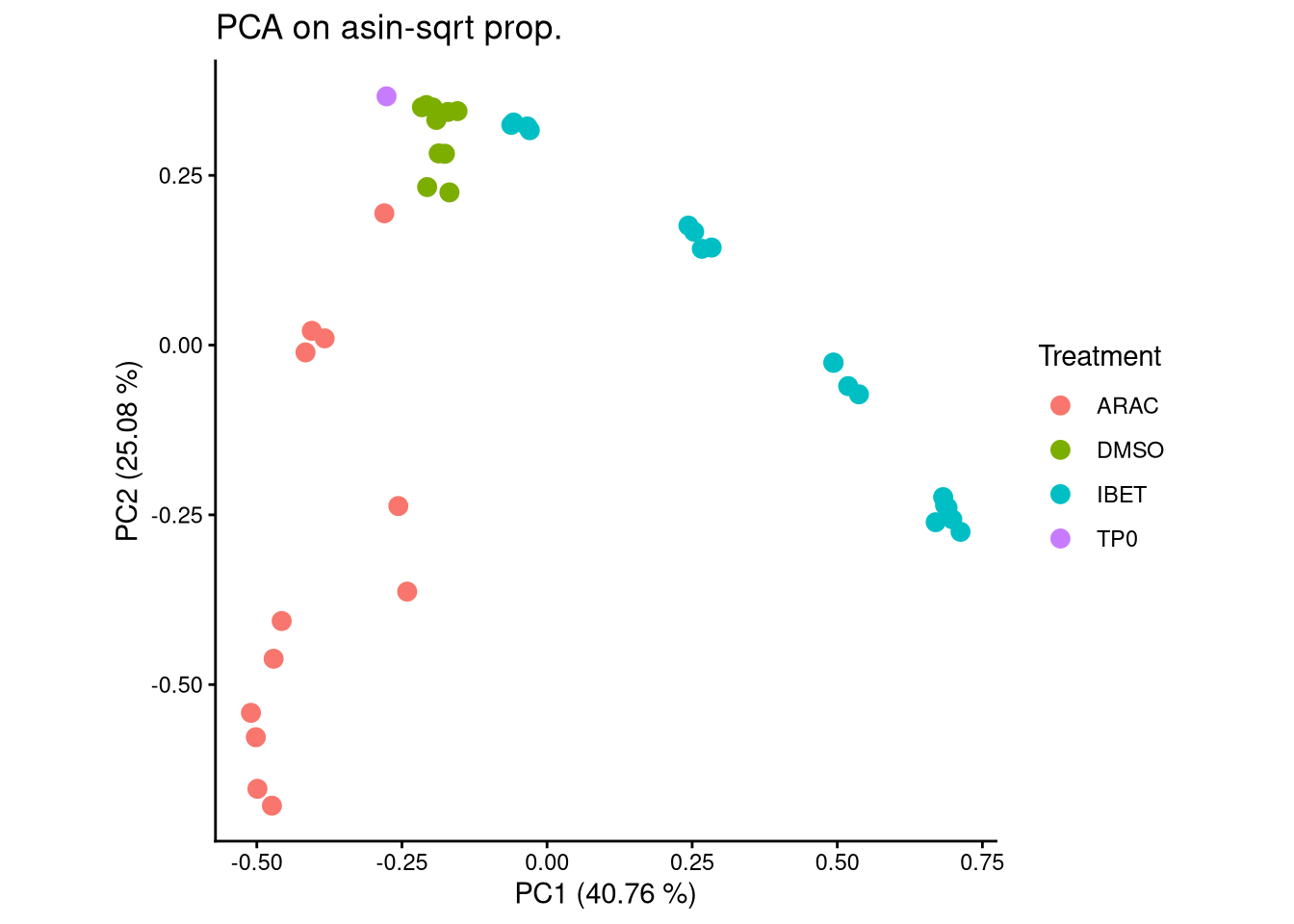
ggplot(pca_df, aes(x = PC1, y = PC2, color = Dose)) +
geom_point(size = 3) +
labs(title = "PCA on asin-sqrt prop.",
x = paste0("PC1 (", round(var_pct[1], 2), " %)"),
y = paste0("PC2 (", round(var_pct[2], 2), " %)")) +
theme_classic() +
theme(aspect.ratio = 1)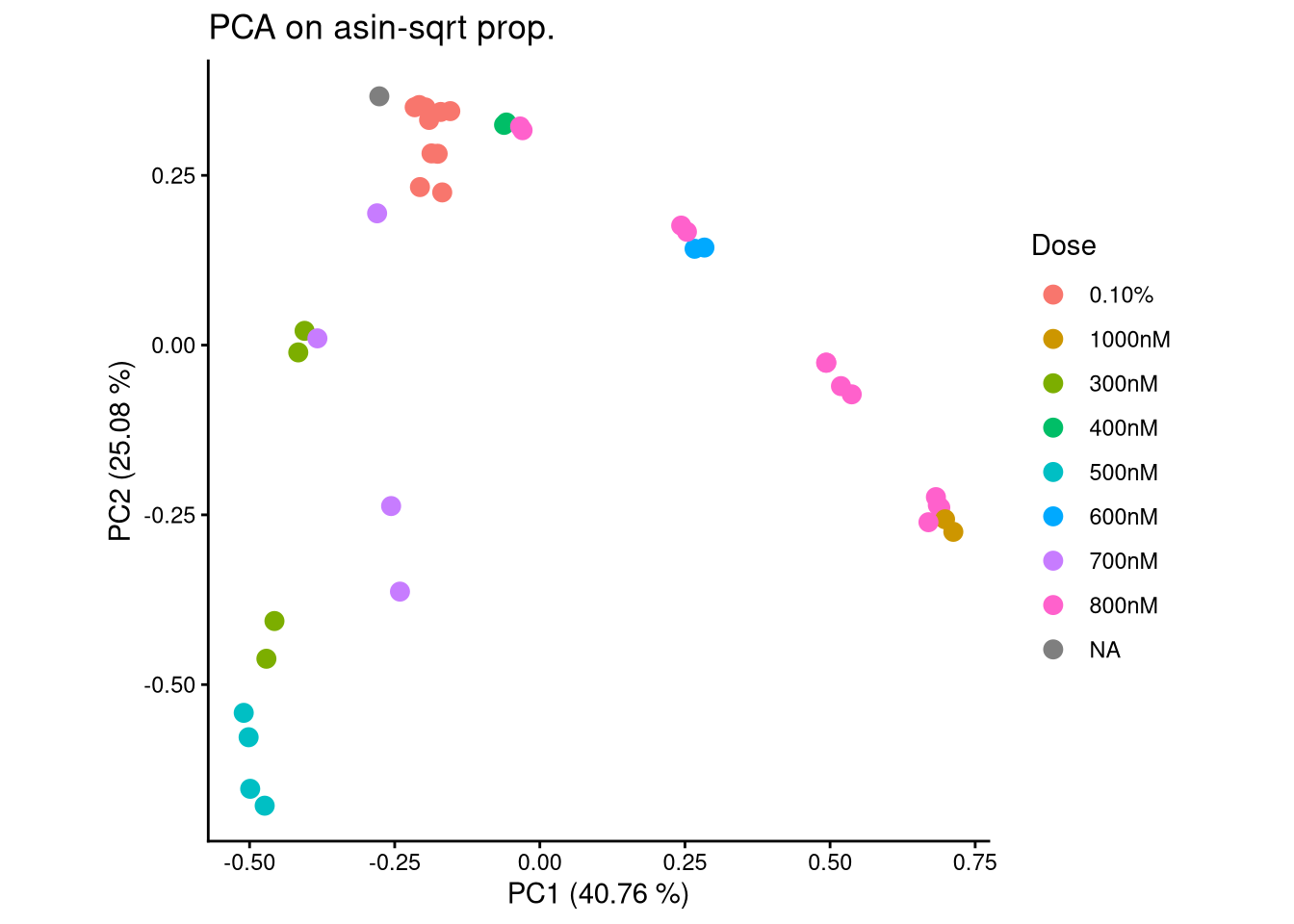
ggplot(pca_df, aes(x = PC1, y = PC2, color = Timepoint)) +
geom_point(size = 3) +
labs(title = "PCA on asin-sqrt prop.",
x = paste0("PC1 (", round(var_pct[1], 2), " %)"),
y = paste0("PC2 (", round(var_pct[2], 2), " %)")) +
theme_classic() +
theme(aspect.ratio = 1)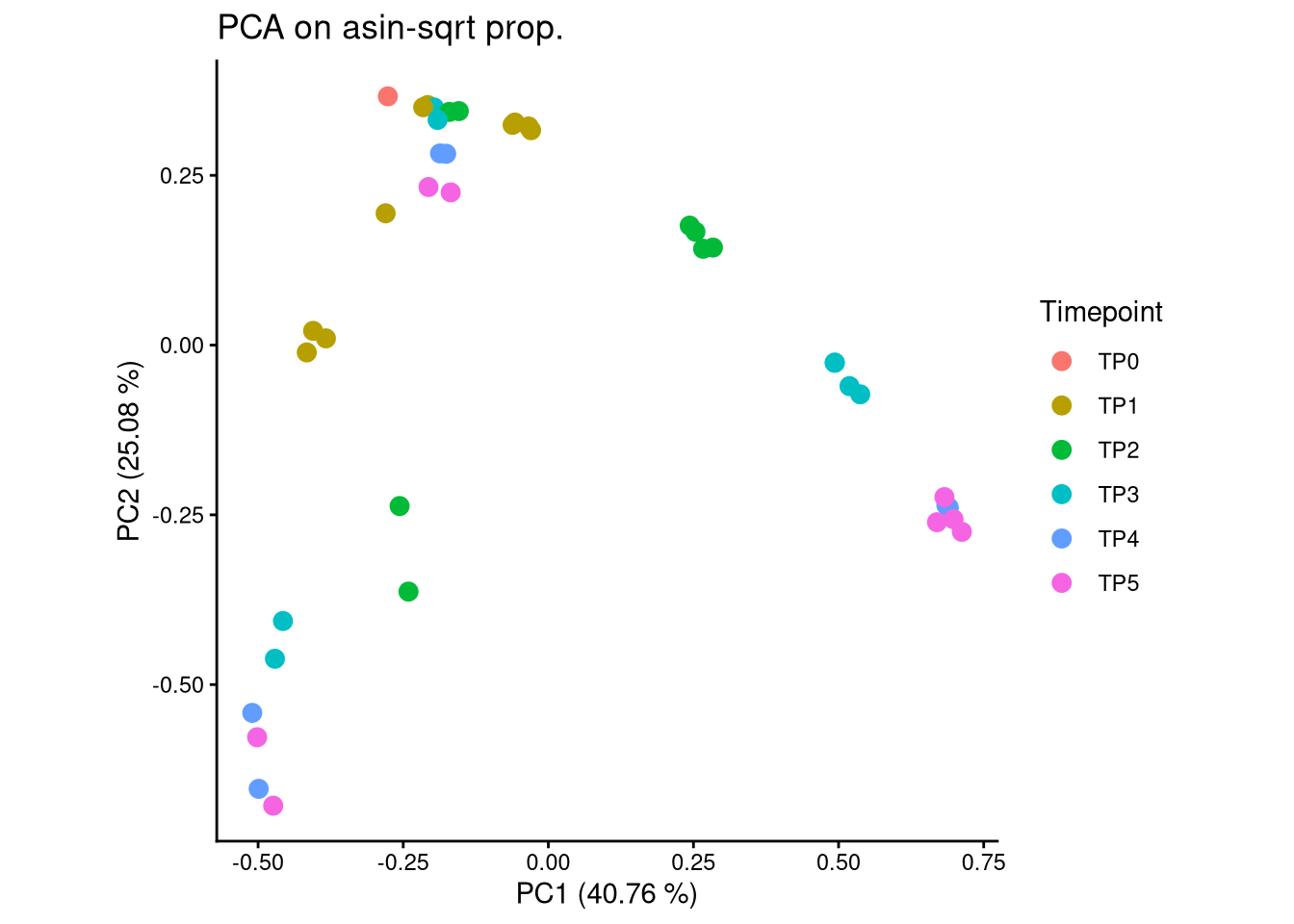
5.2 Plot propportion
## order samples
order_sample <- AML_top$sampleMetadata %>% with(order(Treatment, Dose, Timepoint))
## create bartools object
AML_dge <- DGEList(
counts = assay(AML_top),
group = AML_top$sampleMetadata$Treatment)
## plot
plotBarcodeHistogram(AML_dge, orderSamples = colnames(AML_top)[order_sample])Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"barcode"` instead of `.data$barcode`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"value"` instead of `.data$value`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"name"` instead of `.data$name`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.Warning: Use of .data in tidyselect expressions was deprecated in tidyselect 1.2.0.
ℹ Please use `"freq"` instead of `.data$freq`
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.Warning: The `guide` argument in `scale_*()` cannot be `FALSE`. This was deprecated in
ggplot2 3.3.4.
ℹ Please use "none" instead.
This warning is displayed once every 8 hours.
Call `lifecycle::last_lifecycle_warnings()` to see where this warning was
generated.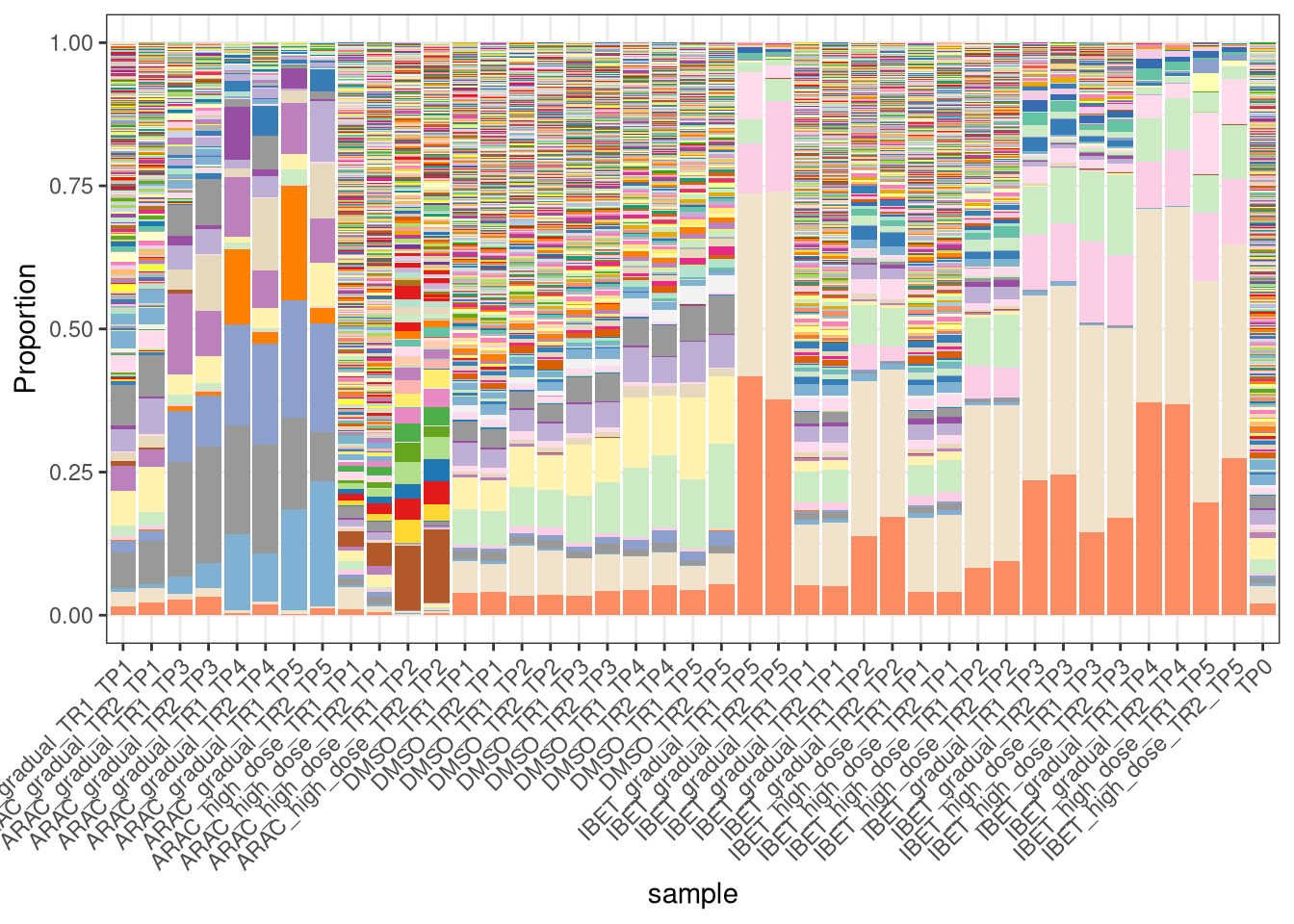
5.3 Subet DMSO
DMSO <- AML_top[, AML_top$sampleMetadata$Treatment == "DMSO"]pca_results <- (assays(DMSO)$proportion) %>% sqrt() %>% asin() %>%
t() %>%
prcomp()
var_pct <- pca_results$sdev^2 / sum(pca_results$sdev^2) * 100
var_pct [1] 7.752146e+01 9.395912e+00 4.157909e+00 3.336706e+00 2.553178e+00
[6] 1.079995e+00 8.330679e-01 7.617977e-01 3.599713e-01 4.692581e-30pca_dmso <- data.frame(
PC1 = pca_results$x[, 1],
PC2 = pca_results$x[, 2],
DMSO$sampleMetadata
)
ggplot(pca_dmso, aes(x = PC1, y = PC2, color = Timepoint)) +
geom_point(size = 3) +
labs(title = "PCA on asin-sqrt prop.",
x = paste0("PC1 (", round(var_pct[1], 2), " %)"),
y = paste0("PC2 (", round(var_pct[2], 2), " %)")) +
theme_classic() +
theme(aspect.ratio = 1)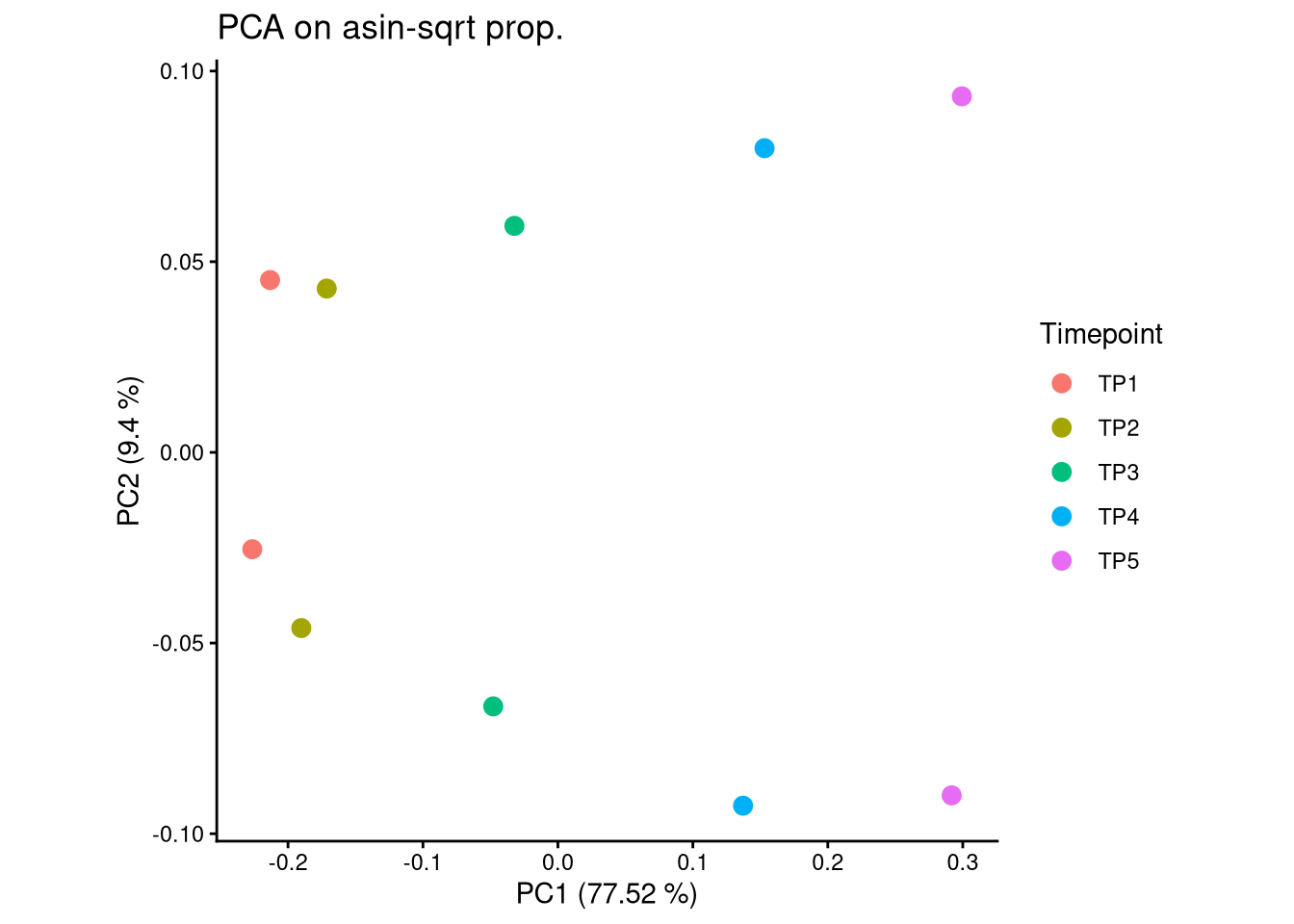
5.4 FS2G: prop. DMSO
## order samples
order_sample <- DMSO$sampleMetadata %>% with(order(Timepoint))
## create bartools object
DMSO_dge <- DGEList(
counts = assay(DMSO),
group = DMSO$sampleMetadata$Timepoint)
## plot
plotBarcodeHistogram(DMSO_dge, orderSamples = colnames(DMSO)[order_sample]) +
labs(title = "AML: DMSO") -> p_prop_DMSO
p_prop_DMSO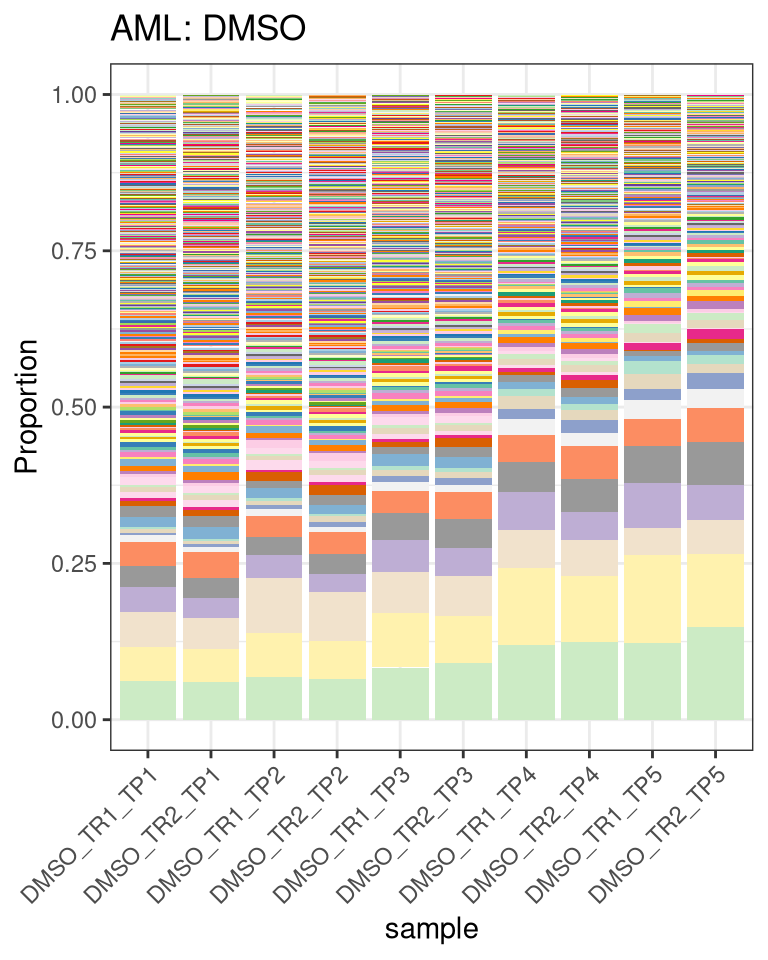
6 Type I error rate
6.1 Negative simulation
avoid running this chunk during rendering.
global_all_loops was saved from the first run.
results_negsi <- suppressMessages({
random_sampling(barbieQ = DMSO, loop_times = 100)
})
## `random_sampling()` save the simulations to global environment as `global_all_loops`
save(global_all_loops, file = "output/AML_negative_simulation.rda")- load results
## loading random sampling results to avoid running it repeatedly
load("output/AML_negative_simulation.rda")
# extract results from the loops
end_sampling <- floor(ncol(DMSO)/2)
## extract FPR
all_FPR <- lapply(seq(3:end_sampling), function(n) {
global_all_loops[[n]]$FPR
})
df_FPR_DMSO <- do.call(rbind, all_FPR)- plot FPR
df_FPR_DMSO$N_Samples <- as.factor(df_FPR_DMSO$N_Samples)
## Diff_Prop
## reshape the data to fit methods for testing
df_FPR_DMSO_long <- df_FPR_DMSO %>%
pivot_longer(cols = starts_with("Prop_"),
names_to = "Method",
values_to = "Pval_Prop") %>%
mutate(Method = factor(Method, levels = c("Prop_asin", "Prop_logit", "Prop_noTrans"),
labels = c("asin-sqrt", "logit", "no trans.")))
P_FPR_prop_DMSO <- ggplot(
df_FPR_DMSO_long, aes(x = factor(N_Samples), y = Pval_Prop, color = Method)) +
geom_boxplot(outlier.shape = 1, position = position_dodge(width = 0.7)) +
# geom_jitter(size = 1, position = position_dodge(width = 0.7)) +
geom_hline(yintercept = 0.05, linetype = "dashed", color = "black") +
# coord_cartesian(ylim = c(0, 0.2)) +
labs(
y = "Fraction of P-Value < 0.05",
x = "Number of samples per group",
# title = "DMSO",
subtitle = "Differential proportion test") +
# facet_wrap(~Method, scales = "free_y") +
theme_classic() +
theme(legend.position = "top")
## Diff_Occ
P_FPR_occ_DMSO <- ggplot(
df_FPR_DMSO, aes(x = factor(N_Samples), y = Occ_firth)) +
geom_boxplot(outlier.shape = 1, position = position_dodge(width = 0.7)) +
# geom_jitter(size = 1, position = position_dodge(width = 0.7)) +
geom_hline(yintercept = 0.05, linetype = "dashed", color = "black") +
scale_color_manual(values = c("Differential occurrrence test" = "black")) +
# coord_cartesian(ylim = c(0, 0.2)) +
labs(
y = "Fraction of P-Value < 0.05",
x = "Number of samples per group",
titlte = "",
subtitle = "Differential occurrence test") +
theme_classic() +
theme(legend.position = "top")6.2 FS2H: FPR
(P_FPR_prop_DMSO + theme(legend.position = "top")) + (P_FPR_occ_DMSO) -> P_FPR_DMSO
P_FPR_DMSOIgnoring unknown labels:
• titlte : ""Warning: No shared levels found between `names(values)` of the manual scale and the
data's colour values.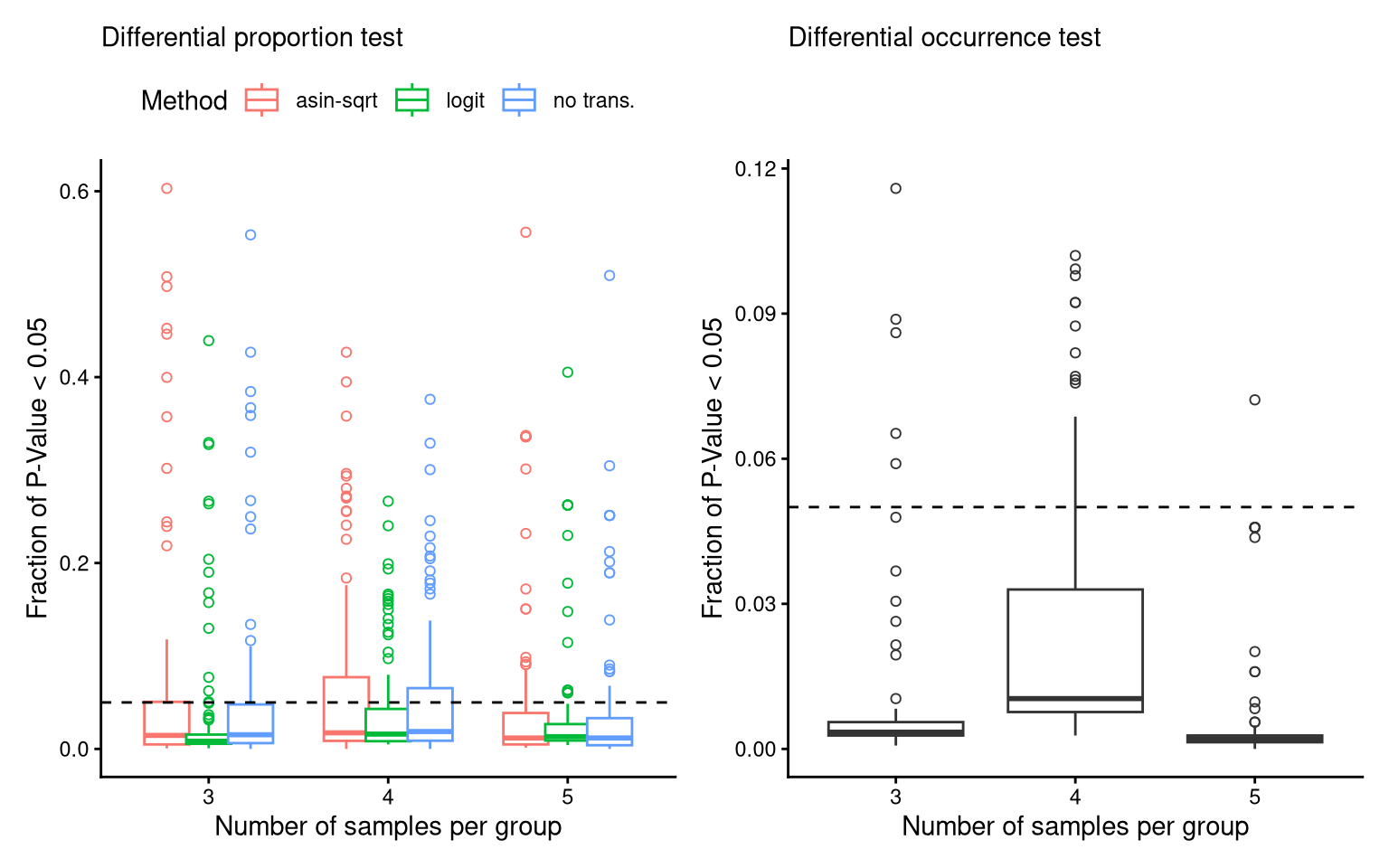
7 Figure S2-AML
layout = "
GHH
"
fs2_aml <- (
wrap_elements(p_prop_DMSO + theme(plot.margin = unit(c(0,0,0,0), "line"))) +
wrap_elements(P_FPR_DMSO + theme(legend.position = "top") + theme(plot.margin = unit(rep(0,4), "cm")))
) +
plot_layout(design = layout) +
plot_annotation(tag_levels = list(c("G", "H"))) &
theme(
plot.tag = element_text(size = 20, face = "bold", family = "arial"),
axis.title = element_text(size = 17),
axis.text = element_text(size = 12),
legend.title = element_text(size = 13),
legend.text = element_text(size = 11))
fs2_amlIgnoring unknown labels:
• titlte : ""Warning: No shared levels found between `names(values)` of the manual scale and the
data's colour values.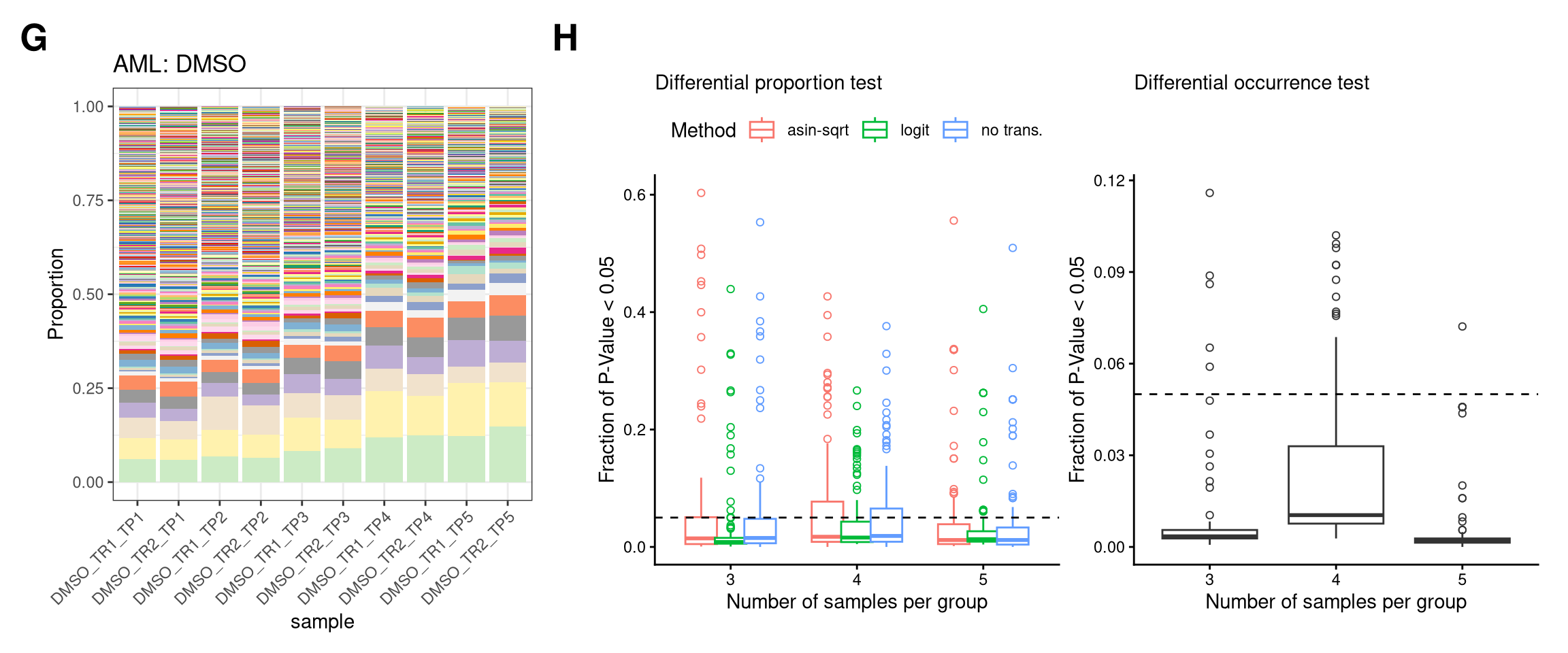
ggsave(
filename = "output/fs2_aml.png",
plot = fs2_aml,
width = 12,
height = 5,
units = "in", # for Rmd r chunk fig size, unit default to inch
dpi = 350
)Ignoring unknown labels:
• titlte : ""Warning: No shared levels found between `names(values)` of the manual scale and the
data's colour values.Saving this figure in fs2_AML
sessionInfo()R version 4.5.0 (2025-04-11)
Platform: x86_64-pc-linux-gnu
Running under: Red Hat Enterprise Linux 9.6 (Plow)
Matrix products: default
BLAS/LAPACK: FlexiBLAS OPENBLAS-OPENMP; LAPACK version 3.9.0
locale:
[1] LC_CTYPE=en_AU.UTF-8 LC_NUMERIC=C
[3] LC_TIME=en_AU.UTF-8 LC_COLLATE=en_AU.UTF-8
[5] LC_MONETARY=en_AU.UTF-8 LC_MESSAGES=en_AU.UTF-8
[7] LC_PAPER=en_AU.UTF-8 LC_NAME=C
[9] LC_ADDRESS=C LC_TELEPHONE=C
[11] LC_MEASUREMENT=en_AU.UTF-8 LC_IDENTIFICATION=C
time zone: Australia/Melbourne
tzcode source: system (glibc)
attached base packages:
[1] stats4 grid stats graphics grDevices utils datasets
[8] methods base
other attached packages:
[1] barbieQ_1.1.3 devtools_2.4.6
[3] usethis_3.2.1 SEtools_1.22.0
[5] sechm_1.16.0 SummarizedExperiment_1.38.1
[7] Biobase_2.68.0 GenomicRanges_1.60.0
[9] GenomeInfoDb_1.44.3 IRanges_2.42.0
[11] S4Vectors_0.48.0 BiocGenerics_0.54.0
[13] generics_0.1.4 MatrixGenerics_1.20.0
[15] matrixStats_1.5.0 edgeR_4.6.3
[17] limma_3.64.3 ComplexHeatmap_2.24.1
[19] ggVennDiagram_1.5.4 scales_1.4.0
[21] patchwork_1.3.2 ggplot2_4.0.0
[23] knitr_1.50 tibble_3.3.0
[25] tidyr_1.3.1 dplyr_1.1.4
[27] magrittr_2.0.4 readxl_1.4.5
[29] workflowr_1.7.2
loaded via a namespace (and not attached):
[1] splines_4.5.0 later_1.4.4 ggplotify_0.1.3
[4] cellranger_1.1.0 polyclip_1.10-7 rpart_4.1.24
[7] XML_3.99-0.20 lifecycle_1.0.4 Rdpack_2.6.4
[10] formula.tools_1.7.1 doParallel_1.0.17 rprojroot_2.1.1
[13] processx_3.8.6 lattice_0.22-6 MASS_7.3-65
[16] backports_1.5.0 openxlsx_4.2.8.1 sass_0.4.10
[19] rmarkdown_2.30 jquerylib_0.1.4 yaml_2.3.10
[22] remotes_2.5.0 httpuv_1.6.16 zip_2.3.3
[25] sessioninfo_1.2.3 pkgbuild_1.4.8 minqa_1.2.8
[28] DBI_1.2.3 RColorBrewer_1.1-3 abind_1.4-8
[31] pkgload_1.4.1 Rtsne_0.17 purrr_1.1.0
[34] ggraph_2.2.2 nnet_7.3-20 yulab.utils_0.2.1
[37] tweenr_2.0.3 rappdirs_0.3.3 git2r_0.36.2
[40] sva_3.56.0 circlize_0.4.16 seriation_1.5.8
[43] GenomeInfoDbData_1.2.14 ggrepel_0.9.6 genefilter_1.90.0
[46] pheatmap_1.0.13 annotate_1.86.1 codetools_0.2-20
[49] DelayedArray_0.34.1 ggforce_0.5.0 tidyselect_1.2.1
[52] shape_1.4.6.1 aplot_0.2.9 UCSC.utils_1.4.0
[55] farver_2.1.2 lme4_1.1-37 viridis_0.6.5
[58] TSP_1.2.6 jsonlite_2.0.0 GetoptLong_1.0.5
[61] mitml_0.4-5 ellipsis_0.3.2 tidygraph_1.3.1
[64] ggbreak_0.1.6 randomcoloR_1.1.0.1 survival_3.8-3
[67] iterators_1.0.14 systemfonts_1.3.1 foreach_1.5.2
[70] tools_4.5.0 ragg_1.5.0 Rcpp_1.1.0
[73] glue_1.8.0 pan_1.9 gridExtra_2.3
[76] SparseArray_1.8.1 xfun_0.53 mgcv_1.9-1
[79] DESeq2_1.48.2 logistf_1.26.1 ca_0.71.1
[82] withr_3.0.2 fastmap_1.2.0 boot_1.3-31
[85] callr_3.7.6 digest_0.6.37 R6_2.6.1
[88] gridGraphics_0.5-1 textshaping_1.0.3 mice_3.18.0
[91] colorspace_2.1-2 RSQLite_2.4.5 data.table_1.17.8
[94] graphlayouts_1.2.2 httr_1.4.7 S4Arrays_1.8.1
[97] whisker_0.4.1 pkgconfig_2.0.3 gtable_0.3.6
[100] blob_1.2.4 registry_0.5-1 S7_0.2.0
[103] XVector_0.48.0 htmltools_0.5.8.1 clue_0.3-66
[106] png_0.1-8 reformulas_0.4.1 ggfun_0.2.0
[109] rstudioapi_0.17.1 rjson_0.2.23 nloptr_2.2.1
[112] nlme_3.1-168 curl_7.0.0 cachem_1.1.0
[115] GlobalOptions_0.1.2 stringr_1.5.2 operator.tools_1.6.3
[118] parallel_4.5.0 AnnotationDbi_1.70.0 pillar_1.11.1
[121] vctrs_0.6.5 promises_1.3.3 jomo_2.7-6
[124] xtable_1.8-4 cluster_2.1.8.1 evaluate_1.0.5
[127] cli_3.6.5 locfit_1.5-9.12 compiler_4.5.0
[130] rlang_1.1.6 crayon_1.5.3 labeling_0.4.3
[133] ps_1.9.1 getPass_0.2-4 fs_1.6.6
[136] stringi_1.8.7 viridisLite_0.4.2 BiocParallel_1.42.2
[139] Biostrings_2.76.0 glmnet_4.1-10 V8_8.0.1
[142] Matrix_1.7-3 bit64_4.6.0-1 KEGGREST_1.48.1
[145] statmod_1.5.0 rbibutils_2.3 broom_1.0.10
[148] igraph_2.1.4 memoise_2.0.1 bslib_0.9.0
[151] bit_4.6.0